Abstract
Ribosomes are translational machineries that catalyse protein synthesis. Ribosome structures from various species are known at the atomic level, but obtaining the structure of the human ribosome has remained a challenge; efforts to address this would be highly relevant with regard to human diseases. Here we report the near-atomic structure of the human ribosome derived from high-resolution single-particle cryo-electron microscopy and atomic model building. The structure has an average resolution of 3.6 Å, reaching 2.9 Å resolution in the most stable regions. It provides unprecedented insights into ribosomal RNA entities and amino acid side chains, notably of the transfer RNA binding sites and specific molecular interactions with the exit site tRNA. It reveals atomic details of the subunit interface, which is seen to remodel strongly upon rotational movements of the ribosomal subunits. Furthermore, the structure paves the way for analysing antibiotic side effects and diseases associated with deregulated protein synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Anger, A. M. et al. Structures of the human and Drosophila 80S ribosome. Nature 497, 80–85 (2013)
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334, 1524–1529 (2011)
Rabl, J., Leibundgut, M., Ataide, S. F., Haag, A. & Ban, N. Crystal structure of the eukaryotic 40S ribosomal subunit in complex with initiation factor 1. Science 331, 730–736 (2011)
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948 (2011)
Spahn, C. M. et al. Cryo-EM visualization of a viral internal ribosome entry site bound to human ribosomes: the IRES functions as an RNA-based translation factor. Cell 118, 465–475 (2004)
Boehringer, D., Thermann, R., Ostareck-Lederer, A., Lewis, J. D. & Stark, H. Structure of the hepatitis C virus IRES bound to the human 80S ribosome: remodeling of the HCV IRES. Structure 13, 1695–1706 (2005)
Chandramouli, P. et al. Structure of the mammalian 80S ribosome at 8.7 Å resolution. Structure 16, 535–548 (2008)
Voorhees, R. M., Fernandez, I. S., Scheres, S. H. & Hegde, R. S. Structure of the mammalian ribosome-Sec61 complex to 3.4 Å resolution. Cell 157, 1632–1643 (2014)
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014)
Greber, B. J. et al. The complete structure of the large subunit of the mammalian mitochondrial ribosome. Nature 515, 283–286 (2014)
Brown, A. et al. Structure of the large ribosomal subunit from human mitochondria. Science 346, 718–722 (2014)
Li, X., Zheng, S. Q., Egami, K., Agard, D. A. & Cheng, Y. Influence of electron dose rate on electron counting images recorded with the K2 camera. J. Struct. Biol. 184, 251–260 (2013)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Bai, X. C., Fernandez, I. S., McMullan, G. & Scheres, S. H. Ribosome structures to near-atomic resolution from thirty thousand cryo-EM particles. eLife 2, e00461 (2013)
Khatter, H. et al. Purification, characterization and crystallization of the human 80S ribosome. Nucleic Acids Res. 42, e49 (2014)
DeLaBarre, B. & Brunger, A. T. Considerations for the refinement of low-resolution crystal structures. Acta Crystallogr. D 62, 923–932 (2006)
Afonine, P. V. et al. FEM: feature-enhanced map. Acta Crystallogr. D 71, 646–666 (2015)
Ben-Shem, A., Jenner, L., Yusupova, G. & Yusupov, M. Crystal structure of the eukaryotic ribosome. Science 330, 1203–1209 (2010)
Fei, J., Kosuri, P., MacDougall, D. D. & Gonzalez, R. L., Jr Coupling of ribosomal L1 stalk and tRNA dynamics during translation elongation. Mol. Cell 30, 348–359 (2008)
Cornish, P. V. et al. Following movement of the L1 stalk between three functional states in single ribosomes. Proc. Natl Acad. Sci. USA 106, 2571–2576 (2009)
Feng, S., Chen, Y. & Gao, Y. G. Crystal structure of 70S ribosome with both cognate tRNAs in the E and P sites representing an authentic elongation complex. PLoS ONE 8, e58829 (2013)
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003)
Frank, J. & Agrawal, R. K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000)
Budkevich, T. V. et al. Regulation of the mammalian elongation cycle by subunit rolling: a eukaryotic-specific ribosome rearrangement. Cell 158, 121–131 (2014)
Budkevich, T. et al. Structure and dynamics of the mammalian ribosomal pretranslocation complex. Mol. Cell 44, 214–224 (2011)
Chen, J., Petrov, A., Tsai, A., O’Leary, S. E. & Puglisi, J. D. Coordinated conformational and compositional dynamics drive ribosome translocation. Nature Struct. Mol. Biol. 20, 718–727 (2013)
Leontis, N. B. & Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 7, 499–512 (2001)
Klaholz, B. & Moras, D. C.-H. O hydrogen bonds in the nuclear receptor RARgamma–a potential tool for drug selectivity. Structure 10, 1197–1204 (2002)
Liu, A. et al. NMR detection of bifurcated hydrogen bonds in large proteins. J. Am. Chem. Soc. 130, 2428–2429 (2008)
Jiang, L. & Lai, L. C. H. O hydrogen bonds at protein-protein interfaces. J. Biol. Chem. 277, 37732–37740 (2002)
Schuwirth, B. S. et al. Structures of the bacterial ribosome at 3.5 Å resolution. Science 310, 827–834 (2005)
Cate, J. H., Yusupov, M. M., Yusupova, G. Z., Earnest, T. N. & Noller, H. F. X-ray crystal structures of 70S ribosome functional complexes. Science 285, 2095–2104 (1999)
Yusupov, M. M. et al. Crystal structure of the ribosome at 5.5 Å resolution. Science 292, 883–896 (2001)
Shenvi, C. L., Dong, K. C., Friedman, E. M., Hanson, J. A. & Cate, J. H. Accessibility of 18S rRNA in human 40S subunits and 80S ribosomes at physiological magnesium ion concentrations–implications for the study of ribosome dynamics. RNA 11, 1898–1908 (2005)
Moore, M. N. & Spremulli, L. L. Effects of cations and cosolvents on eukaryotic ribosomal subunit conformation. Biochemistry 24, 191–196 (1985)
Sperrazza, J. M., Russell, D. W. & Spremulli, L. L. Reversible dissociation of wheat germ ribosomal subunits: cation-dependent equilibria and thermodynamic parameters. Biochemistry 19, 1053–1058 (1980)
Zhang, W., Dunkle, J. A. & Cate, J. H. Structures of the ribosome in intermediate states of ratcheting. Science 325, 1014–1017 (2009)
Dunkle, J. A. et al. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science 332, 981–984 (2011)
Tourigny, D. S., Fernandez, I. S., Kelley, A. C. & Ramakrishnan, V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science 340, 1235490 (2013)
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006)
Garreau de Loubresse, N. et al. Structural basis for the inhibition of the eukaryotic ribosome. Nature 513, 517–522 (2014)
Jenner, L., Rees, B., Yusupov, M. & Yusupova, G. Messenger RNA conformations in the ribosomal E site revealed by X-ray crystallography. EMBO Rep. 8, 846–850 (2007)
Rhodin, M. H. & Dinman, J. D. A flexible loop in yeast ribosomal protein L11 coordinates P-site tRNA binding. Nucleic Acids Res. 38, 8377–8389 (2010)
Yamamoto, H. et al. Structure of the mammalian 80S initiation complex with initiation factor 5B on HCV-IRES RNA. Nature Struct. Mol. Biol. 21, 721–727 (2014)
Hinnebusch, A. G. The scanning mechanism of eukaryotic translation initiation. Annu. Rev. Biochem. 83, 779–812 (2014)
Pestova, T. V. et al. Molecular mechanisms of translation initiation in eukaryotes. Proc. Natl Acad. Sci. USA 98, 7029–7036 (2001)
Demeshkina, N., Jenner, L., Westhof, E., Yusupov, M. & Yusupova, G. A new understanding of the decoding principle on the ribosome. Nature 484, 256–259 (2012)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
van Heel, M. & Schatz, M. Fourier shell correlation threshold criteria. J. Struct. Biol. 151, 250–262 (2005)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nature Methods 11, 63–65 (2014)
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Ban, N. et al. A new system for naming ribosomal proteins. Curr. Opin. Struct. Biol. 24, 165–169 (2014)
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012)
Selmer, M. et al. Structure of the 70S ribosome complexed with mRNA and tRNA. Science 313, 1935–1942 (2006)
DeLano, W. L. The PyMOL Molecular Graphic System. (DeLano Scientiic, 2002).
Acknowledgements
We thank A. Urzhumtsev and P. Afonine for discussions on FEM maps; R. Fritz and J. Michalon for IT support; R. Drui for constant high-quality engineers support; J.-F. Ménétret for technical support; H. Stark for discussions on Cs-correction; D. Moras and J.-C. Thierry for continuous encouragement in pushing structural biology frontiers; the IGBMC cell culture facilities for HeLa cell production; and the High Performance Computing Center of the University of Strasbourg funded by the Equipex Equip@Meso project. This long-term project (since 2003) was supported by the CNRS and the European Research Council (ERC Starting Grant N_243296 TRANSLATIONMACHINERY), and the electron microscope facility was supported by the Alsace Region, the FRM, the IBiSA platform program, INSERM, CNRS and the Association pour la Recherche sur le Cancer (ARC) and by the French Infrastructure for Integrated Structural Biology (FRISBI) ANR-10-INSB-05-01, and Instruct as part of the European Strategy Forum on Research Infrastructures (ESFRI).
Author information
Authors and Affiliations
Contributions
H.K. conducted purification, optimization of samples for cryo-EM and cryo-EM data processing. A.G.M. performed cryo-EM data acquisition, image processing, structure refinement and model building. S.K.N. performed structure refinement and model building. B.P.K supervised the study. All authors analysed the data. B.P.K and H.K. wrote the manuscript, with input from A.G.M. and S.K.N.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Resolution estimation.
The average resolution of the cryo-EM map as estimated from Fourier shell correlation according to the 0.143 criterion.
Extended Data Figure 2 Features of the refined cryo-EM 3D reconstruction.
Final cryo-EM map of the peptidyl-transferase centre region illustrating the high level of details observed.
Extended Data Figure 3 Typical electron density map regions of the human ribosome structure.
These final electron density maps were obtained by combining experimental amplitudes derived from the cryo-EM map and phases calculated from the iteratively refined atomic model, as done in standard refinement procedures in X-ray crystallography (see main text).
Extended Data Figure 4 Comparison of maps determined by cryo-EM and X-ray crystallography.
Top and middle, cryo-EM map and Phenix map of the human 80S ribosome (this study, A4546 region); bottom, crystal structure of a 70S ribosome at 2.8 Å resolution59 (corresponding A2600 region).
Extended Data Figure 5 Ribosomal protein structures.
a, b, Structure of ribosomal proteins uS2 and eS17.
Extended Data Figure 6 The decoding centre on the 40S.
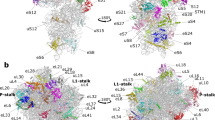
a, Contact regions of the E-site tRNA on the 40S subunit. b, Catalytic PTC region of the ribosome, highlighting the partial structural disorder of a region of protein uL16 (loop), and disorder of residue U4548 (28S rRNA) in the absence of P-site tRNA, while all other residues are well-ordered. This suggests that the P-site pocket is largely pre-defined while U4548 and the loop participate in tRNA accommodation.
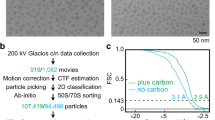
Extended Data Figure 7 Particle sorting scheme.
Particle sorting was done by 3D classification using six classes starting from 75,000 particles, resulting in two dominant classes with 10,000 and 45,000 particles, in rotated and non-rotated ribosome conformations respectively; the highest-resolution close-to-focus data set was refined using movie processing (see Methods).
Rights and permissions
About this article
Cite this article
Khatter, H., Myasnikov, A., Natchiar, S. et al. Structure of the human 80S ribosome. Nature 520, 640–645 (2015). https://doi.org/10.1038/nature14427
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14427
This article is cited by
-
A guideline for the distance measurement plans of site-directed spin labels for structural prediction of nucleic acids
Journal of Molecular Modeling (2024)
-
RPL21 interacts with LAMP3 to promote colorectal cancer invasion and metastasis by regulating focal adhesion formation
Cellular & Molecular Biology Letters (2023)
-
The La-related protein PsLARP4_5 is crucial for zoospore production and pathogenicity in Phytophthora sojae
Phytopathology Research (2023)
-
Ratcheting synthesis
Nature Reviews Chemistry (2023)
-
Selecting optimal support grids for super-resolution cryogenic correlated light and electron microscopy
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.