Abstract
Fundamental to all living organisms is the capacity to coordinate cell division and cell differentiation to generate appropriate numbers of specialized cells. Whereas eukaryotes use cyclins and cyclin-dependent kinases to balance division with cell fate decisions1, equivalent regulatory systems have not been described in bacteria. Moreover, the mechanisms used by bacteria to tune division in line with developmental programs are poorly understood. Here we show that Caulobacter crescentus, a bacterium with an asymmetric division cycle, uses oscillating levels of the second messenger cyclic diguanylate (c-di-GMP) to drive its cell cycle. We demonstrate that c-di-GMP directly binds to the essential cell cycle kinase CckA to inhibit kinase activity and stimulate phosphatase activity. An upshift of c-di-GMP during the G1–S transition switches CckA from the kinase to the phosphatase mode, thereby allowing replication initiation and cell cycle progression. Finally, we show that during division, c-di-GMP imposes spatial control on CckA to install the replication asymmetry of future daughter cells. These studies reveal c-di-GMP to be a cyclin-like molecule in bacteria that coordinates chromosome replication with cell morphogenesis in Caulobacter. The observation that c-di-GMP-mediated control is conserved in the plant pathogen Agrobacterium tumefaciens suggests a general mechanism through which this global regulator of bacterial virulence and persistence coordinates behaviour and cell proliferation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Malumbres, M. Cyclin-dependent kinases. Genome Biol. 15, 122 (2014).
Blanpain, C. & Simons, B. D. Unravelling stem cell dynamics by lineage tracing. Nature Rev. Mol. Cell Biol. 14, 489–502 (2013).
Ishidate, T., Elewa, A., Kim, S., Mello, C. C. & Shirayama, M. Divide and differentiate: CDK/Cyclins and the art of development. Cell Cycle 13, 1384–1391 (2014).
Morgan, D. O. Cyclin-dependent kinases: Engines, clocks, and microprocessors. Annu. Rev. Cell Dev. Biol. 13, 261–291 (1997).
Römling, U., Galperin, M. Y. & Gomelsky, M. Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Microbiol. Mol. Biol. Rev. 77, 1–52 (2013).
Abel, S. et al. Bi-modal distribution of the second messenger c-di-GMP controls cell fate and asymmetry during the Caulobacter cell cycle. PLoS Genet. 9, e1003744 (2013).
Paul, R. et al. Cell cycle-dependent dynamic localization of a bacterial response regulator with a novel di-guanylate cyclase output domain. Genes Dev. 18, 715–727 (2004).
Childers, W. S. et al. Cell fate regulation governed by a repurposed bacterial histidine kinase. PLoS Biol. 12, e1001979 (2014).
Tsokos, C. G., Perchuk, B. S. & Laub, M. T. A dynamic complex of signaling proteins uses polar localization to regulate cell-fate asymmetry in Caulobacter crescentus. Dev. Cell 20, 329–341 (2011).
Jacobs, C., Domian, I. J., Maddock, J. R. & Shapiro, L. Cell cycle-dependent polar localization of an essential bacterial histidine kinase that controls DNA replication and cell division. Cell 97, 111–120 (1999).
Biondi, E. G. et al. Regulation of the bacterial cell cycle by an integrated genetic circuit. Nature 444, 899–904 (2006).
Quon, K. C., Yang, B., Domian, I. J., Shapiro, L. & Marczynski, G. T. Negative control of bacterial DNA replication by a cell cycle regulatory protein that binds at the chromosome origin. Proc. Natl Acad. Sci. USA 95, 120–125 (1998).
Domian, I. J., Quon, K. C. & Shapiro, L. Cell type-specific phosphorylation and proteolysis of a transcriptional regulator controls the G1-to-S transition in a bacterial cell cycle. Cell 90, 415–424 (1997).
Chen, Y. E., Tsokos, C. G., Biondi, E. G., Perchuk, B. S. & Laub, M. T. Dynamics of two phosphorelays controlling cell cycle progression in Caulobacter crescentus. J. Bacteriol. 191, 7417–7429 (2009).
Duerig, A. et al. Second messenger-mediated spatiotemporal control of protein degradation regulates bacterial cell cycle progression. Genes Dev. 23, 93–104 (2009).
Kim, J., Heindl, J. E. & Fuqua, C. Coordination of division and development influences complex multicellular behavior in Agrobacterium tumefaciens. PLoS ONE 8, e56682 (2013).
Huang, Y.-H., Liu, X.-Y., Du, X.-X., Jiang, Z.-F. & Su, X.-D. The structural basis for the sensing and binding of cyclic di-GMP by STING. Nat. Struct. Mol. Biol. 19, 728–730 (2012).
Wheeler, R. T. & Shapiro, L. Differential localization of two histidine kinases controlling bacterial cell differentiation. Mol. Cell. 4, 683–694 (1999).
Paul, R. et al. Allosteric regulation of histidine kinases by their cognate response regulator determines cell fate. Cell 133, 452–461 (2008).
Choi, Y. J. & Anders, L. Signaling through cyclin D-dependent kinases. Oncogene 33, 1890–1903 (2014).
Hochegger, H., Takeda, S. & Hunt, T. Cyclin-dependent kinases and cell-cycle transitions: does one fit all? Nature Rev. Mol. Cell Biol. 9, 910–916 (2008).
Angelastro, P. S., Sliusarenko, O. & Jacobs-Wagner, C. Polar localization of the CckA histidine kinase and cell cycle periodicity of the essential master regulator CtrA in Caulobacter crescentus. J. Bacteriol. 192, 539–552 (2010).
Chen, Y. E. et al. Spatial gradient of protein phosphorylation underlies replicative asymmetry in a bacterium. Proc. Natl Acad. Sci. USA 108, 1052–1057 (2011).
Robinett, C. C. et al. In vivo localization of DNA sequences and visualization of large-scale chromatin organization using lac operator/repressor recognition. J. Cell Biol. 135, 1685–1700 (1996).
Paul, R. et al. Activation of the diguanylate cyclase PleD by phosphorylation-mediated dimerization. J. Biol. Chem. 282, 29170–29177 (2007).
Zähringer, F., Lacanna, E., Jenal, U., Schirmer, T. & Boehm, A. Structure and signaling mechanism of a zinc-sensory diguanylate cyclase. Structure 21, 1149–1157 (2013).
Danilchanka, O. & Mekalanos, J. J. Cyclic dinucleotides and the innate immune response. Cell 154, 962–970 (2013).
Capra, E. J. & Laub, M. T. Evolution of two-component signal transduction systems. Annu. Rev. Microbiol. 66, 325–347 (2012).
Brilli, M. et al. The diversity and evolution of cell cycle regulation in alpha-proteobacteria: a comparative genomic analysis. BMC Syst. Biol. 4, 52 (2010).
Ely, B. Genetics of Caulobacter crescentus. Methods Enzymol. 204, 372–384 (1991).
Thanbichler, M., Iniesta, A. A. & Shapiro, L. A comprehensive set of plasmids for vanillate- and xylose-inducible gene expression in Caulobacter crescentus. Nucleic Acids Res. 35, e137 (2007).
Viollier, P. H. et al. Rapid and sequential movement of individual chromosomal loci to specific subcellular locations during bacterial DNA replication. Proc. Natl Acad. Sci. USA 101, 9257–9262 (2004).
Bernhardt, T. G. & de Boer, P. A. J. Screening for synthetic lethal mutants in Escherichia coli and identification of EnvC (YibP) as a periplasmic septal ring factor with murein hydrolase activity. Mol. Microbiol. 52, 1255–1269 (2004).
Arellano, B. H., Ortiz, J. D., Manzano, J. & Chen, J. C. Identification of a dehydrogenase required for lactose metabolism in Caulobacter crescentus. Appl. Environ. Microbiol. 76, 3004–3014 (2010).
Navarre, W. W. et al. PoxA, yjeK, and elongation factor P coordinately modulate virulence and drug resistance in Salmonella enterica. Mol. Cell 39, 209–221 (2010).
Skerker, J. M., Prasol, M. S., Perchuk, B. S., Biondi, E. G. & Laub, M. T. Two-component signal transduction pathways regulating growth and cell cycle progression in a bacterium: a system-level analysis. PLoS Biol. 3, e334 (2005).
Christen, M. et al. DgrA is a member of a new family of cyclic diguanosine monophosphate receptors and controls flagellar motor function in Caulobacter crescentus. Proc. Natl Acad. Sci. USA 104, 4112–4117 (2007).
Pervushin, K., Riek, R., Wider, G. & Wüthrich, K. Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc. Natl Acad. Sci. USA 94, 12366–12371 (1997).
Salzmann, M., Pervushin, K., Wider, G., Senn, H. & Wüthrich, K. TROSY in triple-resonance experiments: new perspectives for sequential NMR assignment of large proteins. Proc. Natl Acad. Sci. USA 95, 13585–13590 (1998).
Zuiderweg, E. R. & Fesik, S. W. Heteronuclear three-dimensional NMR spectroscopy of the inflammatory protein C5a. Biochemistry 28, 2387–2391 (1989).
Quisel, J. D., Lin, D. C. & Grossman, A. D. Control of development by altered localization of a transcription factor in B. subtilis. Mol. Cell 4, 665–672 (1999).
Taylor, J. A., Ouimet, M.-C., Wargachuk, R. & Marczynski, G. T. The Caulobacter crescentus chromosome replication origin evolved two classes of weak DnaA binding sites. Mol. Microbiol. 82, 312–326 (2011).
Hung, D. Y. & Shapiro, L. A signal transduction protein cues proteolytic events critical to Caulobacter cell cycle progression. Proc. Natl Acad. Sci. USA 99, 13160–13165 (2002).
Kjaergaard, M. & Poulsen, F. M. Sequence correction of random coil chemical shifts: correlation between neighbor correction factors and changes in the Ramachandran distribution. J. Biomol. NMR 50, 157–165 (2011).
Hildebrand, A., Remmert, M., Biegert, A. & Söding, J. Fast and accurate automatic structure prediction with HHpred. Proteins 77 (Suppl. 9). 128–132 (2009).
Šali, A. & Blundell, T. L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Acknowledgements
We thank T. Sharp for help with protein analysis and F. Hamburger for plasmid constructions. S.O. is a recipient of a Japan Society for the Promotion of Science (JSPS) Postdoctoral Fellowships for research abroad. This work was supported by Swiss National Science Foundation grant 310030B_147090 to U.J. and an ERC Advanced Research Grant to U.J.
Author information
Authors and Affiliations
Contributions
C.L., S.O., S.A., S.S. and U.J. initiated the project. All authors designed experiments. S.O. carried out genetic experiments. C.L. and S.S. carried out biochemical experiments. C.L. and S.O. carried out microscopy experiments. R.B. and S.H. carried out NMR experiments. B.N.D. and T.S. contributed to structural analysis. U.J., S.O. and C.L. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Characterization of PdivK::Tn and Pxyl::divK derivatives.
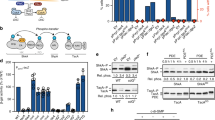
a, Schematic of the synthetic lethality screen. pBlue-pleD is a low copy number plasmid carrying both pleD and lacA genes, each with its own promoter. The lacA gene encodes a subunit of the LacABC dehydrogenase responsible for the breakdown of β-galactosides in C. crescentus34. An open arrowhead in the top panel indicates a representative blue colony on an X-gal agar plate. Out of 142,000 independent transformants, representative white colonies or colonies with blue sectors indicating segregation of the unstable pBlue-pleD plasmid are indicated in the upper and lower panel. Genomic organizations of the divK locus in strains SH100 and SH111 are shown schematically. The exact position of the transposon insertion (PdivK::Tn) in the divK promoter region adjacent to the CtrA box is indicated by closed arrowheads. The transcription start site (+1) and the −10 and −35 elements are shown43. Mapping of the transposon to the CtrA-binding site in the divK promoter region might imply that this lesion reduces divK expression by interfering with CtrA-mediated positive control11. b, Cell morphology and chromosome replication activity. Indicated strains were analysed microscopically and by flow cytometry to measure DNA content. Cells were grown with or without rifampicin (rif) as indicated. Chromosome equivalents (N) are indicated. Phase-contrast images are shown with scale bars of 5 µm. Representatives of two biological replicates are shown. c, DivK levels deduced by immunoblot analysis. Cells grown in peptone yeast extract were harvested at an OD660 of ∼0.2 and subjected to SDS-13% PAGE, followed by immunoblot analysis using anti-DivK antibodies. The intensities of the DivK bands were quantified using ImageJ and are shown as relative values to NA1000 wild-type levels. Representative of two biological replicates is shown. d, Subcellular levels of PleD, DivK and CtrA in the Pxyl::divK derivatives. Cells of strains NA1000, UJ5065, UJ8012 and UJ8013 grown in peptone yeast extract (none) or peptone yeast extract supplemented with 0.2% glucose or 0.03% xylose were analysed by immunoblots as indicated. The intensities of the protein bands were quantified using ImageJ and are shown as relative values to wild-type NA1000. The vector control (pMR20) is indicated. Representatives of two biological replicates are shown. e, Chromosome replication activity of wild-type (UJ8012) and cdG0 (UJ8013) strains expressing divK from the Pxyl promoter. Strains were grown exponentially in peptone yeast extract (none) or peptone yeast extract supplemented with glucose (Gluc), followed by flow cytometry analysis. Representatives of two biological replicates are shown. f, Effect of PdivK::Tn on cell morphology in strains lacking pleD, popA, or both. Scale bar, 5 µm. Representative of two biological replicates is shown.
Extended Data Figure 2 C-di-GMP binds to CckA to induce phosphatase activity.
a, C-di-GMP inhibits the CckA phosphorelay. In vitro phosphorylation reactions with purified proteins (+) in the presence or absence of c-di-GMP (75 µM). A CtrA mutant (D51E) lacking the phosphate-acceptor site is shown as a control. Phosphorylated proteins are marked ∼P. The weak band with a size similar to CtrA (lines 3 and 6) corresponds to a phosphorylated breakdown product of ChpT. Representatives of three technical replicates are shown. b, C-di-GMP stimulates CckA dephosphorylation. Phosphorylation reactions with purified CckA were carried out as outlined in Fig. 2a. After reaching saturation, dephosphorylation was initiated by the addition of increasing concentrations of c-di-GMP. Reactions were run for 15 min and were analysed by autoradiography. Representative results of two technical replicates are shown. c, C-di-GMP inhibits CckA autophosphorylation. Purified wild-type CckA and phosphatase mutant (V366P) were incubated with [32P]ATP and with (+) or without (−) c-di-GMP (75 µM) as indicated. C-di-GMP was added at the time points indicated. Representatives of three technical replicates are shown. d, C-di-GMP specifically binds to CckA. Purified CckA protein was incubated with 33P-labelled c-di-GMP and cross-linked with ultraviolet light in the presence or absence of a tenfold or 100-fold excess of competing non-labelled nucleotides as indicated. Representatives of three technical replicates are shown. e, The CA domain of CckA specifically binds c-di-GMP. Purified full length CckA (FL, lacking N-terminal transmembrane domains) and the minimal binding unit (see f) was incubated with 33P-labelled c-di-GMP and cross-linked with ultraviolet light in the presence or absence of a 100-fold excess of non-labelled ATP or c-di-GMP as indicated. Representatives of three technical replicates are shown. f, Schematic of the domain architecture and truncated constructs of CckA. Amino acids marking the boundaries of each construct are indicated. Constructs marked by green and red bars showed c-di-GMP binding or failed to bind c-di-GMP, respectively. g, Truncated versions of the CckA proteins indicated in a were expressed, purified and analysed for c-di-GMP binding using ultraviolet cross-linking of 33P-labelled c-di-GMP37 (10 µM) in the presence (+) or absence (–) of a 100-fold excess of non-labelled c-di-GMP (1 mM). Samples were analysed by SDS–PAGE and autoradiography as indicated. Representatives of two technical replicates are shown. h, Left, AtCckA, the CckA homologue of A. tumefaciens binds c-di-GMP. The c-di-GMP binding affinities of wild-type AtCckA and the AtCckA(Y674D) mutant protein were determined by ultraviolet cross-linking at increasing concentrations of [33P]c-di-GMP. Relative binding units and affinities are shown. Error bars are standard deviations. Averages and standard deviations were obtained from three technical replicates. Right, AtCckA is regulated by c-di-GMP. Wild-type AtCckA and AtCckA(Y674D) mutant were incubated with [32P]ATP (0 min) and supplemented with c-di-GMP and other nucleotides (75 µM) at 30 min. Fractions were removed after 30 min or 60 min as indicated and analysed by autoradiography. Representative of two technical replicates is shown. i, Phosphatase activity of CckA alleles in the absence of ATP. Reactions were allowed to autophosphorylate for 15 min before hexokinase and d-glucose were added to rapidly deplete ATP. A representative gel image for wild-type CckA is shown (top). Kinetic analysis revealed that CckA(Y514D) retains wild-type-like phosphatase activity (bottom). Error bars are standard deviations. Averages and standard deviations were obtained from three technical replicates. j, Mutational analysis of amino acids contributing to c-di-GMP binding and phosphatase control. Purified CckA wild-type and mutant forms were analysed for phosphorylation activity and [33P]c-di-GMP binding as indicated above. Note that the residue D479 is involved in ATP binding. Consequently, the D479A mutant lacks kinase activity, but is unaltered in its ability to bind c-di-GMP. Representatives of two technical replicates are shown.
Extended Data Figure 3 Characterization of the c-di-GMP binding site by NMR spectroscopy.
a, Top, 2D [15N,1H]TROSY spectrum of 0.38 mM CckA-CA recorded at 20 °C. The sequence-specific resonance assignments are indicated. Bottom left, sequence-specific secondary backbone 13C chemical shifts of CckA-CA relative to the random coil values of Kjaergaard et al.44. A 1–2–1 smoothing function was applied to the raw data. Consecutive stretches with positive and negative values indicate α-helical and β-strand secondary structure, respectively. The secondary structure elements inferred from these data are indicated above. Asterisks indicate unassigned residues. Bottom right, profile–profile alignment of the CA domains of CckA and DivL carried out with HHpred45 and formed the basis for the generation of the CckA homology model (shown in Fig. 2G) using the Modeller software46. The sequence identity is 25%. Secondary structure elements of CckA as determined by 13Cα secondary chemical shifts and of DivL, as derived from the crystal structure (PDB ID: 4Q20) are shown below the sequence alignment. The residue numbering of CckA is indicated. b, Chemical shift perturbation of CckA-CA backbone amide moieties upon c-di-GMP binding. Left, combined chemical shift changes of amide moieties, Δδ(HN), are plotted against the residue number. The magnitudes of 1 s.d. (0.021 p.p.m.) and 2 s.d. (0.042 p.p.m.) are indicated by a purple and pink line, respectively. The arrow points to residues I524 and H525 that experience intermediate chemical exchange upon c-di-GMP binding. Asterisks indicate unassigned residues. Right, region of a 2D [15N,1H]TROSY spectrum of a titration of c-di-GMP to CckA-CA at 20 °C. Sequence-specific resonance assignments are indicated.
Extended Data Figure 4 CLUSTALW alignment of the CA domain of CckA.
CLUSTALW was used to align the CA domain of CckA from C. crescentus and from different alphaproteobacteria. A fragment of the CA domain is shown that corresponds to amino acids 467–546 of C. crescentus CckA. Residues involved in c-di-GMP binding are boxed (green) and red bars above the sequence indicate regions with significant chemical shift in NMR spectroscopy upon c-di-GMP titration. CLUSTALW scores for conservation, quality and consensus are indicated.
Extended Data Figure 5 DivK and c-di-GMP convergently control C. crescentus growth and replication.
a, Model for the regulation of chromosome replication by the CckA kinase/phosphatase switch at reduced levels of DivK. Bold and dotted lines indicate strong and weak reactions, respectively. Dark circles indicate c-di-GMP. The kinase (Kin) and phosphatase modes (Pho) of CckA are indicated. i, C-di-GMP authorizes S-phase entry by inducing CckA phosphatase. ii, C-di-GMP is unable to bind to and induce phosphatase activity of CckA(Y514D) resulting in a G1 arrest. iii, C-di-GMP authorizes S-phase entry by reducing kinase activity of the phosphatase mutant CckA(V366P). iv, Cells lacking PleD fail to downregulate CckA(V366P) kinase activity. b, CtrA binding to the origin region is increased in cells containing cckA(Y514D) and PdivK::Tn. CtrA occupancy at the origin was analysed using chromatin immunoprecipitation (ChIP) and quantitative PCR (qPCR) as described in the Methods. Error bars are s.d. c, Combining the cckA(Y514D) and PdivK::Tn alleles leads to a G1 arrest. Exponential cultures of mutants containing different combinations of cckA, PdivK and pleD alleles were analysed by light microscopy and flow cytometry. Representative examples of two biological replicates for phase-contrast images and profiles of DNA content are shown with scale bars of 5 µm. d, Reduced divK expression in PxylX::divK strains containing the cckA(Y514D) allele leads to G1 arrest. Top, schematic of the chromosomal arrangement of cells expressing divK from the Pxyl promoter (PxylX::divK) and harbouring different cckA alleles. The divK gene is fused to the PxylX promoter in the xylX locus. The chromosomal copy of divK at the original locus was replaced with a Ω cassette. Different cckA alleles were introduced at the cckA locus by allelic exchange. Bottom left, cellular levels of DivK and PleD as determined by immunoblot analysis in strains grown in the presence or absence of xylose. Cells expressing divK from its own promoter at the native locus (PdivK) were used as control. Note that for PxylX::divK derivatives grown in peptone yeast extract, twice as many cells were used. Band intensities were determined with ImageJ and the respective values shown as relative units compared to wild type. Bottom right, DNA content per cell mass (DNA/mass) was analysed as described in Fig. 1c and values are shown below the graphs. Fractions of cells containing more than two chromosomes are indicated by brackets. Representatives of two biological replicates are shown. e, Colony-forming ability of PxylX::divK strains carrying different cckA alleles. Fivefold serial dilutions of the indicated strains were spotted and grown for 2 days at 30 °C on peptone yeast extract plates with the supplements indicated. Representatives of two biological replicates are shown. Note that these results are consistent with individual DNA replication profiles shown in Fig. 3 and panel d. f, Reduced divK expression in PxylX::divK strains containing the cckA(Y514D) allele leads to increased CckA phosphorylation levels. The cellular fraction of phosphorylated CckA (CckA∼P) was determined using Phos-tag gel electrophoresis. As a control, CckA and phosphorylated CckA levels were determined in synchronized populations of wild-type cells proceeding through the cell cycle (bottom). PxylX::divK strains harbouring different cckA alleles were analysed during exponential growth at 30 °C in the presence or absence of glucose (0.3%). The addition of glucose reduces leaky expression from the Pxyl promoter. Relative ratios of phosphorylated CckA to total CckA are shown. Error bars are s.d. Averages and standard deviations were obtained from three biological replicates. g, Single-cell analysis of the replication status of mutants with reduced DivK levels and abolished CckA control by c-di-GMP. Strains producing LacI–CFP and harbouring an array of lac operator (lacO) sites near the origin of replication were analysed32. Fluorescent repressor–operator system strains contained wild-type cckA or the cckA(Y514D) mutant allele, as well as PdivK::Tn-tet with wild-type pleD or ΔpleD as indicated. Representative phase-contrast and fluorescence images of two biological replicates are shown. Numbers of origins per cell length units were analysed statistically and the mean value and standard deviations obtained from two biological replicates are shown as a column graph. For each strain, a total of >900 cells were analysed using MicrobeTracker (http://microbetracker.org). Note that these results are consistent with the DNA replication profiles of equivalent strains without the FROS module shown in Fig. 3 and panels c and d.
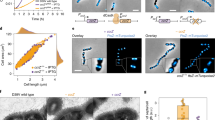
Extended Data Figure 6 C-di-GMP-mediated spatial control of CckA directs replication asymmetry in dividing cells.
a, b, Fluorescent repressor–operator system analysis was used to visualize DNA replication in individual cells. Dividing cells of wild-type C. crescentus (a) and cckA(Y514D) mutant (b) producing TetR–YFP and harbouring an array of tet operator (tetO) sites near the origin of replication were analysed by fluorescence microscopy. Frames from representative time-lapse movies used for panel c are shown. Stalked/old poles of newly divided daughter cells are marked with red arrows; newly replicated origins are marked with blue arrows. c, Spatial patterns of DNA replication were scored using a Tet-based fluorescent repressor–operator system and divided into three classes as indicated: replication initiation at the origin of replication located at the stalked pole (ST, orange), swarmer pole (SW, green), or at both poles (BI, blue). The bar diagram shows the quantification of wild-type and mutant strains with numbers indicating the percentage of cells falling into the three classes. The total number of cells analysed (n) is indicated above each bar. d, Expression cckAΔTM leads to c-di-GMP-dependent over-replication. Wild-type C. crescentus strains expressing different cckAΔTM alleles were analysed by light microscopy and flow cytometry as indicated. The fraction of cells bearing more than two chromosomes is indicated and shown as percentage. Representatives of two biological replicates are shown. e, Expression cckAΔTM leads to c-di-GMP-dependent over-replication. A C. crescentus cdG0 strain expressing cckAΔTM was analysed by light microscopy and flow cytometry as indicated. The fraction of cells bearing more than two chromosomes is indicated and shown as percentage. Representatives of two biological replicates are shown.
Supplementary information
Supplementary Table 1
This table contains the bacterial strains used in this study. (XLSX 36 kb)
Supplementary Table 2
This table contains the plasmids used in this study. (XLSX 26 kb)
Supplementary Table 3
This table contains the oligonucleotides used in this study. (XLSX 31 kb)
Rights and permissions
About this article
Cite this article
Lori, C., Ozaki, S., Steiner, S. et al. Cyclic di-GMP acts as a cell cycle oscillator to drive chromosome replication. Nature 523, 236–239 (2015). https://doi.org/10.1038/nature14473
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14473
This article is cited by
-
Functional diversity of c-di-GMP receptors in prokaryotic and eukaryotic systems
Cell Communication and Signaling (2023)
-
Structural features discriminating hybrid histidine kinase Rec domains from response regulator homologs
Nature Communications (2023)
-
Patterns of abundance, chromosomal localization, and domain organization among c-di-GMP-metabolizing genes revealed by comparative genomics of five alphaproteobacterial orders
BMC Genomics (2022)
-
GGDEF domain as spatial on-switch for a phosphodiesterase by interaction with landmark protein HubP
npj Biofilms and Microbiomes (2022)
-
Heme cross-feeding can augment Staphylococcus aureus and Enterococcus faecalis dual species biofilms
The ISME Journal (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.