Abstract
The mammalian hippocampus is crucial for episodic memory formation1 and transiently retains information for about 3–4 weeks in adult mice and longer in humans2. Although neuroscientists widely believe that neural synapses are elemental sites of information storage3, there has been no direct evidence that hippocampal synapses persist for time intervals commensurate with the duration of hippocampal-dependent memory. Here we tested the prediction that the lifetimes of hippocampal synapses match the longevity of hippocampal memory. By using time-lapse two-photon microendoscopy4 in the CA1 hippocampal area of live mice, we monitored the turnover dynamics of the pyramidal neurons’ basal dendritic spines, postsynaptic structures whose turnover dynamics are thought to reflect those of excitatory synaptic connections5,6. Strikingly, CA1 spine turnover dynamics differed sharply from those seen previously in the neocortex7,8,9. Mathematical modelling revealed that the data best matched kinetic models with a single population of spines with a mean lifetime of approximately 1–2 weeks. This implies ∼100% turnover in ∼2–3 times this interval, a near full erasure of the synaptic connectivity pattern. Although N-methyl-d-aspartate (NMDA) receptor blockade stabilizes spines in the neocortex10,11, in CA1 it transiently increased the rate of spine loss and thus lowered spine density. These results reveal that adult neocortical and hippocampal pyramidal neurons have divergent patterns of spine regulation and quantitatively support the idea that the transience of hippocampal-dependent memory directly reflects the turnover dynamics of hippocampal synapses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Squire, L. R. & Zola-Morgan, S. The medial temporal lobe memory system. Science 253, 1380–1386 (1991)
Frankland, P. W. & Bontempi, B. The organization of recent and remote memories. Nature Rev. Neurosci. 6, 119–130 (2005)
Frey, U. & Morris, R. G. Synaptic tagging and long-term potentiation. Nature 385, 533–536 (1997)
Barretto, R. P. et al. Time-lapse imaging of disease progression in deep brain areas using fluorescence microendoscopy. Nature Med. 17, 223–228 (2011)
Engert, F. & Bonhoeffer, T. Dendritic spine changes associated with hippocampal long-term synaptic plasticity. Nature 399, 66–70 (1999)
Maletic-Savatic, M., Malinow, R. & Svoboda, K. Rapid dendritic morphogenesis in CA1 hippocampal dendrites induced by synaptic activity. Science 283, 1923–1927 (1999)
Holtmaat, A. J. et al. Transient and persistent dendritic spines in the neocortex in vivo. Neuron 45, 279–291 (2005)
Xu, T. et al. Rapid formation and selective stabilization of synapses for enduring motor memories. Nature 462, 915–919 (2009)
Yang, G., Pan, F. & Gan, W. B. Stably maintained dendritic spines are associated with lifelong memories. Nature 462, 920–924 (2009)
Zuo, Y., Yang, G., Kwon, E. & Gan, W. B. Long-term sensory deprivation prevents dendritic spine loss in primary somatosensory cortex. Nature 436, 261–265 (2005)
Yang, G. et al. Sleep promotes branch-specific formation of dendritic spines after learning. Science 344, 1173–1178 (2014)
Harris, K. M. Structure, development, and plasticity of dendritic spines. Curr. Opin. Neurobiol. 9, 343–348 (1999)
Mizrahi, A., Crowley, J. C., Shtoyerman, E. & Katz, L. C. High-resolution in vivo imaging of hippocampal dendrites and spines. J. Neurosci. 24, 3147–3151 (2004)
Barretto, R. P., Messerschmidt, B. & Schnitzer, M. J. In vivo fluorescence imaging with high-resolution microlenses. Nature Methods 6, 511–512 (2009)
Gu, L. et al. Long-term in vivo imaging of dendritic spines in the hippocampus reveals structural plasticity. J. Neurosci. 34, 13948–13953 (2014)
Ziv, Y. et al. Long-term dynamics of CA1 hippocampal place codes. Nature Neurosci. 16, 264–266 (2013)
Harris, K. M. & Stevens, J. K. Dendritic spines of CA 1 pyramidal cells in the rat hippocampus: serial electron microscopy with reference to their biophysical characteristics. J. Neurosci. 9, 2982–2997 (1989)
Moser, M. B., Trommald, M. & Andersen, P. An increase in dendritic spine density on hippocampal CA1 pyramidal cells following spatial learning in adult rats suggests the formation of new synapses. Proc. Natl Acad. Sci. USA 91, 12673–12675 (1994)
Rampon, C. et al. Enrichment induces structural changes and recovery from nonspatial memory deficits in CA1 NMDAR1-knockout mice. Nature Neurosci. 3, 238–244 (2000)
Sanders, J., Cowansage, K., Baumgartel, K. & Mayford, M. Elimination of dendritic spines with long-term memory is specific to active circuits. J. Neurosci. 32, 12570–12578 (2012)
Bourne, J. N. & Harris, K. M. Coordination of size and number of excitatory and inhibitory synapses results in a balanced structural plasticity along mature hippocampal CA1 dendrites during LTP. Hippocampus 21, 354–373 (2011)
Huerta, P. T., Sun, L. D., Wilson, M. A. & Tonegawa, S. Formation of temporal memory requires NMDA receptors within CA1 pyramidal neurons. Neuron 25, 473–480 (2000)
Yasumatsu, N., Matsuzaki, M., Miyazaki, T., Noguchi, J. & Kasai, H. Principles of long-term dynamics of dendritic spines. J. Neurosci. 28, 13592–13608 (2008)
Toni, N., Buchs, P. A., Nikonenko, I., Bron, C. R. & Muller, D. LTP promotes formation of multiple spine synapses between a single axon terminal and a dendrite. Nature 402, 421–425 (1999)
Goshen, I. et al. Dynamics of retrieval strategies for remote memories. Cell 147, 678–689 (2011)
Sorra, K. E. & Harris, K. M. Stability in synapse number and size at 2 hr after long-term potentiation in hippocampal area CA1. J. Neurosci. 18, 658–671 (1998)
Fusi, S., Drew, P. J. & Abbott, L. F. Cascade models of synaptically stored memories. Neuron 45, 599–611 (2005)
Abraham, W. C. & Robins, A. Memory retention—the synaptic stability versus plasticity dilemma. Trends Neurosci. 28, 73–78 (2005)
Wu, X. E. & Mel, B. W. Capacity-enhancing synaptic learning rules in a medial temporal lobe online learning model. Neuron 62, 31–41 (2009)
Poirazi, P. & Mel, B. W. Impact of active dendrites and structural plasticity on the memory capacity of neural tissue. Neuron 29, 779–796 (2001)
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000)
Holtmaat, A. et al. Long-term, high-resolution imaging in the mouse neocortex through a chronic cranial window. Nature Protocols 4, 1128–1144 (2009)
Dombeck, D. A., Harvey, C. D., Tian, L., Looger, L. L. & Tank, D. W. Functional imaging of hippocampal place cells at cellular resolution during virtual navigation. Nature Neurosci. 13, 1433–1440 (2010)
Mishchenko, Y. et al. Ultrastructural analysis of hippocampal neuropil from the connectomics perspective. Neuron 67, 1009–1020 (2010)
Acknowledgements
We thank J. Li, J. Lecoq and E. T. W. Ho for technical assistance, R. P. J. Barretto and B. Messerschmidt for advice, and A. Holtmaat for providing published data sets. The National Institute of Mental Health, The National Institute on Aging, and the Ellison Foundation provided grants (M.J.S.). Fellowships from the National Science Foundation and The Stanford Center for Mind, Brain, and Computation provided graduate research support (J.E.F.).
Author information
Authors and Affiliations
Contributions
A.A. and M.J.S. designed experiments. A.A. performed experiments. A.A. and J.E.F. analysed data. J.E.F. designed and performed the modelling. A.A., J.E.F. and M.J.S. wrote the paper. M.J.S. supervised the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 In vivo imaging of CA1 spine dynamics over extended time intervals.
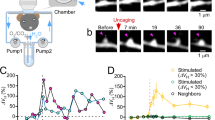
a, In vivo, 80-day-long time-lapse image data set sampled at variable intervals. Each image shown is the maximum projection of 4–8 images acquired at adjacent z-planes. Scale bar, 2 μm. b, c, Direct empirical determinations of spine density (b) and spine survival (c) across the 80 days. A normalized spine density of one corresponds to a measured spine density of 1.16 μm−1. Data points are mean ± s.e.m. for 16 dendrites. d, In vivo, 22-day-long time-lapse image data set sampled every 3 days. Over 50% of the spines underwent visually noticeable dynamic changes. Pie chart shows the proportions of spines that were persistent or exhibited different patterns of turnover (n = 1,075 total spines from 4 mice). Colour coding is the same as in Fig. 1d. Error bars are s.e.m. for four mice. e, Histograms show the distributions, for each class of spines, of the fraction of imaging sessions in which each spine was observed within the same 22-day data set of d. Colour coding is the same as in Fig. 1d. Error bars represent s.d. estimated as counting errors.
Extended Data Figure 2 The chronic CA1 preparation induces a minimal, 5-µm-thick layer of glial activation and does not affect spine density.
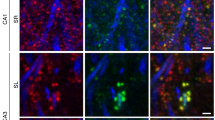
a–c, In two groups of mice, each comprising two Thy1-GFP transgenic animals (8–10 weeks old), we implanted the imaging guide tube just dorsal to hippocampal area CA1 following established procedures4,16,33. a, b, We euthanized, sliced and stained the first group after two further weeks (a), and the second group after five further weeks (b). Confocal fluorescence images of the stained tissue slices revealed activated microglia (CD68 staining, top, red), astrocytes (GFAP staining, bottom, red), GFP-expression pyramidal neurons (green), and permitted quantifications of spine density in CA1 regions both ipsilateral and contralateral to the implant. a, Two weeks after implantation, confocal photomicrographs (maximum intensity projection of four separate z-planes, axially spaced 0.2 µm apart) revealed a limited presence of activated microglia (red arrowheads indicate single cells) on the implanted hemisphere (top right) but were virtually non-existent in the contralateral control CA1 from the same mice (top left). Staining for astrocytes on the implanted hemisphere (bottom right) was almost indistinguishable from the control hemisphere (bottom left), except for a 5–10-µm-thick layer of astrocyte label abutting the optical surface of the imaging guide tube. b, Five weeks after implantation, confocal photomicrographs (maximum intensity projection of four separate z-planes, axially spaced 0.2 µm apart) revealed an almost undetectable presence of activated microglia on the implanted hemisphere (top right), comparable to the contralateral control hemisphere (top left). Staining for astrocytes in the implanted hemisphere (bottom right) was almost indistinguishable from the control hemisphere (bottom left), except for a 5–10-µm-thick layer of astrocyte label abutting the optical surface of the imaging guide tube. c, Mean density of spines on pyramidal cell basal dendrites in the implanted CA1 (white columns) was statistically indistinguishable from the contralateral control CA1 (black columns), at 2 weeks (P = 0.21; Mann–Whitney U-test; n = 11 dendrites) and at five weeks (P = 0.98; Mann–Whitney U-test; n = 11 dendrites) after implantation. Error bars are s.d.
Extended Data Figure 3 Two-photon and STED imaging of the same CA1 spines in fixed tissue reveals that nearby spines can merge in two-photon images.
a–c, Example two-photon microendoscopy (a), STED (b), and overlay (c) images of the same CA1 basal dendrite, acquired in a fixed brain slice from a Thy1-GFP mouse. Nearby spines that are clearly distinguishable in the STED image (green arrowheads) but within the diffraction-resolution limit of two-photon microendoscopy (0.85 NA) appear as single, merged entities within the two-photon image (red arrowheads). The two-photon image shown is the maximum intensity projection of three optical sections axially spaced 0.6 µm apart. The STED image is the maximum intensity projection of six optical sections spaced 0.3 µm apart. Scale, 2 µm. d, e, To attain an approximate measure of spine volume, we quantified each spine’s fluorescence in manually drawn regions of interest (ROIs) and normalized it by the fluorescence value in the nearby dendritic shaft, within an ROI of identical shape and size within a single axial section of the two-photon image stack. To ascertain whether each of the spines scored in the two-photon images was actually a merged spine or not, we consulted the STED images of the same dendrite. Plotted are the normalized fluorescence values (d) for unitary spines as well as doublet and triplet merged spines (black lines: mean values ± s.d.; coloured points: data from individual spines), and the cumulative distributions of these measurements (e). The distributions of normalized fluorescence were statistically indistinguishable (P > 0.06; Kruskal–Wallis ANOVA; N = 100, 30 and 7, respectively, for unitary, doublet and triplet spines), probably reflecting the substantial range of CA1 spine geometries (Extended Data Fig. 9).
Extended Data Figure 4 The asymptotic value of the surviving fraction of spines exceeds the fraction of permanent spines.
a, Surviving fraction curves for models in which the fraction of permanently stable spines is γ = 0 (red curve) and γ = 0.3 (purple curve) (Supplementary Methods). The timescale of spine survival was τ = 1, and the filling fraction value was f = 0.2. The surviving fraction asymptotes to a value, S∞, that encodes the fraction of stable spines. (Supplementary Information has a list of all mathematical variables used in this work, and their definitions). b, The time asymptotic value of the surviving fraction (green curve) exceeds the fraction of stable spines (dashed blue line). Coloured circles correspond to the surviving fraction curves plotted in a.
Extended Data Figure 5 Examples of simulated imaging data sets and their scoring.
a, Example simulated images (Supplementary Methods) of dendrites for which the spine density was 0.6 µm−1 (top), 1.5 µm−1 (middle) and 2.4 µm−1 (bottom). b, Ground truth and manual scoring of a simulated dendrite for which the spine density was 1.5 µm−1. Green arrowheads indicate counting errors originating from the optical merging of spines (Supplementary Methods). Blue arrowheads indicate counting errors that occur when the spine’s projection into the optical plane is too short (Supplementary Methods). Scale bar, 2 μm. c, The visually scored surviving fraction (red circles) differed from the true spine surviving fraction (blue triangles), but the departures were well predicted by the kinetic model (solid black curve) (Supplementary Methods). d, We simulated and scored a long-term lapse imaging data set with kinetic parameters that matched the best-fit model of Fig. 4b (Supplementary Methods). Even though the data set lacked stable spines, many simulated spines appeared to persist for long time intervals. Scale bar, 2 μm. e, Although the spine density in the simulated data was 2.56 µm−1, visual scoring yielded a lower spine density. We used the measured and true spine densities to estimate the extent of merging and the counting resolution (Supplementary Methods). Data points are mean ± s.e.m for 20 simulated dendrites (c) or 10 simulated dendrites (e).
Extended Data Figure 6 Variability of imaging angles has virtually no impact on determinations of spine turnover.
a–g, We examined empirically whether variations in dendritic angle across different imaging sessions might impact determinations of spine turnover. However, the variations in dendritic angle that were actually present in our data sets were insufficient to cause illusory turnover. a, b, Dendritic spines can be detected when the angle (θ) between a spine and the normal vector is large (Supplementary Methods). View of a dendrite and spine in the (x, z) (a) and optical (x, y) (b) planes. c, d, Dendritic spines cannot be detected when the angle between a spine and the normal vector (θ) is small (Supplementary Methods). View of a dendrite and spine in the (x, z) (c) and optical (x, y) (d), planes. e, For every dendrite and time point, we estimated the dendrite’s angle with respect to the optical plane using the three-dimensional coordinates of two manually labelled points on the dendrite chosen to flank the region of dendrite containing the scored spines. Over time, individual dendrites varied about their initial angle (n = 55 dendrites tracked over 16 sessions; data set of Fig. 3d). f, Distribution of the fluctuations in angle, pooled across the 55 dendrites, relative to the initial angle as seen in the first imaging session. The average magnitude of an angular fluctuation was 4.5°, and 90% of angular fluctuations were <10° in magnitude. Thus, a 5° fluctuation was typical in our data set, whereas a 10° fluctuation was atypically large. g, To determine if variability in the imaging angle might impact determinations of spine turnover, we imaged 18 dendrites in fixed slices while deliberately tilting the imaging plane by 0°, 5° and 10°. We made a total of 989 spine observations. Over 95% of spines scored in the 0° condition were also correctly scored when the specimen was tilted by 5° or 10°. Overall, the level of angular fluctuations in the in vivo imaging data has virtually no impact on turnover scores.
Extended Data Figure 7 Kinetic modelling well describes how optical merging affects spine turnover dynamics as monitored with finite optical resolution.
a, Diagram of the kinetic scheme used to describe merged spine dynamics (Supplementary Methods). Each state is labelled by the number of actual spines that have merged in appearance to a single spine (a quantity that we call the merged spine order; Supplementary Methods). State ‘0’ indicates the absence of a spine, and state ‘1’ indicates a spine that is truly unitary. Transitions occur between adjacent states in the kinetic ladder diagram with rate constants rmn. b, The rate constants governing increases in merged spine order depend on two parameters (Supplementary Methods): (1) the initial state or merged spine order; and (2) the overall degree of merging in the spine image data set, which is proportional to the product of the spine density and the shortest resolvable interspine interval (denoted L) (Supplementary Methods). By contrast, the rate constants governing decreases in merged spine order (inset) depend only on the initial merged spine order (Supplementary Methods). c, In the case when all spines are labile, a collapsed kinetic scheme in which a single state (Ψ) combines all merged spine orders above zero approximates the complete model (d) and can be solved mathematically (Supplementary Methods). d, The surviving fraction curve generated from the collapsed kinetic scheme (blue curve) fits the empirically observed surviving fraction (black data points) as well as the best-fit model (red). e, The asymptotic value of the surviving fraction is a function of the degree of merging (Supplementary Methods). A large degree of merging (as in CA1, blue circle) produces a larger asymptotic value of the merged spine surviving fraction than a small degree of merging (as in the neocortex, purple circle). f, The estimated lifetime of merged spines is a function of the degree of merging (Supplementary Methods). A large degree of merging (as in the hippocampal CA1, blue circle) produces a longer relative lifetime of merged spines than a small degree of merging (as in the neocortex, purple circle).
Extended Data Figure 8 Dynamic spine geometries induce modest levels of apparent spine turnover that cannot explain the turnover measured in vivo.
a–f, To study potential effects of fluctuations in spine geometry, we used values for the means and variances of dendrite radius, spine length and spine radius that were determined by electron microscopy17,34. We then computationally examined how time-dependent fluctuations in these parameters would affect determinations of spine surviving fraction (Supplementary Methods). a, We examined how fluctuations in dendrite radius, spine radius, spine length and spine angle—individually (coloured data points) and all together (black data)—affect the spine surviving fraction when the fluctuating geometric parameters are chosen stochastically in each of two imaging sessions, as a function of the parameter’s time correlation between the two sessions (Supplementary Methods). As expected, when the two sessions involved image pairs that were perfectly correlated, the surviving fraction reached 100%. Fluctuations in all four parameters had greater effects than fluctuations in individual geometric parameters. b, To estimate the time dependence of the surviving fraction of scorable spines from a, we assumed all geometric parameters evolved according to the time-correlation function that we empirically determined from in vivo imaging data (Extended Data Fig. 9d). c, The apparent surviving fraction is the product of the true surviving fraction and the surviving fraction of scorable spines (Supplementary Methods). For the best-fit kinetic model, the apparent surviving fraction is very close to the true surviving fraction. d, The difference between the fitted timescale of the apparent surviving fraction and the true survival timescale is small across the range of model parameters consistent with the in vivo data (Supplementary Methods). e, The graph plots the lower bound of the surviving fraction of scorable merged spines as a function of the time-correlation function shared by all four geometric parameters, for different merged spine orders. As this lower bound increases rapidly with the merged spine order, artefactual turnover due to unscorable spines is unlikely when spine merging is common (Supplementary Methods). f, We combined Fig. 4c and Extended Data Fig. 8e to bound the turnover that could result from unscorable spines (Supplementary Methods). As the empirically measured surviving fraction falls below the lower bound obtained for the surviving fraction of scorable merged spines, ongoing changes in the geometric parameters of spines cannot account for the observed spine turnover.
Extended Data Figure 9 Dynamics of spine geometries measured in vivo.
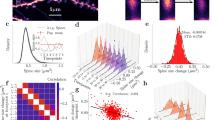
a, Time courses of the spine length, measured from the border of the dendritic shaft to the centre of the spine, for five example spines tracked over eight imaging sessions (left). Distribution of spine lengths (right; n = 344 spine observations). b, Time courses of the spine radius, measured from the border to the centre of the spine, for five example spines tracked over eight imaging sessions (left). Distribution of spine radii (right; n = 344 spine observations). c, Time courses of the dendritic radius, measured from the border to the centre of the dendrite, at the location of five example spines tracked over eight imaging sessions (left). Distribution of all dendritic radii (right; n = 344 dendrite observations). d, Experimental spine length time‐correlation function and its exponential fit (Supplementary Methods).
Extended Data Figure 10 Volumes of stable spines fluctuate minimally over time.
a–d, To attain an approximate measure of spine volume for stable spines, we quantified each spine’s fluorescence in manually drawn regions of interest (ROIs) and normalized it by the fluorescence value in the nearby dendritic shaft, as determined within a ROI of identical shape and size in a single z-section image acquired by two-photon microendoscopy. In addition, each spine’s fluorescence value at each time point is normalized to its own mean over the entire experiment. a, b, Mean (± s.e.m.) fluorescence intensities of all spines (a), and individual spines (b), from a set of 43 stable spines, across a 21-day span during which mice (n = 4) were in their home cages (same data as for Fig. 3a). Dashed black line in a indicates the mean over all imaging sessions. c, The correlation functions of spine radius (green), length (blue), fluorescence (black) and dendritic radius (red) are indistinguishable from each other. d, Mean (± s.e.m.) fluorescence intensities of 61 stable spines across a 46-day span during which mice (n = 3) initially resided in their home cages (black data points) but later moved to an enriched environment (red points) (same data set as for Fig. 3d). Dashed black and red lines respectively denote the mean values over the baseline and enriched periods.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary Appendices 1-3 and Supplementary References. (PDF 3399 kb)
Rights and permissions
About this article
Cite this article
Attardo, A., Fitzgerald, J. & Schnitzer, M. Impermanence of dendritic spines in live adult CA1 hippocampus. Nature 523, 592–596 (2015). https://doi.org/10.1038/nature14467
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14467
This article is cited by
-
Liprin-α proteins are master regulators of human presynapse assembly
Nature Neuroscience (2024)
-
Coordinated drift of receptive fields in Hebbian/anti-Hebbian network models during noisy representation learning
Nature Neuroscience (2023)
-
The Memory Orchestra: Contribution of Astrocytes
Neuroscience Bulletin (2023)
-
An interactive time series image analysis software for dendritic spines
Scientific Reports (2022)
-
A subpopulation of cortical VIP-expressing interneurons with highly dynamic spines
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.