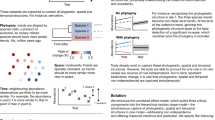
Abstract
A major challenge in theoretical ecology is understanding how natural microbial communities support species diversity1,2,3,4,5,6,7,8, and in particular how antibiotic-producing, -sensitive and -resistant species coexist9,10,11,12,13,14,15. While cyclic ‘rock–paper–scissors’ interactions can stabilize communities in spatial environments9,10,11, coexistence in unstructured environments remains unexplained12,16. Here, using simulations and analytical models, we show that the opposing actions of antibiotic production and degradation enable coexistence even in well-mixed environments. Coexistence depends on three-way interactions in which an antibiotic-degrading species attenuates the inhibitory interactions between two other species. These interactions enable coexistence that is robust to substantial differences in inherent species growth rates and to invasion by ‘cheating’ species that cease to produce or degrade antibiotics. At least two antibiotics are required for stability, with greater numbers of antibiotics enabling more complex communities and diverse dynamic behaviours ranging from stable fixed points to limit cycles and chaos. Together, these results show how multi-species antibiotic interactions can generate ecological stability in both spatially structured and mixed microbial communities, suggesting strategies for engineering synthetic ecosystems and highlighting the importance of toxin production and degradation for microbial biodiversity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hutchinson, G. E. The paradox of the plankton. Am. Nat. 95, 137–145 (1961).
May, R. M. Will a large complex system be stable? Nature 238, 413–414 (1972).
Levin, S. A. Dispersion and population interactions. Am. Nat. 108, 207–228 (1974).
Armstrong, R. A. & McGehee, R. Competitive exclusion. Am. Nat. 115, 151–170 (1980).
Levin, S. A. Community equilibria and stability, and an extension of the competitive exclusion principle. Am. Nat. 104, 413–423 (1970).
Stewart, F. M. & Levin, B. R. Partitioning of resources and the outcome of interspecific competition: a model and some general considerations. Am. Nat. 107, 171–198 (1973).
Levin, B. R. Coexistence of two asexual strains on a single resource. Science 175, 1272–1274 (1972).
Shoresh, N., Hegreness, M. & Kishony, R. Evolution exacerbates the paradox of the plankton. Proc. Natl Acad. Sci. USA 105, 12365–12369 (2008).
Durrett, R. & Levin, S. Spatial aspects of interspecific competition. Theor. Popul. Biol. 53, 30–43 (1998).
Czárán, T. L., Hoekstra, R. F. & Pagie, L. Chemical warfare between microbes promotes biodiversity. Proc. Natl Acad. Sci. USA 99, 786–790 (2002).
Kerr, B., Riley, M. A., Feldman, M. W. & Bohannan, B. J. M. Local dispersal promotes biodiversity in a real-life game of rock-paper-scissors. Nature 418, 171–174 (2002).
Reichenbach, T., Mobilia, M. & Frey, E. Mobility promotes and jeopardizes biodiversity in rock–paper–scissors games. Nature 448, 1046–1049 (2007).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nature Rev. Microbiol. 8, 15–25 (2010).
Chait, R., Vetsigian, K. & Kishony, R. What counters antibiotic resistance in nature? Nature Chem. Biol. 8, 2–5 (2012).
Conlin, P. L., Chandler, J. R. & Kerr, B. Games of life and death: antibiotic resistance and production through the lens of evolutionary game theory. Curr. Opin. Microbiol. 21, 35–44 (2014).
Green, J. & Bohannan, B. J. M. Spatial scaling of microbial biodiversity. Trends Ecol. Evol. 21, 501–507 (2006).
Vetsigian, K., Jajoo, R. & Kishony, R. Structure and evolution of Streptomyces interaction networks in soil and in silico. PLoS Biol. 9, e1001184 (2011).
Cordero, O. X. et al. Ecological populations of bacteria act as socially cohesive units of antibiotic production and resistance. Science 337, 1228–1231 (2012).
Raynaud, X. & Nunan, N. Spatial ecology of bacteria at the microscale in soil. PLoS ONE 9, e87217 (2014).
Perlin, M. H. et al. Protection of Salmonella by ampicillin-resistant Escherichia coli in the presence of otherwise lethal drug concentrations. Proc. R. Soc. Lond. B 276, 3759–3768 (2009).
Wright, G. D. Bacterial resistance to antibiotics: enzymatic degradation and modification. Adv. Drug Deliv. Rev. 57, 1451–1470 (2005).
Wootton, J. T. Indirect effects in complex ecosystems: recent progress and future challenges. J. Sea Res. 48, 1–16 (2002).
Ohgushi, T., Schmitz, O. & Holt, R. D. Trait-Mediated Indirect Interactions. (Cambridge Univ. Press, 2012).
Bollenbach, T., Quan, S., Chait, R. & Kishony, R. Nonoptimal microbial response to antibiotics underlies suppressive drug interactions. Cell 139, 707–718 (2009).
Dugatkin, L. A., Perlin, M., Lucas, J. S. & Atlas, R. Group-beneficial traits, frequency-dependent selection and genotypic diversity: an antibiotic resistance paradigm. Proc. R. Soc. Lond. B 272, 79–83 (2005).
Yurtsev, E. A., Chao, H. X., Datta, M. S., Artemova, T. & Gore, J. Bacterial cheating drives the population dynamics of cooperative antibiotic resistance plasmids. Mol. Syst. Biol. 9, 683 (2013).
Lenski, R. E. & Hattingh, S. E. Coexistence of two competitors on one resource and one inhibitor: a chemostat model based on bacteria and antibiotics. J. Theor. Biol. 122, 83–93 (1986).
Hsu, S. B. & Waltman, P. A survey of mathematical models of competition with an inhibitor. Math. Biosci. 187, 53–91 (2004).
Levin, S. A., Segel, L. A. & Adler, F. R. Diffuse coevolution in plant-herbivore communities. Theor. Popul. Biol. 37, 171–191 (1990).
Großkopf, T. & Soyer, O. S. Synthetic microbial communities. Curr. Opin. Microbiol. 18, 72–77 (2014).
D’Costa, V. M. Sampling the antibiotic resistome. Science 311, 374–377 (2006).
Walsh, F. & Duffy, B. The culturable soil antibiotic resistome: a community of multi-drug resistant bacteria. PLoS ONE 8, e65567 (2013).
Brook, I. The role of beta-lactamase-producing-bacteria in mixed infections. BMC Infect. Dis. 9, 202 (2009).
Shaw, K. J., Rather, P. N., Hare, R. S. & Miller, G. H. Molecular genetics of aminoglycoside resistance genes and familial relationships of the aminoglycoside-modifying enzymes. Microbiol. Rev. 57, 138–163 (1993).
Wetzstein, H. G., Stadler, M., Tichy, H. V., Dalhoff, A. & Karl, W. Degradation of ciprofloxacin by basidiomycetes and identification of metabolites generated by the brown rot fungus Gloeophyllum striatum. Appl. Environ. Microbiol. 65, 1556–1563 (1999).
Livermore, D. M. β-Lactamases in laboratory and clinical resistance. Clin. Microbiol. Rev. 8, 557–584 (1995).
Rafii, F. & Hansen, E. B. Isolation of nitrofurantoin-resistant mutants of nitroreductase-producing Clostridium sp. strains from the human intestinal tract. Antimicrob. Agents Chemother. 42, 1121–1126 (1998).
Hoshino, Y. et al. Monooxygenation of rifampicin catalyzed by the rox gene product of Nocardia farcinica: structure elucidation, gene identification and role in drug resistance. J. Antibiot. (Tokyo) 63, 23–28 (2010).
Chait, R., Craney, A. & Kishony, R. Antibiotic interactions that select against resistance. Nature 446, 668–671 (2007).
Chait, R., Shrestha, S., Shah, A. K., Michel, J.-B. & Kishony, R. A differential drug screen for compounds that select against antibiotic resistance. PLoS ONE 5, e15179 (2010).
Acknowledgements
We thank R. Ward, S. Levin, M. Elowitz, G. Bunin, A. Amir, S. Kryazhimskiy, Y. Gerardin, J. Meyer, N. Yin, E. Bairey, H. Chung and A. Palmer for critical feedback and discussions; M. Baym, M. Fischbach, M. Traxler, R. Chait, L. Stone and E. Gontag for strains. E.D.K. acknowledges government support under and awarded by the Department of Defense, Office of Naval Research, National Defense Science and Engineering Graduate (NDSEG) Fellowship, 32 CFR 168a. J.Z. acknowledges support from the Research Science Institute at MIT during the summer of 2011. K.V. acknowledges start-up funds from University of Wisconsin-Madison. R.K. acknowledges the support of a James S. McDonnell Foundation 21st Century Science Initiative in Studying Complex Systems Research Award 220020169, National Institutes of Health Grant R01GM081617, European Research Council Seventh Framework Programme ERC Grant 281891, Israeli Centers of Research Excellence I-CORE Program ISF Grant No. 152/11.
Author information
Authors and Affiliations
Contributions
E.D.K., K.V. and R.K. designed the study; E.D.K., K.V. and J.Z. did the experiments; E.D.K. and K.V. did the simulations and modelling; E.D.K., K.V. and R.K. analysed the data and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Antibiotic attenuation is widespread among natural antibiotic producing species isolated from soil.
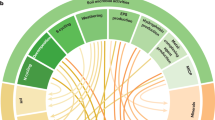
a, Diagram of three-species assays: measuring the antibiotic inhibition zone of E. coli around a producer strain enables quantification of three-species interactions caused by a modulator strain. b, Attenuating interactions dominate among a set of 54 Streptomyces producer–modulator combinations. Triangle and square markers show scoring from example images in Fig. 1b, left and right respectively. c, Antibiotic modulation assays: attenuating interactions dominate among combinations of soil species with a panel of ten pure antibiotics. Combinations are coloured by modulation index as in b and marked with a dot where the modulation index is significantly different from zero (n = 3, Methods). Strains 1 and 2 are the strongest modulators from the three-species assays, strain three is Streptomyces coelicolor, strains 4–18 are additional soil isolates (Extended Data Table 1). DOX, doxycycline; CMP, chloramphenicol; TOB, tobramycin; CPR, ciprofloxacin; FOX, cefoxitin; PIP, piperacillin; PEN, penicillin; NIT, nitrofurantoin; RIF, rifampicin; TMP, trimethoprim. d, Scatter plots of average modulation index M for 12 species from panel c, with and without β-lactamase inhibitors. Points occur off the diagonal for the β-lactams piperacillin and penicillin but near the diagonal for the structurally unrelated antibiotics rifampin and nitrofurantoin, consistent with attenuation of the β-lactams through a mechanism of antibiotic degradation. Error bars are standard error of the mean for technical replicas with inhibitor (n = 3) or without inhibitor (control, n = 6). Control is addition of H2O instead of inhibitor. Clav, clavulanic acid; tazB, tazobactum; sulB, sulbactam.
Extended Data Figure 2 Example images from three-species interaction assays and antibiotic assays.
a, Images and scoring for three-species interaction assays from Extended Data Fig. 1b. p indicates producer species, m indicates potential modulator species. b, Example images and scoring from the antibiotic modulation assays from Extended Data Fig. 1c. Each plate shows tests for modulation of antibiotic inhibition for three different species against a different antibiotic. Colour lines show the size of the inhibition zone at the location of the radially positioned modulator species; the white circles show the radius of the zone of inhibition as inferred from the left side of the plate that contains no modulators. c, Example images from an antibiotic modulation assay with β-lactamase inhibitors and the β-lactam antibiotic cefoxitin. Left and right side of each plate is inoculated with a line of species four from Extended Data Fig. 1c. Attenuation is significantly reduced by the β-lactamase inhibitors (especially clavulanic acid and tazobactam) when compared to controls (H2O).
Extended Data Figure 3 Illustration of the spatial inhibition-zone model.
The simulation is performed on a grid of size L × L. A single individual occupies each grid location. During each generation: (1) Individuals from species of type P produce antibiotics within a circle of radius rproduction; (2) individuals of type RD remove antibiotics within a circle of radius rdegradation. This process is repeated for each of the different antibiotics; (3) all sensitive individuals (type S) are killed at any locations that still contain the corresponding antibiotics (crossed out S; antibiotic values at each location are calculated at centre positions); (4) empty locations of a new grid are filled by randomly choosing from any surviving individuals within a radius rdispersal. If there are no surviving individuals within rdispersal then an empty location remains empty.
Extended Data Figure 4 Dependence of coexistence on degradation in the mixed inhibition-zone model.
a, Stability analysis of the full parameter space in the mixed inhibition-zone model. Using simulations, we tested the stability of cyclic three-species, three-antibiotic communities with dense sampling of all possible parameters for the inhibition-zone model, varying strengths of KP and KD, initial abundances of species 1–3 and growth ratios g2/g1 and g3/g1. As in Fig. 2b, each grid shows a 100-fold range of growth rate ratios from 0.1 to 10. Large basins of attraction exist across a wide range of parameter values, with maximal stability at high levels of antibiotic production and intermediate levels of degradation. b–d, Intuition for why coexistence depends on degradation. b, The inhibition-zone model calculates the probability of a given sensitive species being inhibited by an antibiotic producer (blue), or being protected by a degrading species (red), given the relative strengths of production (KP) and degradation (KD). The expected area covered by the overlapping circles (left) is used to calculate the corresponding inhibition and attenuation probabilities (percentage area of filled boxes, right; Methods). c, Focusing on one antibiotic in a stable three-species community, increasing the abundance of the yellow species creates negative feedback by decreasing the abundance of blue and red, which results in more inhibition of yellow. d, Communities are not stable at low levels of degradation due to positive feedback, whereby increasing the abundance of the yellow species results in a decrease of inhibition.
Extended Data Figure 5 Coexistence of antibiotic degrading communities in a well-mixed chemostat model with three antibiotics.
a, Stability of the three-species, three-antibiotic community in a single resource chemostat model in which species and antibiotics are completely homogeneous. Communities in the chemostat model may coexist through fixed points or limit cycles. For parameter sets in which the fixed point of the chemostat model was unstable, we started simulations close to the fixed point to determine if the community coexists through a limit cycle. Limit cycles occur in the areas between the dashed and solid black lines. Chemostat parameters: g1 = 3, g2 = 6, g3 = 9, kz = 0.5, P = 1. b, Communities coexist in the chemostat for a wide range of assumptions regarding antibiotic mechanisms. We simulated the chemostat model while changing how the action of the antibiotics is modelled and observed robust coexistence across all models. For each simulation we started all species at equal concentration (Xi = 0.2), ran the simulation until t = 100 generations and calculated the Shannon diversity of the final species levels. The default chemostat model assumes exponential inhibition of species by antibiotics, but similar coexistence is observed for Monod-like inhibition, for linear inhibition (for Gi < 0 we set Gi = 0), when species are only partially inhibited or when each species is sensitive at some level to all antibiotics. G1 is the growth rate of species 1 under inhibition, while g1 is its maximal rate of growth; R(x) captures resource dependence on current species levels; C1, C2 and C3 are the concentrations of antibiotics produced by species 1–3 respectively (Methods). Equations for the growth of species 2 and 3 have the same form as G1. All other parameters are the same as for panel a.
Extended Data Figure 6 Coexistence of communities with three species and two antibiotics.
a, Comparing community diversity in the mixed inhibition-zone model and the chemostat model. For each simulation we started all species at equal concentration (Xi = 1/3 for the inhibition-zone model, Xi = 0.2 for the chemostat model), ran the simulation for until t = 100 generations and calculated the Shannon diversity of the final species levels. Other parameters are the same as in Fig. 2 for the inhibition-zone model or Extended Data Fig. 5 for the chemostat model. b, Three species communities require two antibiotics for stability. When only one antibiotic is degraded the community either lacks stability (first panel), or is stable only for a small number of growth rates and initial conditions (second panel). When two antibiotics are degraded the community is robustly stable to differences in species growth rates and initial conditions (third panel), provided that the antibiotics inhibit the faster-growing species (species 2 and 3). Basin colours as in Fig. 2b; grey shows parameters for which no initial conditions were stable; KP = 40, KD = 4.
Extended Data Figure 7 Robustness of three species communities to invasion by cheaters in the mixed inhibition-zone model.
Analysis of production cheaters (P→RI, left) and degradation cheaters (RD→S, right). As in Fig. 3, we plot the final abundance of each cheater as a function of its growth advantage ε over its parent species. a, Cheaters cannot invade the three-species, three-antibiotic network when their growth advantage is small, except for the production cheater of the species with the fastest inherent growth rate (species 3, green line), which replaces its parent generating a new stable community of three species interacting through two antibiotics. b, This resulting three-species, two-antibiotic community is resilient to invasion by all cheaters; cheaters must have a substantial growth advantage to invade and take over. Parameters for both networks are the same as Fig. 3. Shaded areas indicate the maximum and minimum abundance when the community reaches stable oscillations. Note, the analysis above is for networks with g1 < g2 < g3. The alternative network of g1 > g2 > g3, is less robust to invasion by production cheaters. Two cheaters can invade this community even with small ε: the production cheater for species 2 invades the network in a and gives rise to a stable three-species two-antibiotic community, while the production cheater for species 1 can take over the communities shown in a and b. For degradation cheaters this alternative network is similarly robust to cheaters as shown.
Extended Data Figure 8 Complex network topologies support coexistence of larger numbers of species in the mixed inhibition-zone model.
For given initial numbers of species and antibiotics, sets of up to 106 communities with random networks were simulated and the final number of surviving species recorded. The number inside each square shows how many networks resulted in the specified number of final species (after removing networks that did not use all of the initial antibiotics, Methods). Colours show the frequency of each outcome within all simulated networks, with grey where no stable networks were found. We sparsely sampled parameters for species growth rates and antibiotic production and degradation levels (Methods). The sparse sampling means that a given network topology may exhibit stability for parameter combinations that were not tested.
Extended Data Figure 9 A community with chaotic dynamics.
Plotting the abundance of species 1–3 for the network from Fig. 4b. We show the last 30,000 steps of a simulation with 40,000 total generations, coloured with a slowly changing gradient. The trajectories form a strange attractor.
Supplementary information
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 3
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 3. (AVI 5928 kb)
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 6
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 6. (AVI 2996 kb)
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 20
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 20. (AVI 2996 kb)
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 80
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 80. (AVI 1041 kb)
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 200
Spatial inhibition-zone simulations, intrinsically resistant communities, dispersal radius = 200. (AVI 1041 kb)
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 3
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 3. (AVI 5928 kb)
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 6
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 6. (AVI 2996 kb)
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 20
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 20. (AVI 2996 kb)
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 80
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 80. (AVI 1041 kb)
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 200
Spatial inhibition-zone simulations, antibiotic degrading communities, dispersal radius = 200. (AVI 1041 kb)
Rights and permissions
About this article
Cite this article
Kelsic, E., Zhao, J., Vetsigian, K. et al. Counteraction of antibiotic production and degradation stabilizes microbial communities. Nature 521, 516–519 (2015). https://doi.org/10.1038/nature14485
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14485
This article is cited by
-
Wet-dry cycles protect surface-colonizing bacteria from major antibiotic classes
The ISME Journal (2022)
-
Synchronization induced by directed higher-order interactions
Communications Physics (2022)
-
Steering ecological-evolutionary dynamics to improve artificial selection of microbial communities
Nature Communications (2021)
-
Cooperation and competition between pair and multi-player social games in spatial populations
Scientific Reports (2021)
-
Bacterial siderophores in community and host interactions
Nature Reviews Microbiology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.