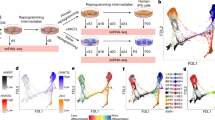
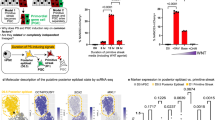
Abstract
In the context of most induced pluripotent stem (iPS) cell reprogramming methods, heterogeneous populations of non-productive and staggered productive intermediates arise at different reprogramming time points1,2,3,4,5,6,7,8,9,10,11. Despite recent reports claiming substantially increased reprogramming efficiencies using genetically modified donor cells12,13, prospectively isolating distinct reprogramming intermediates remains an important goal to decipher reprogramming mechanisms. Previous attempts to identify surface markers of intermediate cell populations were based on the assumption that, during reprogramming, cells progressively lose donor cell identity and gradually acquire iPS cell properties1,2,7,8,10. Here we report that iPS cell and epithelial markers, such as SSEA1 and EpCAM, respectively, are not predictive of reprogramming during early phases. Instead, in a systematic functional surface marker screen, we find that early reprogramming-prone cells express a unique set of surface markers, including CD73, CD49d and CD200, that are absent in both fibroblasts and iPS cells. Single-cell mass cytometry and prospective isolation show that these distinct intermediates are transient and bridge the gap between donor cell silencing and pluripotency marker acquisition during the early, presumably stochastic, reprogramming phase2. Expression profiling reveals early upregulation of the transcriptional regulators Nr0b1 and Etv5 in this reprogramming state, preceding activation of key pluripotency regulators such as Rex1 (also known as Zfp42), Dppa2, Nanog and Sox2. Both factors are required for the generation of the early intermediate state and fully reprogrammed iPS cells, and thus represent some of the earliest known regulators of iPS cell induction. Our study deconvolutes the first steps in a hierarchical series of events that lead to pluripotency acquisition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
20 May 2015
The labels in the key in Fig. 3g were corrected.
References
Brambrink, T. et al. Sequential expression of pluripotency markers during direct reprogramming of mouse somatic cells. Cell Stem Cell 2, 151–159 (2008)
Buganim, Y. et al. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 150, 1209–1222 (2012)
Golipour, A. et al. A late transition in somatic cell reprogramming requires regulators distinct from the pluripotency network. Cell Stem Cell 11, 769–782 (2012)
Chen, J. et al. H3K9 methylation is a barrier during somatic cell reprogramming into iPSCs. Nature Genet. 45, 34–42 (2013)
Hanna, J. et al. Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462, 595–601 (2009)
Ho, R., Papp, B., Hoffman, J. A., Merrill, B. J. & Plath, K. Stage-specific regulation of reprogramming to induced pluripotent stem cells by Wnt signaling and T cell factor proteins. Cell Rep. 3, 2113–2126 (2013)
O'Malley, J. et al. High-resolution analysis with novel cell-surface markers identifies routes to iPS cells. Nature 499, 88–91 (2013)
Polo, J. M. et al. A molecular roadmap of reprogramming somatic cells into iPS cells. Cell 151, 1617–1632 (2012)
Sridharan, R. et al. Role of the murine reprogramming factors in the induction of pluripotency. Cell 136, 364–377 (2009)
Stadtfeld, M., Maherali, N., Breault, D. T. & Hochedlinger, K. Defining molecular cornerstones during fibroblast to iPS cell reprogramming in mouse. Cell Stem Cell 2, 230–240 (2008)
Tanabe, K., Nakamura, M., Narita, M., Takahashi, K. & Yamanaka, S. Maturation, not initiation, is the major roadblock during reprogramming toward pluripotency from human fibroblasts. Proc. Natl Acad. Sci. USA 110, 12172–12179 (2013)
Rais, Y. et al. Deterministic direct reprogramming of somatic cells to pluripotency. Nature 502, 65–70 (2013)
Di Stefano, B. et al. C/EBPα poises B cells for rapid reprogramming into induced pluripotent stem cells. Nature 506, 235–239 (2014)
Hou, P. et al. Pluripotent stem cells induced from mouse somatic cells by small-molecule compounds. Science 341, 651–654 (2013)
Zunder, E. R., Lujan, E., Goltsev, Y., Wernig, M. & Nolan, G. P. A continuous molecular roadmap to iPSC reprogramming through progression analysis of single cell mass cytometry. Cell Stem Cell 16, 323–337 (2015)
Ichida, J. K. et al. A small-molecule inhibitor of Tgf-β signaling replaces Sox2 in reprogramming by inducing Nanog. Cell Stem Cell 5, 491–503 (2009)
Li, R. et al. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 7, 51–63 (2010)
Maherali, N. & Hochedlinger, K. Tgfβ signal inhibition cooperates in the induction of iPSCs and replaces Sox2 and cMyc. Curr. Biol. 19, 1718–1723 (2009)
Mikkelsen, T. S. et al. Dissecting direct reprogramming through integrative genomic analysis. Nature 454, 49–55 (2008)
Samavarchi-Tehrani, P. et al. Functional genomics reveals a BMP-driven mesenchymal-to-epithelial transition in the initiation of somatic cell reprogramming. Cell Stem Cell 7, 64–77 (2010)
Meissner, A., Wernig, M. & Jaenisch, R. Direct reprogramming of genetically unmodified fibroblasts into pluripotent stem cells. Nature Biotechnol. 25, 1177–1181 (2007)
Bandura, D. R. et al. Mass cytometry: technique for real time single cell multitarget immunoassay based on inductively coupled plasma time-of-flight mass spectrometry. Anal. Chem. 81, 6813–6822 (2009)
Qiu, P. et al. Extracting a cellular hierarchy from high-dimensional cytometry data with SPADE. Nature Biotechnol. 29, 886–891 (2011)
Zhang, J. et al. Dax1 and Nanog act in parallel to stabilize mouse embryonic stem cells and induced pluripotency. Nature Commun. 5, 5042 (2014)
Takahashi, K. et al. Induction of pluripotency in human somatic cells via a transient state resembling primitive streak-like mesendoderm. Nature Commun. 5, 3678 (2014)
Vierbuchen, T. et al. Direct conversion of fibroblasts to functional neurons by defined factors. Nature 463, 1035–1041 (2010)
Ellis, P. et al. SOX2, a persistent marker for multipotential neural stem cells derived from embryonic stem cells, the embryo or the adult. Dev. Neurosci. 26, 148–165 (2004)
Rubinson, D. A. et al. A lentivirus-based system to functionally silence genes in primary mammalian cells, stem cells and transgenic mice by RNA interference. Nature Genet. 33, 401–406 (2003)
Fienberg, H. G., Simonds, E. F., Fantl, W. J., Nolan, G. P. & Bodenmiller, B. A platinum-based covalent viability reagent for single-cell mass cytometry. Cytometry 81, 467–475 (2012)
Finck, R. et al. Normalization of mass cytometry data with bead standards. Cytometry 83, 483–494 (2013)
Linderman, M. D. et al. CytoSPADE: high-performance analysis and visualization of high-dimensional cytometry data. Bioinformatics 28, 2400–2401 (2012)
Acknowledgements
We thank P. Lovelace, R. Finck, K. M. Loh, K. Tanabe and S. Marro for advice. We thank J. Hanna for the secondary Mbd3fl/− MEFs. We also thank S. Knöbel for the mEF-SK4 antibody. This work was supported by the California Institute of Regenerative Medicine grant RB2-01592 (to G.P.N.), the Institute for Stem Cell Biology and Regenerative Medicine at Stanford, and a New York Stem Cell Foundation-Robertson Investigator Award. E.L. was supported by the California Institute for Regenerative Medicine Predoctoral Fellowship TG2-01159 and National Science Foundation Graduate Research Fellowship DGE-114747. E.R.Z. was supported by National Institutes of Health National Research Service Award F32 GM093508-01. M.W. is a New York Stem Cell Foundation-Robertson Investigator and a Tashia and John Morgridge Faculty Scholar at the Child Health Research Institute at Stanford.
Author information
Authors and Affiliations
Contributions
E.L., E.R.Z., G.P.N. and M.W. designed research. E.L. conducted reprogramming and sorting experiments. E.R.Z. conducted mass cytometry analysis and data processing. Y.H.N. and I.N.G. assisted with sample processing. E.L., E.R.Z., G.P.N. and M.W. analysed data. E.L., E.R.Z., G.P.N. and M.W. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Results from surface marker screen.
a, Shown are surface markers detected in MEFs, partially reprogrammed cells (PR) or ESCs analysed by flow cytometry. Numbers indicate the percentage of each population positive for the marker of interest, relative to isotype control samples. Markers are grouped for enrichment in single populations or shared between multiple populations. b, SPADE analysis for MEFs, mESCs and PRCs for surface markers analysed by mass cytometry (continued from Fig. 1b). Colour bars (bottom) represent ArcSinh-transformed counts for each marker.
Extended Data Figure 2 SPADE and biaxial analysis for MEF reprogramming.
a, SPADE analysis of lentiviral-infected MEF reprogramming populations analysed by mass cytometry (continued from Fig. 1c). Colours bars (bottom) represent ArcSinh-transformed counts for each marker. b, Biaxial plots for selected markers in control populations and during MEF reprogramming.
Extended Data Figure 3 SPADE analysis for MEF reprogramming.
a, SPADE analysis of lentiviral-infected MEF reprogramming populations analysed by mass cytometry (continued from Fig. 1c). Colours bars (bottom) represent ArcSinh-transformed counts for each marker. b, Day 12 and 16 time points for markers shown in Fig. 1c). Coloured bars for percentage total represent absolute percentages.
Extended Data Figure 4 Details for sorting experiments and chemical treatment assay.
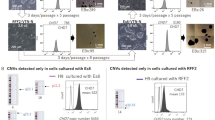
a, Ninety-six-well reprogramming assay. Twenty cells per well sorted at days 3, 6 and 9. Sox2–eGFP+ colonies were assayed on day 24. b, Gating strategy for SSEA1 in controls and day 6 reprogramming population. High- and low-expressing populations were determined on the basis of MEF and ESC control levels. c, Ten thousand SSEA1high (black bar) or SSEA1low (white bar) were sorted onto 3 cm gelatinized plates with feeders. Sox2–eGFP+ colonies were counted on day 24 (n = 3 independent experiments). d, Gating strategy for CD73 and CD49d in controls and day 6 reprogramming population. High- and low-expressing populations were determined on the basis of MEF, PRC and ESC control levels. e, f, Treatment of reprogramming populations with compounds affecting CD73 and CD49d. Shown are 96-well reprogramming efficiencies for infected Rosa-rtTA Sox2–eGFP MEFs (e) or secondary Mbd3fl/− MEFs (f). The y axis displays wells with Sox2–eGFP+ colonies 24 days after infection (e) or wells with Oct4–eGFP+ colonies 8 days after transgene induction (f) and treated with the indicated compounds for the days (D) indicated (n = 2 independent experiments).
Extended Data Figure 5 Day 6 continuation analysis on day 16.
a, Schematic of continuation analysis. Reprogramming populations were sorted for poised (CD73high or CD49high) and non-poised (CD73low) populations on day 6, cultured for 10 days on a 3 cm plate and analysed by mass cytometry on day 16. b, Morphology of CD49dhigh, CD73high or CD73low cells sorted on day 6 and inspected on day 10. Poised CD49dhigh or CD73high cells form compact colonies within several days of sorting while non-poised CD73low cells fail to do so. c, SPADE analysis (day 16) of cells sorted at day 6 for CD73high/CD49dhigh CD73high, CD49high and CD73low expression (continued from Fig. 3h). Boxes highlight a SSEA1high CD326high branch that is unique to the poised populations. Colours bars (bottom) ArcSinh-transformed counts for each marker.
Extended Data Figure 6 Continuation analysis replicates confirm a SSEA1high CD326high branch that is unique to poised populations.
Continuation analysis replicates for reprogramming-prone (CD73high/CD49dhigh, CD73high, CD49dhigh) and non-prone (CD73low) populations. Boxes highlight a SSEA1high CD326high branch that is unique to the poised populations. Colours bars (bottom) represent absolute percentages (left panel) and ArcSinh-transformed counts for each marker.
Extended Data Figure 7 Molecular characterization of reprogramming-prone intermediates.
a, Genes differentially expressed between reprogramming-prone (day 6 or day 9 CD73high or CD49dhigh) and non-prone (CD73low) populations. Genes with more than twofold differential expression between reprogramming-prone and non-prone were selected and k-means clustered (k = 5) with control and total reprogramming population expression values. b, Heat map of pluripotency-associated genes shown in Fig. 4a (log2). c, d, Quantitative PCR verification of Etv5 (c) and Nr0b1 (d) expression levels (n = 3 technical replicates). e, f, Etv5 (e) and Nr0b1 (f) knockdown qPCRs (n = 3 technical replicates). g, Representative FACS plots for day 9 CD73high/CD49dhigh quantification shown in Fig. 4b. h, Demonstration of ESC self-renewal after infection with Etv5 and Nr0b1 hairpins All infected ESCs continue to express Oct4 after passaging except ESCs infected with shEtv5–8 (n = 1).
Extended Data Figure 8 Characterization of high-efficiency reprogramming systems.
a, b, Expression analysis for CD49d, CD73 and CD104 for previously reported highly efficient reprogramming systems generated by transient expression of C/EBPα13 (a) or Mbd3 depletion12 (b). c, Oct4-GFP transgene reporter signal and d, SSEA1 and CD326 levels for the Mbd3fl/− secondary reprogramming MEFs for untreated (left) and 9 days after induction (right). e, SPADE analysis for reprogramming Mbd3fl/− secondary MEFs at days 0, 3, 6, 9 and 12 using all surface markers by mass cytometry. Percentage totals of cells and representative markers are shown for each time point. Remaining markers are shown in Extended Data Figs 9 and 10. Colours bars represent absolute percentages (left) and ArcSinh-transformed counts for each marker. f, Verification of Mbd3 loss in passage 3 Rosa26-CreER, Mbd3fl/− secondary MEFs after treatment with 4OH-tamoxifen. Mbd3 levels were compared with passage 3 Rosa-rtTA± MEFs. While there are several unspecific bands, there is clearly one band around the expected size of Mbd3 absent in 4OH-tamoxifen-treated cells (arrow).
Extended Data Figure 9 SPADE and biaxial analysis for secondary Mbd3fl/− MEFs.
a, SPADE analysis for 2° Mbd3fl/− MEF reprogramming populations (continued from Extended Data Fig. 8e). Colour bars (bottom) represent ArcSinh-transformed counts for each marker. b, Biaxial plots for selected markers.
Extended Data Figure 10 SPADE analysis for secondary Mbd3fl/− MEFs.
SPADE analysis for 2° Mbd3fl/−reprogramming populations (continued from Extended Data Fig. 8e). Colours bars (bottom) represent ArcSinh-transformed counts for each marker.
Supplementary information
Supplementary Table 1
This table contains antibody panels, sheet 1 shows the CyTOF staining panel and sheet 2 shows the FACS staining panel. (XLSX 51 kb)
Supplementary Table 2
This table contains cell counts for CyTOF experiments. (XLSX 40 kb)
Supplementary Table 3
This table contains hairpin and quantitative PCR sequences. (XLSX 34 kb)
Source data
Rights and permissions
About this article
Cite this article
Lujan, E., Zunder, E., Ng, Y. et al. Early reprogramming regulators identified by prospective isolation and mass cytometry. Nature 521, 352–356 (2015). https://doi.org/10.1038/nature14274
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14274
This article is cited by
-
Comparative roadmaps of reprogramming and oncogenic transformation identify Bcl11b and Atoh8 as broad regulators of cellular plasticity
Nature Cell Biology (2022)
-
Hybrid MoS2/g-C3N4-assisted LDI mass spectrometry for rapid detection of small molecules and polyethylene glycols and direct determination of uric acid in complicated biological samples
Microchimica Acta (2021)
-
A high throughput screening system for studying the effects of applied mechanical forces on reprogramming factor expression
Scientific Reports (2020)
-
Etv5 safeguards trophoblast stem cells differentiation from mouse EPSCs by regulating fibroblast growth factor receptor 2
Molecular Biology Reports (2020)
-
KLF4 inhibition promotes the expansion of keratinocyte precursors from adult human skin and of embryonic-stem-cell-derived keratinocytes
Nature Biomedical Engineering (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.