Abstract
Multicellularity is often considered a prerequisite for morphological complexity, as seen in the camera-type eyes found in several groups of animals. A notable exception exists in single-celled eukaryotes called dinoflagellates, some of which have an eye-like ‘ocelloid’ consisting of subcellular analogues to a cornea, lens, iris, and retina1. These planktonic cells are uncultivated and rarely encountered in environmental samples, obscuring the function and evolutionary origin of the ocelloid. Here we show, using a combination of electron microscopy, tomography, isolated-organelle genomics, and single-cell genomics, that ocelloids are built from pre-existing organelles, including a cornea-like layer made of mitochondria and a retinal body made of anastomosing plastids. We find that the retinal body forms the central core of a network of peridinin-type plastids, which in dinoflagellates and their relatives originated through an ancient endosymbiosis with a red alga2. As such, the ocelloid is a chimaeric structure, incorporating organelles with different endosymbiotic histories. The anatomical complexity of single-celled organisms may be limited by the components available for differentiation, but the ocelloid shows that pre-existing organelles can be assembled into a structure so complex that it was initially mistaken for a multicellular eye3. Although mitochondria and plastids are acknowledged chiefly for their metabolic roles, they can also be building blocks for greater structural complexity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
Transcriptomic data from Warnowia sp. and Erythropsidinium sp. have been deposited in GenBank under accession numbers KR632763–KR632773. Plastid genomic data from Nematodinium sp. have been deposited in GenBank under accession numbers KP765301–KP765306.
References
Greuet, C. Organisation ultrastructurale de l’ocelle de deux Peridiniens Warnowiidae, Erythropsis pavillardi Kofoid et Swezy et Warnowia pulchra Schiller. Protistologica 4, 209–230 (1968)
Janouskovec, J. et al. A common red algal origin of the apicomplexan, dinoflagellate, and heterokont plastids. Proc. Natl Acad. Sci. USA 107, 10949–10954 (2010)
Kofoid, C. A. & Swezy, O. The free-living, unarmoured dinoflagellates. Mem. Univ. Calif. 5, 1–562 (1921)
Dodge, J. D. The functional and phylogenetic significance of dinoflagellate eyespots. Biosystems 16, 259–267 (1984)
Kreimer, G. Reflective properties of different eyespot types in dinoflagellates. Protist 150, 311–323 (1999)
Greuet, C. Structural and ultrastructural evolution of ocelloid of Erythropsidinium-pavillardi, Kofoid-and-Swezy (dinoflagellate Warnowiidae, Lindemann) during division and palintomic divisions. Protistologica 13, 127–143 (1977)
Greuet, C. Structure fine de locelle d’Erythropsis pavillardi Hertwig, peridinien Warnowiidae Lindemann. C.R. Acad. Sci. 261, 1904–1907 (1965)
Hoppenrath, M. et al. Molecular phylogeny of ocelloid-bearing dinoflagellates (Warnowiaceae) as inferred from SSU and LSU rDNA sequences. BMC Evol. Biol. 9, 116 (2009)
Leander, B. S. Different modes of convergent evolution reflect phylogenetic distances. Trends Ecol. Evol. 23, 481–482 (2008)
Gehring, W. J. New perspectives on eye development and the evolution of eyes and photoreceptors. J. Hered. 96, 171–184 (2005)
Gomez, F., Lopez-Garcia, P. & Moreira, D. Molecular phylogeny of the ocelloid-bearing dinoflagellates Erythropsidinium and Warnowia (Warnowiaceae, Dinophyceae). J. Eukaryot. Microbiol. 56, 440–445 (2009)
Yoon, H. S. et al. Single-cell genomics reveals organismal interactions in uncultivated marine protists. Science 332, 714–717 (2011)
Lasken, R. S. Genomic sequencing of uncultured microorganisms from single cells. Nature Rev. Microbiol. 10, 631–640 (2012)
Kolisko, M. et al. Single-cell transcriptomics for microbial eukaryotes. Curr. Biol. 24, R1081–R1082 (2014)
Hofmann, E. et al. Structural basis of light harvesting by carotenoids: peridinin-chlorophyll-protein from Amphidinium carterae. Science 272, 1788–1791 (1996)
Keeling, P. J. The number, speed, and impact of plastid endosymbioses in eukaryotic evolution. Annu. Rev. Plant Biol. 64, 583–607 (2013)
Saldarriaga, J. F. et al. Dinoflagellate nuclear SSU rRNA phylogeny suggests multiple plastid losses and replacements. J. Mol. Evol. 53, 204–213 (2001)
Mornin, L. & Francis, D. Fine structure of Nematodinium armatum, a naked dinoflagellate. J. Microsc. 6, 759–772 (1967)
Lindberg, K., Moestrup, O. & Daugbjerg, N. Studies on woloszynskioid dinoflagellates I: Woloszynskia coronata re-examined using light and electron microscopy and partial LSU rDNA sequences, with description of Tovellia gen. nov. and Jadwigia gen. nov. (Tovelliaceae fam. nov.). Phycologia 44, 416–440 (2005)
Moestrup, O., Hansen, G. & Daugbjerg, N. Studies on woloszynskioid dinoflageflates III: on the ultrastructure and phylogeny of Borghiella dodgei gen. et sp nov., a cold-water species from Lake Tovel, N. Italy, and on B. tenuissima comb. nov. (syn. Woloszynskia tenuissima). Phycologia 47, 54–78 (2008)
Hausmann, K. Extrusive organelles in protists. Int. Rev. Cytol. 52, 197–276 (1978)
Abrahams, M. V. & Townsend, L. D. Bioluminescence in dinoflagellates: a test of the burglar alarm hypothesis. Ecology 74, 258–260 (1993)
Liu, J. & Kattawar, G. W. Detection of dinoflagellates by the light scattering properties of the chiral structure of their chromosomes. J. Quant. Spectrosc. Radiat. Transf. 131, 24–33 (2013)
Hayakawa, S. et al. Function and evolutionary origin of unicellular camera-type eye structure. PLoS ONE 10, http://dx.doi.org/10.1371/journal.pone.0118415 (2015)
Boisvert, S. et al. Ray: simultaneous assembly of reads from a mix of high-throughput sequencing technologies. J. Comput. Biol. 17, 1519–1533 (2010)
Langmead, B. & Salzberg, S. Fast gapped-read alignment with Bowtie 2. Nature Methods 9, 357–359 (2012)
Edgar, R. C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004)
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000)
Tamura, K. et al. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013)
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006)
Ronquist, F. et al. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012)
Acknowledgements
This work was supported by grants from the Natural Sciences and Engineering Research Council of Canada (2014-05258 to B.S.L., and 227301 to P.J.K.) and the Tula Foundation (Centre for Microbial Diversity and Evolution). We thank G. Owen for his operation of the FIB-SEM and G. Martens for preparing our samples for tomography. G.S.G. thanks S. Maslakova, C. Young, A. Lehman, and D. Blackburn for training in developmental biology, marine systems, electron microscopy, and ultrastructure, respectively. C.A.S., P.J.K. and B.S.L. are Senior Fellows of the Canadian Institute for Advanced Research.
Author information
Authors and Affiliations
Contributions
G.S.G., S.H., P.J.K. and B.S.L. designed the experiments. G.S.G. performed light microscopy, TEM, FIB-SEM, dissected-organelle and single-cell genomics, and phylogenetic analyses on specimens he collected in Canada, with resources and funding from B.S.L. and P.J.K. S.H. performed light microscopy, TEM, and transcriptomics on specimens she collected in Japan with resources and funding from T.G., and was supported in Canada by P.J.K. and B.S.L. R.A.W. prepared genomic libraries for sequencing and participated in single-cell genomics with funding from C.A.S. G.S.G. and B.S.L. wrote the manuscript and all authors participated in the drafting process.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 TEM of thylakoid membranes in Nematodinium sp.
a, A small, peripheral plastid in Nematodinium sp. with typical thylakoids resembling peridinin plastids in other dinoflagellates. b, Thylakoids in the iris region of the ocelloid. c, Thylakoids in the iris positioned beside waveform membranes (w) of the retinal body, during interphase. d, A retinal body towards the end of interphase, in which the waveform membranes de-differentiate and are continuous with the typical thylakoids. Typical thylakoids are marked by arrows.
Extended Data Figure 2 Development in warnowiids
a, b, Light micrographs of several cells of Nematodinium sp., and Erythropsidinium sp., progressing from interphase (left) to division (right). Scale bars, 10 µm. c, TEM of membranes in the retinal body, during differentiated (left, Nematodinium sp.), transitional (middle, Erythropsidinium sp.), and de-differentiated modes (right, Nematodinium sp.). Scale bars, 200 nm. The double arrowhead marks a typical plastid; arrowheads mark the retinal bodies; arrows mark lenses that are de-differentiating.
Extended Data Figure 3 Transient thylakoids in the retinal body viewed with TEM.
a, b, Ocelloid in a cell of Nematodinium sp. near division. c–e, Ocelloid in cells of Erythropsidinium sp. during division. L, lens; t, thylakoids; asterisks, lipid droplets; arrows, waveform membranes.
Extended Data Figure 4 Light micrographs of warnowiids used in this study.
a, Still frame from a video of Warnowia sp. b, Erythropsidinium sp. c, Nematodinium sp. with a nematocyst (arrowhead). d, The ventral side of Nematodinium sp. showing red pigmentation of the retinal body. e, Epifluorescence image of the same cell and angle, showing red fluorescence of the retinal body excited by 505 nm light. f, Nematodinium sp. showing a bright spot of reflectivity (that is, ‘eyeshine’) (arrowhead) in the ocelloid. Scale bars, 10 μm.
Extended Data Figure 5 TEM of the cornea-like layer of mitochondria in the ocelloid of Nematodinium sp.
a, Low-magnification TEM of the ocelloid, with rectangles delimiting the areas of higher magnification shown in b–d. b–d, High magnifications of structures bordering the lens (L). Mitochondria, m; pigmented ring, p; retinal body, r.
Extended Data Figure 6 Individual ribosomal gene and photosystem protein gene trees.
For c and d, the photosystem genes for Nematodinium sp. were amplified from the retinal body of the ocelloid. Support values for all phylogenies were calculated from 100 bootstraps using maximum likelihood analysis. a, 18S ribosomal DNA gene phylogeny derived from a 1,717-bp alignment across 33 dinoflagellate taxa. b, 28S ribosomal DNA gene phylogeny derived from a 970-bp alignment across 43 dinoflagellate taxa. For both a and b, warnowiids are highlighted in yellow and Nematodinium sp. is highlighted in black. c, Photosystem I P700 apoprotein A2 (PsaB) protein phylogeny derived from a 508 amino acid (AA) alignment across 42 photosynthetic taxa. d, Photosystem II protein D1 (PsbA) protein phylogeny derived from a 360 AA alignment across 39 photosynthetic taxa. For c and d, dinoflagellates are shaded in grey, and Nematodinium sp. is highlighted in black.
Extended Data Figure 7 Individual photosystem protein trees.
All the photosystem genes from Nematodinium sp. were amplified from the retinal body of the ocelloid. Support values for all phylogenies were calculated from 100 bootstraps using maximum likelihood analysis. a, Photosystem II CP47 (PsbB) protein phylogeny derived from a 504 AA alignment across 38 photosynthetic taxa. b, Photosystem II protein D1 (PsbD) phylogeny derived from a 342 AA alignment across 42 photosynthetic taxa. c, Cytochrome b6 (PetB) protein phylogeny derived from a 216 AA alignment across 32 photosynthetic taxa. d, Cytochrome b6/f complex subunit 4 (PetD) protein phylogeny derived from an 161 AA alignment across 31 photosynthetic taxa. Dinoflagellates are shaded in grey, and Nematodinium sp. is highlighted in black.
Extended Data Figure 8 Continuity between the retinal body and the plastid network in Nematodinium sp.
a, FIB-SEM slice of plastids attached to retinal body. b, TEM overview of ocelloid in a high-pressure frozen cell. c, FIB-SEM overview of ocelloid in a high-pressure frozen cell. d, Three-dimensional reconstruction of the ocelloid shown halved. e, Three-dimensional reconstruction of the ocelloid in full. f, Fusion site between plastids joined to the retinal body as seen in TEM. g, Site where the waveform-membrane region of the ocelloid joins to a region with thylakoids as seen in TEM. Inset shows thylakoids, and corresponds to the box in the main image. h, Fusion site as seen through FIB-SEM. i, Tracing of membrane continuity in Amira. j, Partial reconstruction of the ocelloid in Amira. Arrowheads point to fusion zones between sites bounded by the plastid membrane (reconstructed in red), blue denotes mitochondria, yellow denotes the surface of the lens. L, lens; w, waveform membranes; t, thylakoids.
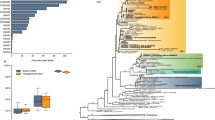
Extended Data Figure 9 Dinoflagellate eyespot types within a phylogenetic context.
Diagrams of whole cells and eyespots are shown for all dinoflagellates for which both ultrastructural descriptions and 18S and 28S ribosomal DNA sequences have been published. Eyespot diagrams highlight plastid-like structures (crimson), as well as mitochondria (dark blue), lens-like vesicles (light blue), lipid droplets (red dots), and crystalline layers (grey dashes). The phylogenetic tree was inferred from a 2,331-nucleotide alignment of concatenated 18S and 28S ribosomal DNA sequences across 36 genera; statistical support was evaluated with 500 bootstraps using maximum likelihood and 10,000 generations of Bayesian analysis. Bootstrap values above 60% are shown. For some taxa, 18S and 28S ribosomal sequences were concatenated from different species within the genus. Only the genus is shown for these taxa.
Extended Data Figure 10 Light micrographs and TEM showing food vacuoles in Nematodinium sp.
a, Differential interference contrast light micrographs showing a cell with prey (P) visible as green tinted food vacuole. b, Differential interference contrast light micrographs showing a cell in which the condensed dinoflagellate-type nuclei (n) are visible as birefringent chromosomes both in the predator and in the prey. c, Differential interference contrast light micrographs of a Nematodinium sp. cell containing digested prey (arrowhead) and co-occurring with potential prey, a smaller dinoflagellate. d, TEM showing a food vacuole inclusion consisting of a bolus of discharged trichocysts. e, TEM of undischarged dinoflagellate-type trichocysts showing their characteristic square shape in transverse section. f, TEM of discharged dinoflagellate-type trichocysts showing their characteristic striation pattern in longitudinal section.
Supplementary information
Erythropsidinium sp. in vivo.
Video of an Erythropsidinium cell moving with its characteristic “piston” appendage. (MOV 2386 kb)
Warnowia sp. in vivo.
Video of a Warnowia cell moving. (MOV 1612 kb)
FIB-SEM reconstruction of the ocelloid from Nematodinium sp.
From a stack of two-dimensional FIB-SEM images, the mitochondria (blue), lens (yellow), plastids (red), and flagellum (grey) were reconstructed as three-dimensional surfaces in Amira. (MPG 23047 kb)
Rights and permissions
About this article
Cite this article
Gavelis, G., Hayakawa, S., White III, R. et al. Eye-like ocelloids are built from different endosymbiotically acquired components. Nature 523, 204–207 (2015). https://doi.org/10.1038/nature14593
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14593
This article is cited by
-
Biophotonic probes for bio-detection and imaging
Light: Science & Applications (2021)
-
Dinoflagellate nucleus contains an extensive endomembrane network, the nuclear net
Scientific Reports (2019)
-
Flexible control of movement in plants
Scientific Reports (2019)
-
Towards a Dynamic Interaction Network of Life to unify and expand the evolutionary theory
BMC Biology (2018)
-
The evolution of eyes: major steps. The Keeler lecture 2017: centenary of Keeler Ltd
Eye (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.