Abstract
Congenital heart disease (CHD) is the most prevalent birth defect, affecting nearly 1% of live births1; the incidence of CHD is up to tenfold higher in human fetuses2,3. A genetic contribution is strongly suggested by the association of CHD with chromosome abnormalities and high recurrence risk4. Here we report findings from a recessive forward genetic screen in fetal mice, showing that cilia and cilia-transduced cell signalling have important roles in the pathogenesis of CHD. The cilium is an evolutionarily conserved organelle projecting from the cell surface with essential roles in diverse cellular processes. Using echocardiography, we ultrasound scanned 87,355 chemically mutagenized C57BL/6J fetal mice and recovered 218 CHD mouse models. Whole-exome sequencing identified 91 recessive CHD mutations in 61 genes. This included 34 cilia-related genes, 16 genes involved in cilia-transduced cell signalling, and 10 genes regulating vesicular trafficking, a pathway important for ciliogenesis and cell signalling. Surprisingly, many CHD genes encoded interacting proteins, suggesting that an interactome protein network may provide a larger genomic context for CHD pathogenesis. These findings provide novel insights into the potential Mendelian genetic contribution to CHD in the fetal population, a segment of the human population not well studied. We note that the pathways identified show overlap with CHD candidate genes recovered in CHD patients5, suggesting that they may have relevance to the more complex genetics of CHD overall. These CHD mouse models and >8,000 incidental mutations have been sperm archived, creating a rich public resource for human disease modelling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Reller, M. D., Strickland, M. J., Riehle-Colarusso, T., Mahle, W. T. & Correa, A. Prevalence of congenital heart defects in metropolitan Atlanta, 1998–2005. J. Pediatr. 153, 807–813 (2008)
Hoffman, J. I. Incidence of congenital heart disease: II. Prenatal incidence. Pediatr. Cardiol. 16, 103–113 (1995)
Tennstedt, C., Chaoui, R., Korner, H. & Dietel, M. Spectrum of congenital heart defects and extracardiac malformations associated with chromosomal abnormalities: results of a seven year necropsy study. Heart 82, 34–39 (1999)
Øyen, N. et al. Recurrence of congenital heart defects in families. Circulation 120, 295–301 (2009)
Zaidi, S. et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 498, 220–223 (2013)
Liu, X. et al. Interrogating congenital heart defects with noninvasive fetal echocardiography in a mouse forward genetic screen. Circ Cardiovasc Imaging 7, 31–42 (2014)
Atkinson, D. E. & Drant, S. Diagnosis of heterotaxy syndrome by fetal echocardiography. Am. J. Cardiol. 82, 1147–1149 (1998)
Berg, C. et al. Prenatal diagnosis of cardiosplenic syndromes: a 10-year experience. Ultrasound Obstet. Gynecol. 22, 451–459 (2003)
Taketazu, M., Lougheed, J., Yoo, S. J., Lim, J. S. & Hornberger, L. K. Spectrum of cardiovascular disease, accuracy of diagnosis, and outcome in fetal heterotaxy syndrome. Am. J. Cardiol. 97, 720–724 (2006)
Tan, S. Y. et al. Heterotaxy and complex structural heart defects in a mutant mouse model of primary ciliary dyskinesia. J. Clin. Invest. 117, 3742–3752 (2007)
Leigh, M. W. et al. Clinical and genetic aspects of primary ciliary dyskinesia/Kartagener syndrome. Genet. Med. 11, 473–487 (2009)
Shiratori, H. & Hamada, H. The left–right axis in the mouse: from origin to morphology. Development 133, 2095–2104 (2006)
Damerla, R. R. et al. Ion torrent sequencing for conducting genome-wide scans for mutation mapping analysis. Mamm. Genome 25, 120–128 (2014)
Coghill, E. L. et al. A gene-driven approach to the identification of ENU mutants in the mouse. Nature Genet. 30, 255–256 (2002)
Bultman, S. et al. A Brg1 null mutation in the mouse reveals functional differences among mammalian SWI/SNF complexes. Mol. Cell 6, 1287–1295 (2000)
Kasarskis, A., Manova, K. & Anderson, K. V. A phenotype-based screen for embryonic lethal mutations in the mouse. Proc. Natl Acad. Sci. USA 95, 7485–7490 (1998)
Chen, J., Knowles, H. J., Hebert, J. L. & Hackett, B. P. Mutation of the mouse hepatocyte nuclear factor/forkhead homologue 4 gene results in an absence of cilia and random left–right asymmetry. J. Clin. Invest. 102, 1077–1082 (1998)
Sung, C. H. & Leroux, M. R. The roles of evolutionarily conserved functional modules in cilia-related trafficking. Nature Cell Biol. 15, 1387–1397 (2013)
Ware, S. M., Aygun, M. G. & Hildebrandt, F. Spectrum of clinical diseases caused by disorders of primary cilia. Proc. Am. Thorac. Soc. 8, 444–450 (2011)
Fliegauf, M., Benzing, T. & Omran, H. When cilia go bad: cilia defects and ciliopathies. Nature Rev. Mol. Cell Biol. 8, 880–893 (2007)
Clement, C. A. et al. TGF-β signaling is associated with endocytosis at the pocket region of the primary cilium. Cell Rep. 3, 1806–1814 (2013)
Sang, L. et al. Mapping the NPHP-JBTS-MKS protein network reveals ciliopathy disease genes and pathways. Cell 145, 513–528 (2011)
Stagner, E. E., Bouvrette, D. J., Cheng, J. & Bryda, E. C. The polycystic kidney disease-related proteins Bicc1 and SamCystin interact. Biochem. Biophys. Res. Commun. 383, 16–21 (2009)
Hoff, S. et al. ANKS6 is a central component of a nephronophthisis module linking NEK8 to INVS and NPHP3. Nature Genet. 45, 951–956 (2013)
Donoso, M. et al. Polarized traffic of LRP1 involves AP1B and SNX17 operating on Y-dependent sorting motifs in different pathways. Mol. Biol. Cell 20, 481–497 (2009)
Shen, Y. et al. Cardiovascular phenotyping of fetal mice by noninvasive high-frequency ultrasound facilitates recovery of ENU-induced mutations causing congenital cardiac and extracardiac defects. Physiol. Genomics 24, 23–36 (2005)
Kim, A. J. et al. Microcomputed tomography provides high accuracy congenital heart disease diagnosis in neonatal and fetal mice. Circ Cardiovasc Imaging 6, 551–559 (2013)
Pazour, G. J. et al. The intraflagellar transport protein, IFT88, is essential for vertebrate photoreceptor assembly and maintenance. J. Cell Biol. 157, 103–114 (2002)
Inglis, P. N., Boroevich, K. A. & Leroux, M. R. Piecing together a ciliome. Trends Genet. 22, 491–500 (2006)
Maere, S., Heymans, K. & Kuiper, M. BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21, 3448–3449 (2005)
Acknowledgements
We thank R. Ramirez for early assistance with the mutagenesis breeding pipeline, R. Subramanian and D. Farkas for early assistance with necropsy and pathology examination of mutants, A. Srinivasan for early assistance with exome sequencing, S. Fatakia for assistance with sequencing data maintenance, M. Wong and C. Krise for assistance with mouse curation, B. Beutler for advice on mapping mutations using intercrosses with the C57BL/10J strain and whole-mouse exome sequencing analysis, D. Weeks and Y. Shan for assistance in statistical modelling of target gene size estimates, E. Goldmuntz for helpful discussions and critical review of the manuscript, and the New England Research Institutes (NERI) for constructing the CHD Mouse Mutation Database. The project was supported by award numbers U01HL098180 (to C.W.L.) and U01HL098188 (to NERI) from the National Heart, Lung, and Blood Institute, R01MH094564 (to M.K.G.) from the National Institute of Mental Health, and HG000330 (to J.E.) from the National Human Genome Research Institute. Funding was also provided by the University of Pittsburgh School of Medicine. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Heart, Lung, and Blood Institute, the National Human Genome Research Institute or the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Study design: C.W.L. ENU mutagenesis, line cryopreservation and JAX strain datasheet construction: H.P., L.R., J.M., L.H. Mouse breeding, sample collection, sample tracking: S.A., C.L., K.L., G.C.G., A.V., C.W.L. Electronic database construction and maintenance: C.C. MGI curation: K.T., G.C.G., L.L., C.W.L., C.L.S., J.T.E. CHD phenotyping: X.L., K.L., Y.C., G.C.G., A.J.K., S.A., W.D., C.W.L., L.L., K.T., R.F. Cilia immunostain and histology: J.T.S.A., G.J.P., R.F. Analysis of airway and node cilia motility: R.F., K.L., G.C.G., A.J.K. Exome sequencing analysis: Y.L. Mutation validation: N.T.K., B.C., R.R.D., H.Y., Y.L. Mutation mapping: R.R.D., N.T.K., B.C., Y.L. Interactome analysis: M.K.G., M.T. Ciliome and pathway annotation: C.W.L., G.J.P., G.C.G., N.T.K., Y.L. Manuscript preparation: C.W.L., Y.L., N.T.K., G.C.G.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
All mutant mouse lines recovered in this mouse mutagenesis screen and their phenotype description and causative mutations are curated in the MGI database (http://www.informatics.jax.org) and can be retrieved by entering “b2b” in the search box. All mutant mouse lines curated in MGI can be reanimated from sperm cryopreserved in the Jackson Laboratory (JAXMice) Repository. All mutations recovered by mouse exome sequencing analysis are searchable together with phenotype information via the public Bench to Bassinet Congenital Heart Disease Mouse Mutation Database (http://benchtobassinet.com/ForResearchers/BasicScienceDataResourceSharing/GeneDiscoveryinMouseModels.aspx) The mouse exome datasets are available from the GNomEx Cardiovascular Development Consortium Datahub (https://b2b.hci.utah.edu/gnomex/gnomexGuestFlex.jsp?topicNumber=67).
Extended data figures and tables
Extended Data Figure 1 Breeding, phenotyping and mutation recovery pipeline for mouse forward genetic screen.
a, Two generation backcross breeding scheme used to generate G3 mutants with recessive mutations causing congenital heart defects, with all offspring from a single G1 male defined as a distinct pedigree or mutant line. b, Pipeline for recovery and curation and cryopreservation of CHD mutant mouse models and the recovery of pathogenic CHD causing mutations.
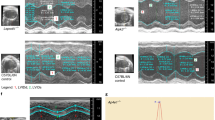
Extended Data Figure 2 Situs anomalies and congenital heart defects in Ap1b1b2b1660 mutants.
a–c, Mutants from line 1660, identified with an Ap1b1 mutation, exhibit situs solitus (a), situs inversus (b) or heterotaxy (c). Situs solitus, characterized by normal left–right visceral organ positioning, the heart apex (arrow) points to the left (levocardia), four lung lobes are on the right and one on the left, stomach is to the left, and the dominant liver lobe is on the right. With situs inversus, there is complete mirror reversal of organ situs, while with heterotaxy, visceral organ situs is randomized, such as dextrocardia with levogastria shown in c. d–g, The heterotaxy mutant in c exhibits complex CHD with atrioventricular septal defect (AVSD) (d), ventricular septal defect (VSD) (e), duplicated inferior vena cava (IVC) (f) and left pulmonary isomerism with bilateral single lung lobes (g). Ao, aorta; L1–5, lung lobes 1–5; Lv1–3, live lobes 1–3; mLA, morphologic left atrium; mLV, morphologic left ventricle; mRA, morphologic right atrium; mRV, morphologic right ventricle; PA, pulmonary artery; Stm, stomach.
Extended Data Figure 3 Distribution of pathogenic mutations recovered from the forward genetic screen.
a, Distribution of all incidental coding mutations (left), pathogenic mutations (middle) and ciliome CHD genes (right) recovered from 113 CHD mouse mutant lines. b, Recovery of pathogenic mutations and associated CHD phenotypes. Grey-filled boxes indicate CHD mutations in genes not previously identified to cause CHD. Ao, aorta; AVSD, atrioventricular septal defect; BVH, biventricular hypertrophy; DORV, double outlet right ventricle; IAA, interrupted aortic arch; MAPCA, major aortopulmonary collateral artery; PA, pulmonary artery; PTA, persistent truncus arteriosus; RAA, right aortic arch; TGA, transposition of the great arteries; VSD, ventricular septal defect; VS, vascular sling.
Extended Data Figure 4 Pathogenic splicing mutations causing CHD.
a, The 19 pathogenic splicing mutations recovered are shown, with mutations located beyond the 2-base canonical splice junction highlighted in grey. b, Schematic diagram showing the anomalous Dnah5c.133290-10T>A mutant transcript observed, with the polymerase chain reaction (PCR) primer location and anomalous PCR product size indicated. c, Sanger sequencing profile showing point mutation in Dnah5c.133290-10T>A mutant transcript versus that of wild type. d, PCR amplification of Dnah5c.133290-10T>A heterozygous mutant (m/+) showed the expected 514 bp wild-type and 271 bp mutant PCR product, while only the 271 bp mutant product was observed in the homozygous mutant (m/m) sample.
Extended Data Figure 5 Ciliome mutations causing CHD with and without laterality defects.
A flow chart showing the distribution of ciliome versus non-ciliome CHD genes among laterality versus non-laterality CHD lines, and further stratification of ciliome CHD genes affecting primary versus motile cilia function.
Extended Data Figure 6 CHD phenotypes associated with mutations affecting endocytic trafficking.
a–c, Outflow tract malalignment defects with double outlet right ventricle (DORV) and overriding aorta (Ao) were observed in Ap2b1 (a), Dnm2 (b) and Snx17 (c) mutants, with Ap2b1 (a) mutant also showing anterior positioning of the aorta (Taussig–Bing type DORV). Snx17 mutant also has AVSD. d, e, Lrp2 (d) and Lrp1 (e) mutants both exhibited outflow tract septation defect with persistent truncus arteriosus (PTA). Scale bar, 0.5 mm.
Extended Data Figure 7 CHD genes associated with axon guidance.
Diagram illustrating the biological context of several CHD genes known to be involved in axonal guidance (colour highlighting indicates CHD genes recovered from the present screen). Adapted from QIAGEN’s Ingenuity Pathway Analysis (http://www.qiagen.com/ingenuity).
Supplementary information
Supplementary Data 1
This data file contains CHD mutations and comparison to other mutant alleles and human diseases. (XLSX 1524 kb)
Supplementary Data 2
This data file contains position and sequence conservation of CHD mutations. (XLSX 19888 kb)
Supplementary Data 3
This data file contains gene ontology and pathway analyses of CHD interactome genes. (XLSX 167 kb)
Supplementary Data 4
This data file contains ciliome and pathway annotation for 28 CHD candidate genes recovered from Pediatric Cardiac Genomics Consortium human CHD patient exome sequencing analysis. (XLSX 74 kb)
Ultrasound imaging of normal fetus shown in Figure 1a.
Vevo 2100 color flow Doppler imaging in coronal view show normal alignment of the two great arteries with normal connection to the two ventricles. (MOV 1838 kb)
Ultrasound imaging of mutant fetus from line b2b2025 shows DORV and presence of a VSD
Vevo2100 color flow imaging in coronal view of fetus in Figure 1e-k showed aorta and pulmonary artery side-by side, both emerging from the right ventricle (RV) indicating DORV, and a shunting of blood between the two ventricles indicating ventricular septal defect (VSD). (MOV 2312 kb)
Ultrasound imaging of mutant fetus from line b2b2025 shows AVSD and muscular VSD
Vevo 2100 color flow imaging in transverse view of fetus in Figure 1e-k detected forward blood flow and regurgitation from a common atrioventricular valve suggesting atrioventricular septal defect. Also observed was a muscular ventricular septal defect. (MOV 2342 kb)
Ultrasound imaging of mutant fetus from line b2b2025 shows heterotaxy
Vevo2100 2D imaging in coronal view of fetus in Figure 1e-k detected heart apex pointing to left suggesting levocardia, but stomach (Stom) located on right, which together indicated this fetus has heterotaxy. (MOV 2318 kb)
Videomicroscopy of ciliary motion of tracheal epithelia of newborn Foxj1 mutant mice
Tracheal airway epithelium in a newborn homozygous Foxj1b2b774 mutant mouse shows normal ciliary motion. (MOV 3304 kb)
Videomicroscopy of ciliary motion in embryonic node of Foxj1 mutant embryo
Cilia in the embryonic node of a homozygous Foxj1b2b774 mutant embryo shows dyskinetic ciliary motion and no nodal flow. (MOV 8174 kb)
Rights and permissions
About this article
Cite this article
Li, Y., Klena, N., Gabriel, G. et al. Global genetic analysis in mice unveils central role for cilia in congenital heart disease. Nature 521, 520–524 (2015). https://doi.org/10.1038/nature14269
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14269
This article is cited by
-
Standardisation and future of preclinical echocardiography
Mammalian Genome (2023)
-
The second heart field: the first 20 years
Mammalian Genome (2023)
-
De novo disruptive heterozygous MMP21 variants are potential predisposing genetic risk factors in Chinese Han heterotaxy children
Human Genomics (2022)
-
A change of heart: new roles for cilia in cardiac development and disease
Nature Reviews Cardiology (2022)
-
Flow blockage disrupts cilia-driven fluid transport in the epileptic brain
Acta Neuropathologica (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.