Abstract
Pluripotent stem cells provide a powerful system to dissect the underlying molecular dynamics that regulate cell fate changes during mammalian development. Here we report the integrative analysis of genome-wide binding data for 38 transcription factors with extensive epigenome and transcriptional data across the differentiation of human embryonic stem cells to the three germ layers. We describe core regulatory dynamics and show the lineage-specific behaviour of selected factors. In addition to the orchestrated remodelling of the chromatin landscape, we find that the binding of several transcription factors is strongly associated with specific loss of DNA methylation in one germ layer, and in many cases a reciprocal gain in the other layers. Taken together, our work shows context-dependent rewiring of transcription factor binding, downstream signalling effectors, and the epigenome during human embryonic stem cell differentiation.
Similar content being viewed by others
Main

Human embryonic stem (ES) cells hold great promise for tissue engineering and disease modelling; yet a key challenge to deriving mature, functional cell types is understanding the molecular mechanisms that underlie cellular differentiation. There has been much progress in understanding how core regulators such as OCT4 (also known as POU5F1), SOX2, and NANOG as well as transcriptional effector proteins of signalling pathways, such as SMAD1, TCF3, and SMAD2/3, control the molecular circuitry that maintains human ES cells in a pluripotent state1,2. While the genomic binding sites of many of these factors have also been mapped in mouse ES cells, cross-species comparison of OCT4 and NANOG targets showed that only 5% of regions are conserved and occupied across species3. Together with more general assessment of divergent transcription factor (TF) binding4, those results highlight the importance of obtaining binding data in the respective species.
It is well understood that epigenetic modifications, such as DNA methylation and posttranslational modifications of the various histone tails, are essential for normal development5,6. TF binding sites are overlapping with regions of dynamic changes in DNA methylation and are linked to its targeted regulation7,8. More generally, TFs orchestrate the overall remodelling of the epigenome, including the priming of loci that will change expression only at later stages6,9,10. It has also been shown that lineage-specific TFs and signalling pathways collaborate with the core regulators of pluripotency to exit the ES cell state and activate the transcriptional networks governing cellular specification11,12. However, how the handoff between the central regulators occurs and what role individual TFs and signalling cues play in rewiring the epigenome to control proper lineage specification and stabilize commitment is still poorly understood.
TF binding maps across human ES cell differentiation
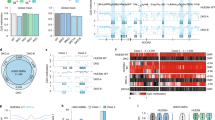
To dissect the dynamic rewiring of TF circuits, we used human ES cells to derive early stages of endoderm (dEN), mesoderm (dME) and ectoderm (dEC)13,14,15 along with a mesendoderm (dMS) intermediate (Fig. 1a, Supplementary Information). We defined and collected the dMS population at 12 h owing to maximal expression of BRACHYURY (T) (Fig. 1b), and carried out chromatin immunoprecipitation followed by sequencing (ChIP-seq) for four of the Roadmap Epigenomics Project16 core histone modifications (H3K4me1, H3K4me3, H3K27Ac and H3K27me) as well as RNA sequencing (RNA-seq) of polyadenylated transcripts (Supplementary Table 1). As expected, we observe upregulation of key TFs including FOXA2 and HNF4A in dEN, HAND1 and SNAI2 in dME, and OTX2 and PAX6 in dEC (Fig. 1b,c)9,17. We identified high-quality antibodies for 38 factors (Fig. 1c) and provide detailed information, including their validation and use in other studies, in Supplementary Table 2.
a, Schematic of the human ES cell differentiation system including timeline and key signalling pathways that are modulated. b, Normalized RNA expression of selected TFs over the differentiation timeline towards endoderm. c, RNA-seq data of the selected TFs. Factors are generally ordered by condition where they are most active: ES cells on top, followed by dMS, dEN, dME, and dEC.
Using a micrococcal nuclease (MNase)-based ChIP-seq (MNChIP-seq) protocol18 we obtained binding patterns as well as reproducibility comparable to sonication ChIP-seq with only 1–2 million cells (Extended Data Fig. 1a–e). We quantified the enrichment over background for each experiment (Supplementary Table 3), and show that the level of binding is comparable to TF ChIP-seq data from ENCODE19 (Extended Data Fig. 1f). To evaluate computationally the specificity of the chosen antibodies we searched our binding maps for previously reported motifs of the respective factors20 (Extended Data Fig. 2). Our final data set consists of 6.7 billion aligned sequencing reads that yield 4.2 million total binding events (Supplementary Table 3). The binding spectrum of all TFs averages 21,468 peaks and ranges from 578 to 100,778 binding events. Of these 23% are found in promoters, 44% in distal regions, 30% in introns, and 3% in exons.
Classes of TF dynamics
We first grouped the TF binding dynamics into four main classes (static, dynamic, enhanced and suppressed) similar to prior studies in yeast21 and then further subdivided each of these as either temporal (between successive time-points) or cross-lineage (between germ layers) (Fig. 2a and Extended Data Figs 3 and 4).
a, Classes of dynamics comparing TF binding between successive time points (temporal) or between different germ layers (cross-lineage). The schematics, browser images, and Venn diagrams illustrate examples of each class. b, SMAD4 predominantly binds to unique regions in the three germ layers. c, EOMES binding is enhanced from ES cells to dMS and dynamic in dEN. d, OTX2 binding is dynamic in dEN and dEC when compared to ES cells.
A number of factors, including NANOG, show largely static binding in ES cells and endoderm (Fig. 2a). This could be the result of NANOG’s proposed functions in endoderm, including protection against neuroectoderm specification and buffering TGF-β signalling to avoid premature induction of definitive endoderm11. CTCF is both temporally and cross-lineage static in its binding pattern, showing a similar overlap between cell types as between replicates (Extended Data Figs 1a and 4a). The high similarity in binding is consistent with a previous study that investigated CTCF binding in 19 diverse human cell types22. Although each of the germ layer derivatives exhibits unique expression signatures, they show overall only limited transcriptional dynamics9, which is in agreement with the largely static enrichment for POLII and cMYC (Extended Data Fig. 3a).
In contrast, a number of the selected factors show dynamic binding between two (for example, GATA4) or more (for example, SMAD4) cell types (Fig. 2a, b). EOMES changes its binding profile notably during the dMS to dEN transition, suggesting its function may evolve at different stages of differentiation (Fig. 2c). Also, OTX2 occupies a largely different binding spectrum in the undifferentiated cells compared to dEN and dEC (Fig. 2d). Many factors also exhibit different temporal and cross-lineage dynamics. For example, while NANOG binding is temporally static in dMS and dEN, it is suppressed temporally and cross-lineage in dME (Extended Data Figs 3a, 4b). Meanwhile, OCT4 and SOX2 binding is temporally static in dEN, but cross-lineage dynamic between dEN and dME (Extended Data Figs 3a and 4c). Likewise, TCF4 (a transcriptional effector of WNT signalling) is temporally static in dEN but suppressed in dME and dEC, consistent with the lack of WNT signalling in those germ layers13,14,15 (Extended Data Figs 3a and 4d). Finally, OTX2 is temporally suppressed in dME (Fig. 2a), but temporally dynamic in the other germ layers (Fig. 2d).
To investigate the interplay between TFs across the cell types and how they might collaborate to mediate cellular transitions, we analysed all pairwise TF co-binding relationships. We identify several germ-layer-specific co-binding interactions; for example, GATA4 targets associate significantly (hypergeometric P < 10−300) with SMAD1 binding in dME, but less so in dEN (Fig. 3a, left and Extended Data Fig. 5). To extend this, we clustered all co-binding relationships and identified groups of interactions between factors and developmental time points (Fig. 3a, right). We found both clusters of many regulators in one cell type as well as clusters for individual TFs across cell types. For instance, cluster C1 shows that CTCF binding spectrum is highly similar in all three germ layers. In cluster C2, we find high overlap in binding between key endoderm regulators, while C4 captures primarily pluripotent and dMS binding profiles. Many known mesoderm factors aggregate in clusters C8 and flanking the pluripotent cluster C4 are EOMES, T, and NR5A2 clusters (C3, C5, C6), all known regulators in mesendoderm that are likely to be involved in the transition towards mesoderm and endoderm11.
a, Left, overlap in binding between GATA4 and SMAD1 is greater in dME than in dEN. Similarly, overlap in binding between OTX2 and SMAD1 is greater in dEN than in dEC. Right, highly significant TF co-binding relationships are assigned a dark blue colour, representing –log10 of hypergeometric P value. All TF dynamics and co-binding interactions are clustered and displayed in a matrix, where each row/column represents a single ChIP-seq experiment. The colour code indicates the cell type identity for the majority of ChIP-seq profiles making up each cluster. b, Genomic annotations for factors that bind more than 15,000 regions in multiple conditions. c, GATA4 (top) and OTX2 (bottom) binding is associated with different chromatin marks between lineages.
Interestingly, we noticed that GATA4 and OTX2 binding in the different cell types is not only divergent, but enriched at distinct genomic features (Fig. 3b). In dME 36% of all GATA4 binding sites occur in promoters, compared to only 13.6% in dEN. OTX2’s fraction of binding sites at promoters is larger in dEN (34%) and dEC (28%) than in ES cells (13%). Accompanying GATA4’s shift in binding preference, we also observe higher levels of H3K4me1 at dEN targets and higher H3K27Ac and H3K4me3 enrichment in dME (Fig. 3c). Similarly, OTX2 associates with higher H3K27Ac and H3K4me1 levels in ES cells, and higher H3K4me3 occupancy in dEN and dEC, in line with increased promoter binding in these two germ layers (Fig. 3c). It is worth noting that similar to the distinct GATA4/SMAD1 co-binding, OTX2 co-occupies a higher fraction of loci with SMAD1 in dEN than in dEC (Fig. 3a, left and Extended Data Fig. 5). Although TGF-β signalling is primarily associated with effector proteins SMAD2/3, it also acts through the SMAD1/5/8 complex and may encourage interaction with OTX2 in dEN but not in dEC, where TGF-β signalling is specifically inhibited23.
H3K27Ac domains identify lineage regulators
Extended H3K27Ac domains have recently been termed super-enhancers and were used to describe regulatory regions that enrich for binding sites of master TFs in the respective cell types24,25. Binding of GATA4 in dME indeed coincides with long stretches of H3K27Ac near several mesodermal genes (Fig. 4a). We therefore used the previously described approach24,25 to rank extended H3K27Ac domains in our populations and identify such super-enhancers (Supplementary Table 4), which were indeed predominantly unique to each cell type (Fig. 4b and Extended Data Fig. 6). As expected, in human ES cells, core regulators OCT4, SOX2, NANOG (abbreviated OSN), and OTX2 binding is highly enriched at super-enhancers1,26 (Fig. 4c).
a, Browser tracks for H3K27me3 and H3K27Ac across all five cell types as well as GATA4/SMAD1 enrichment over the HAND1 locus in dME. b, Limited overlap of extended H3K27Ac domains between cell types. c, Top, schematic of different transitions in TF regulation at super-enhancers. OTX2 is present at the dMS stage, but not shown in the schematic due to limited space. Bottom, hypergeometric P values (–log10) displaying the most significant overlaps in H3K27Ac super-enhancers (SE) and TF binding for each cell type.
We used enrichment of binding at super-enhancers for identifying possible master regulators in the germ layers (Fig. 4c); the results were highly robust to different cut-offs for defining the super-enhancers (Supplementary Table 5). Surprisingly, we found that many of the core regulators bound at ES cell super-enhancers also occupy dEN super-enhancers, including OSN, OTX2, SMAD1, TCF4, and SMAD2/3 (Fig. 4c and Extended Data Fig. 6e). In mesoderm, GATA4 and SMAD1 were the most highly enriched factors at dME super-enhancers (Extended Data Figs 6f and 7), consistent with GATA4’s known role in directing cardiomyocyte development downstream of BMP signalling27. OTX2 is known to regulate neuronal subtype specification in the midbrain28 and we found strong enrichment for OTX2 binding at ectoderm super-enhancers (Fig. 4c and Extended Data Fig. 6g, h). Meanwhile, dMS super-enhancers were enriched for known regulators such as EOMES and T, along with OSN and OTX2 (Fig. 4c). At a lower significance level we also find enrichment for a number of endoderm factors, including FOXA1/2, GATA4/6 and SOX17 (Supplementary Table 5). Interestingly, binding of EOMES, T and FOXA1/2 in the undifferentiated ES cells was also enriched (hypergeometric P < 10−6) at dMS super-enhancers (Fig. 4c and Extended Data Fig. 6), suggesting that a number of loci might be already marked before differentiation.
Regulation of poised enhancers across germ layers
As dEN H3K27Ac domains were mostly devoid of known endoderm TFs, we asked if such regulators are instead present at regions that enrich for H3K4me1, as seen at the HNF1B locus (Fig. 5a). H3K4me1 can be found at both active and poised enhancers29 and is known to also form extended enhancer domains that may not overlap with the H3K27Ac domains24,25. Using the same approach as above we identified extended H3K4me1 domains in dEN and then measured enrichment for TF binding in these regions. In contrast to H3K27Ac, the top H3K4me1 domains were enriched for binding of FOXA1/2, GATA4, GATA6, and SOX17 (Extended Data Fig. 8a, b), known regulators of the early endodermal fate30. We then measured the significance in overlap between TF binding and all poised enhancers for each cell type and found strong enrichment for these regulators and PRDM1 in dEN (Extended Data Fig. 8c, d).
a, Selected browser tracks for H3K27Ac and H3K4me1 and normalized binding of FOXA1/2, SOX17, and GATA4/6 over the HNF1b locus. Grey vertical bars highlight regions enriched for H3K4me1 in dEN. b, P values (–log10) for three or more of the most enriched DNA binding motifs (rows) at SMAD1 binding per cell type (columns). c, Gene expression of selected lineage markers in dME (top) and dEN (bottom), comparing three GATA4 KD and control lines, determined by reverse transcription quantitative PCR (RT–qPCR). The mean expression for 22 dEN and 24 dME marker genes (excluding GATA4 and SMAD1) is shown as the last bar in each panel. Error bars display the standard deviation in fold expression change. Asterisk highlights genes with significant (P < 0.05, paired t-test) change in expression between control and KD replicates. d, Normalized SMAD1 (left) and H3K27Ac (right) occupancy decreases in shRNA KD versus control lines in dME but not in dEN.
In concordance with this analysis and global chromatin remodelling trends (Extended Data Fig. 8e), GATA4 is associated with dynamics of H3K4me1 in dEN and H3K27Ac in dME. Given that the SMAD proteins are known to interact with histone acetyltransferases EP300 and CBP31, it is plausible that, through BMP signalling in dME, GATA4 interacts with SMAD1 and recruits EP300 to induce acetylation of H3K27 at target sites. This recruitment relationship is further supported by the higher enrichment of GATA4 motif instances at SMAD1 binding sites in dME versus dEN (Fig. 5b and Extended Data Fig. 8f) and the stronger enrichment of H3K27Ac at GATA4 targets in dME versus dEN (Fig. 3c).
To further explore this, we used several shRNAs to knock down (KD) GATA4 and then measured gene expression following differentiation into dME and dEN (Extended Data Fig. 9a). The mean expression for more than 20 lineage markers is very similar between control and KD cell lines, arguing that the KD cells still differentiate into comparable populations (Fig. 5c, right bar). While the GATA4 KD in dEN does not greatly affect any of the measured endoderm TFs (total P = 0.49, paired t-test), in dME the KD leads to a 1.7–4-fold reduction in the expression of seven key factors (total P = 5.39−5, paired t-test). GATA4 binding in dME and dEN occupies similar loci in control and KD cell lines (Extended Data Fig. 9b, c), and H3K27Ac super-enhancers in dME are largely unaffected by our knockdown (Extended Data Fig. 9d, e). Nonetheless, we observe a significant decrease in SMAD1 and H3K27Ac enrichment in dME at GATA4 target sites in the KD lines (Fig. 5d, P < 10−300, paired t-test). To a lesser degree, we also observe a decrease in mean SMAD1 occupancy at binding sites away from GATA4 (Extended Data Fig. 9f). This could be the result of the general reduction of SMAD1 expression in the dME KDs or linked to other TFs that aid SMAD1 binding, such as factors from the TEAD and GATA family (Fig. 5b).
Loss of DNA methylation at targets of lineage TFs
DNA methylation can silence genomic regions, directly or indirectly, and plays an important role during mammalian development5. Some TFs can modulate DNA methylation levels8, but it is not generally known which factors can alter it in a developmental context and which ones might be sensitive to its presence. In endoderm at a region upstream of SOX17, we observe specific loss of DNA methylation accompanied by epigenetic remodelling to a poised state. We also observe that the loss of DNA methylation associates with lineage-specific binding of several TFs (Fig. 6a and Extended Data Fig. 10a). Interestingly, OTX2 and NANOG show some enrichment already in ES cells that seems to be linked to a very focal depletion of DNA methylation that may serve as a means of initial marking or protecting the region for downstream binding (Extended Data Fig. 10b).
a, Top, browser tracks for H3K4me3 and H3K27me3 as well as enrichment of selected TFs upstream of SOX17. Bottom, each rectangle represents a single CpG and its methylation state. Loss of DNA methylation occurs specifically in dEN, which coincides with changes in chromatin state and specific binding of several known endoderm factors. b, Hypergeometric P values (–log10) for the overlap in TF binding and regions that gain or lose DNA methylation (DNAme) relative to ES cells. Possible transition states are defined at the top. Heat maps display the enrichment of TF binding in ES cells, dMS (left), dEN, dEC (centre), and dME (right) at differentially methylated regions in the three germ layers. c, Whole-genome bisulfite sequencing (WGBS)-based average CpG methylation level of 100-bp tiles over GATA6-bound dEN targets. d, WGBS mean methylation level at OTX2 dEC targets. e, WGBS mean methylation level at GATA4 dEN and dME targets. f, Reduced-representation bisulfite sequencing-based average CpG methylation level of 100-bp tiles over GATA4 targets in control and GATA4 KD cell lines in dEN (left) and dME (right). For comparison, WGBS ES cell mean methylation level is also shown (grey).
We next performed global enrichment analysis for all TF binding at regions that either gained or lost DNA methylation. Many target sites of OSN as well as SMAD1 and TCF4 show gain of DNA methylation in all three lineages, consistent with silencing of their pluripotency-related target genes (Fig. 6b, left). The dMS target sites of T and EOMES also become methylated in the three germ layer populations. Interestingly, we frequently find a reciprocal gain in DNA methylation in the alternative lineages of key dEN and dEC factors (Fig. 6b, middle).
As shown near SOX17, we also find that lineage regulators associate with targeted loss of DNA methylation. For instance, in dEN binding sites of EOMES, FOXA1/2 (Extended Data Fig. 10c, d), GATA4/6, SOX17, and OTX2 display focal and germ layer specific loss of DNA methylation (Fig. 6b, c). We also find strong enrichment for loss of DNA methylation at OTX2 binding sites in dEC (Fig. 6b, d). In dME we find seven partially overlapping TFs that show loss of DNA methylation at their binding sites, especially in regions that also gain H3K27Ac (Fig. 6b, e and Extended Data Fig. 7c). Using reduced-representation bisulfite sequencing32 we measured the DNA methylation level for a representative subset of targets in GATA4 KD and control lines. Both dME and dEN GATA4 KD cells displayed significantly higher methylation level (P < 10−10, paired t-test) (Fig. 6f and Extended Data Fig. 10e), suggesting a possible role for GATA4 in the focal depletion of DNA methylation.
Discussion
Directed differentiation of human ES cells into the three embryonic germ layers coupled with comprehensive TF binding analysis and integration with epigenomic data has allowed us to characterize differentiation-associated regulatory dynamics. We find that targets of many lineage-specific factors associate with loss of DNA methylation in those germ layers, while factors that are expressed in more than one lineage (GATA4, GATA6, OTX2, SOX17) show a corresponding loss of DNA methylation at their targets in multiple cell types. This is in line with the model that some TFs have an intrinsic ability to alter DNA methylation, although more work is needed to determine if all of these can indeed be considered “pioneer factors”33. We also find a specific gain of DNA methylation for the targets of many TFs at later time points or in parallel time-points but along alternate lineages. This might present a possible mechanism for occluding binding sites of certain methylation sensitive factors at past or alternate differentiation paths.
To investigate the interplay between TF binding and the chromatin landscape, we focused on TF dynamics at H3K27Ac super-enhancers, where OTX2 and OSN seem to guide the transition to dEN while GATA4 and OTX2 act as key regulators for dME and dEC, respectively. GATA4 exemplifies a factor with distinct germ layer functions, where in dEN it resides at poised enhancers and in dME it appears to associate with SMAD1/EP300 to establish and maintain H3K27Ac domains. The dual use of GATA4 and OTX2 highlights the modularity in transcriptional networks in development and the complex interaction of downstream signalling effectors, TFs and chromatin in the three germ layers.
Methods
No statistical methods were used to predetermine sample size.
Human ES cell culture
Cell culture was done as reported previously9. Briefly, we chose the NIH approved, male human embryonic stem (ES) cell line HUES64 because it has maintained a stable karyotype over many passages and is able to differentiate well into the three germ layers. HUES64 was routinely tested for Mycoplasma and was negative in all instances. ES cells were maintained on ∼15,000 cells per cm2 irradiated murine embryonic fibroblasts (MEFs, Global Stem) and cultured in 20% knockout serum replacement (KSR, Life Technologies), 200 mM GlutaMAX (Life Technologies), 1× minimal essential media (MEM) non-essential amino acids solution (Life Technologies), 10 μg ml−1 bFGF (Millipore), 55 µM β-mercaptoethanol in knockout Dulbecco’s modified Eagle medium (KO DMEM, Life Technologies). ES cells were passaged every 4–5 days using 1 mg ml−1 collagenase IV (Life Technologies).
Directed differentiation of human ES cells
When human ES cells reached 60–70% confluency on MEFs, the cells were plated as clumps on 6-well plates coated with Matrigel (Life Technologies) in mTeSR1 basal medium (Stem Cell Technologies). We maintained the cells for three days in feeder-free culture and then induced directed differentiation towards mesendoderm, endoderm, mesoderm, and ectoderm using different media conditions. For mesendoderm and endoderm differentiation cells were cultured for 12 and 120 h, respectively, in Roswell Park Memorial Institute (RPMI) medium (Life Technologies) supplemented with 100 ng ml−1 activin A (R&D Systems), 50nM ml−1 WNT3A (R&D Systems), 0.5% FBS (Hyclone), 200 mM GlutaMAX (Life Technologies), 0.2× MEM non-essential amino acids solution (Life Technologies), and 55 µM β-mercaptoethanol. For the first 24 h of mesoderm differentiation, cells were cultured in DMEM/F12 medium supplemented with 100 ng ml−1 activin A (R&D Systems), 10 ng ml−1 bFGF (Millipore), 100 ng ml−1 BMP4 (R&D Systems), 100 ng ml−1 VEGF (R&D Systems), 0.5% FBS (Hyclone), 200 mM GlutaMax (Life Technologies), 0.2× MEM non-essential amino acids solution (Life Technologies), and 55 µM β-mercaptoethanol. From 24 to 120 h of mesoderm differentiation, Activin A was removed from the culture. For ectoderm differentiation cells were cultured in DMEM/F12 medium supplemented with 2 µM TGF-β inhibitor (Tocris, A83-01), 2 µM WNT3A inhibitor (Tocris, PNU-74654), 2 μM dorsomorphin BMP inhibitor (Tocris), 15% KOSR (Life Technologies), 0.2× MEM non-essential amino acids solution (Life Technologies), and 55 µM β-mercaptoethanol. Media was changed daily. Before inducing differentiation, we manually removed the differentiated cell clumps. We routinely obtain greater than 80% differentiated cells based on the presence of the surface marker CD56 (81.7% of mesoderm and 94.4% of ectoderm cells) and greater than 70% differentiated cells based on the surface marker CD184 for endoderm.
RNA extraction and RNA-seq
For measuring expression levels, RNA was isolated from the human ES cells and differentiated cells using TRIzol (Invitrogen, 15596-026), further purified with RNeasy columns (QIAGEN, 74104) and DNase treated. RNA-seq library construction and data analysis was carried out as described previously9.
Antibodies
Supplementary Table 2 lists detailed information for all antibodies used in this study, along with references that validate the specificity and use of each antibody.
MNChIP-seq and library construction
ChIP-seq for all chromatin marks was done as in ref. 9. MNChIP-seq for TFs was carried out as in ref. 9 with several modifications, including the micrococcal nuclease (MNase) digestion. Briefly, cell were grown to a final count of 10 million, resuspended in PBS, and crosslinked in 10% formaldehyde solution for 10 min at room temperature. Following quenching with 0.125M glycine and two PBS washes, we isolated nuclei using cell lysis buffer (20 mM Tris-HCl ph8, 85 mM KCl, 0.5% NP40). Nuclei were then digested using MNase (Worthington, LS004797) as done in ref. 18. Digestion was stopped with 0.05 M EGTA and chromatin was aliquoted into 1–2 million cells per ChIP. Antibodies were added and immunoprecipitation was carried out overnight at 4 °C as done in ref. 9. The next day, protein G beads (Life Technology, 10009D) were added for 2 h at 4 °C to isolate the protein-bound DNA and washed twice using low salt wash buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.1, 150 mM NaCl), high salt wash buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-HCl pH 8.1, 500 mM NaCl), LiCl wash buffer (0.25 M LiCl, 0.5% NP40, 0.5% sodium deoxycholate, 1 mM EDTA, 10 mM Tris-HCl pH 8.1,), and TE buffer pH 8 (10 mM Tris-HCl, pH 8, 1 mM EDTA pH 8). DNA was eluted twice using 100 μl of ChIP elution buffer (1% SDS, 0.1 M NaHCO3) at 65 °C for 15 min. Crosslinking was reversed by addition of 32 µl reverse crosslinking salt mixture (250 mM Tris-HCl pH 6.5, 62.5 mM EDTA pH 8, 1.25 M NaCl, 5 mg ml−1 proteinase K) for 5–18 h at 65 °C. DNA was isolated using phenol/chloroform extraction and treated with DNase-free RNase for 30 min at 37 °C. The whole-cell extract (WCE) control was generated using MNase-treated material that was then reverse-crosslinked and phenol/chloroform-extracted, skipping the immunoprecipitation and washing steps. DNA libraries were constructed using standard Illumina protocols for blunt-ending, polyA extension, and ligation, except each clean-up step was replaced with phenol/chloroform extractions to preserve small fragments as done in ref. 18. Ligated DNA was then PCR-amplified and gel-size-selected for fragments between 30 and 600 bp. Samples were sequenced using Illumina HiSeq at a target sequencing depth of 20 million uniquely aligned reads.
shRNA infection and knockdown experiments
ES cells were maintain MEFs in KSR culture media as described above and passaged onto geltrex coated dishes in mTeSR1 culture media before infection. When cells were ∼75% confluent, cells were collected with accutase as single cells or small clumps. 100,000 ES cells were plated per well of 12-well plate coated with geltrex and in mTeSR1 culture media. After 24 h, ES cells were infected twice on separate days for 3 h with approximately 30 viral particles per cell. 48 h after the last infection, cells were selected with 1 μg ml−1 puromycin until the non-infected ES cells die off (usually within 3 days). Knockdown (KD) and control shRNA-infected ES cell lines were then maintained as described above. We then performed directed differentiation of three control and KD cell lines into 5-day dEN and dME. We collected cells and carried out RNA and DNA extraction as ref. 9. cDNA reaction was set-up from 1 μg of total RNA per sample using High-Capacity cDNA RT Kit (Life Technologies). qPCR was performed on 384-well TaqMan hPSC Scorecard plates using Viia7 RUO software and Applied Biosystems ViiA7 instrument. CT values were normalized using two probes of the ACTN housekeeping gene and averaged for the three GATA4 KD and three control cell lines to obtain fold change in expression. DNA was used for reduced-representation bisulfite sequencing as in ref. 32. We also collected crosslinked cells from the same samples and carried out MNChIP-seq for GATA4, SMAD1, and H3K27Ac as described above. Composite plots display the average normalized occupancy for three GATA4 KD and two control cell lines. We used pLKO.1 cloning vector with the following target sequences for GATA4 KD: CCAGAGATTCTGCAACACGAA, CGAGGAGATGCGTCCCATCAA, CCCGGCTTACATGGCCGACGT. The shRNA control cell lines targeted gene products not present in the human genome using the same cloning vector with the following target sequences: TGACCCTGAAGTTCATCTGCA (GFP) and CACTCGGATATTTGATATGTG (Luciferase).
Selection of transcription factors
Approximately half of the transcription factors (TFs) were chosen because they are known to play an important role in regulation of pluripotent cells or in the transition to mesendoderm (for example, BRACHYURY), endoderm (for example, SOX17), mesoderm (for example, GATA4), and ectoderm (for example, PAX6). Others were chosen computationally based on Nanostring expression analysis and RNA-seq data. Previous work12 identified that OCT4 and SOX2 play distinct roles in the transition from ES cells to mesendoderm and ectoderm based on differential expression of these TFs in the two lineages. We used a similar approach to computationally identify factors that are differentially expressed in mesoderm and endoderm. Another study showed that temporal upregulation of TFs can be indicative of their importance at specific stages of blood differentiation34. We used this approach to identify factors that were upregulated upon transition to mesendoderm, mesoderm and endoderm and included those as well in the study (see Supplementary Table 2 for additional details on the factors).
ChIP-seq and MNChIP-seq data processing
Reads were aligned to the hg19 reference assembly using bwa version 0.5.7 (ref. 35) with default parameter settings. Subsequently, reads were filtered for duplicates and extended by 200 bp. For visualization, extended reads were summed at each base and normalized for sequencing depth by scaling the y axis to represent cumulative reads per 1 million reads sequenced. This normalization was used for browser and heat map visualizations of the data in all figures. We used MACS36 peak calling algorithm with default settings to identify significant binding events for each TF, excluding duplicate reads. Peaks were additionally discarded if they overlapped with regions that MACS detected as peaks in four different WCE samples. Such regions have been shown to cause false-positive peaks in ChIP-seq data due to unannotated high copy number regions37. Peaks were then annotated according to their proximity to transcription start sites (TSSs) using Homer38. Peaks within exons and introns were annotated first. Then, peaks overlapping a region from −2,000 bp to +500 bp of their nearest TSS were annotated as at promoters. Peaks outside of promoters but not in exons or introns were annotated as distal.
Data quality assessment and motif analysis
To quantify enrichment over background in ChIP-seq experiments, we measured the percentage of reads in peaks by counting all unique tags within 1,000 bp regions centred on all binding events, using bedtools multicov function with default parameters. To compare to ENCODE, we downloaded all (n = 1,410) TF ChIP-seq profiles with matching peak and raw data (.bam) files from http://hgdownload.cse.ucsc.edu/goldenPath/hg19/encodeDCC/, and computed the percentage reads in peaks in the same manner. Since ENCODE data was collected in cell types where the factors are known to be active, for Extended Data Fig. 1f we excluded all our TF binding profiles for time points where the factors are not highly expressed and expected to be inactive (middle box plot).
To quantify the specificity of our antibodies computationally, we carried out motif analysis that measured the enrichment of 1,887 known DNA binding sequences at 500 bp regions centred on the peaks of each TF using Centrimo39 and Homer38 (Extended Data Fig. 2). For six factors, (POL2, SALL4, T, NR5A2, THAP11, TRIM28) we did not find a reliable DNA-binding motif in the database of 1,887 motifs combining TRANSFAC and Jolma et al. data sets20. For the remaining 32 TFs, we found that 88% (28/32) of factors significantly (P < 10−75) associate with the known DNA binding motif. Moreover, we carried out de novo motif discovery for these factors (using MEME40 and Homer38) and show that these motifs are highly similar to the known motifs, further supporting the specificity of these antibodies (Extended Data Fig. 2). For the other 4 factors (SRF, REX1, STAT3, TAL1) of the 32, we believe that either the known motifs in the database do not match the in vivo binding affinities for these factors in our cell types or that cross-reactivity of the antibody with other proteins is occurring. To be conservative, we have excluded all these factors from further analyses, figures, and the main manuscript.
The GATA4 and SMAD1 motif enrichment in Extended Data Fig. 8f was also carried out using Centrimo39 with weighted moving average of 50-bp window. Finally, motif enrichment for Fig. 5b was carried out by scanning 1,887 motifs (see above) within 500 bp of binding using Centrimo39 and displaying three or more of the most enriched DNA motifs per cell type.
TF dynamics and co-binding relationships
For quantifying TF dynamics between cell types and co-binding relationships between TFs, peak regions were merged if two peak centres were a distance of 1,000 bp or less, and significance P values were calculated using the hypergeometric distribution and were subsequently corrected for multiple hypothesis testing. For each TF MNChIP in each condition, we calculated a vector of the –log10 P values for interactions with all other experiments. We then clustered all vectors along both rows and columns based on correlation distance using hierarchical clustering algorithm and average linkage (Fig. 3a). We filtered all experiments with no interactions at significance level P value < 10−5 for ease of visualization. To define classes of TF binding dynamics, binding was termed enhanced/suppressed if we observed at least a twofold increase/decrease in binding sites between two different conditions. If the binding sites had not decreased/increased twofold between two conditions, we defined the co-binding relationship as static if P value < 10−300, and dynamic if P value > 10−300.
Defining chromatin state
For differential signal enrichment analysis, we first computed the number of uniquely aligned sequencing tag midpoints for all 1-kb tiles of the genomic black list filtered human genome. Genomic region black lists were obtained from http://hgdownload.cse.ucsc.edu/goldenPath/hg19/encodeDCC/wgEncodeMapability/wgEncodeDacMapabilityConsensusExcludable.bed.gz.
For each histone mark and each condition, we then determined all 1-kb tiles significantly enriched over the whole-cell extract (WCE). To that end, we fitted local Poisson models to the read count normalized WCE tag distribution for each 1-kb tile of the human genome41. Only regions enriched threefold or higher compared to the whole cell extract and significant after correcting multiple testing using the Benjamini–Hochberg method at a q value ≤ 0.05 were retained. In order to identify differentially enriched regions between ES cells and each of the ES-cell-derived populations, we took advantage of a recently published analysis strategy based on mixture models that allows to incorporate replicate information and to correct for differences in IP efficiency and signal to noise ratio42. We used the R implementation in the software package enrich to first fit a latent Poisson mixture model with two components to each ChIP-seq experiment in order to obtain an estimate of the fraction of reads in the signal component. Next, we used the initial parameter estimates from the latter model to fit a joint Poisson mixture model for each group of biological replicates. Finally, we used the obtained models for each sample group to conduct pairwise comparisons accounting for sequencing depth and differences in IP efficiency. To that end, we made the assumption that the true number of enriched regions between two compared conditions for a given mark or factor is similar and set the p parameter in the enrich mix function to 1. Finally, we obtained a list of candidates of differentially enriched regions at an FDR = 0.05 and retained only those regions that exhibited an absolute log2 difference ≥ 1.5 in the estimated tile enrichment levels and that were significantly enriched above background according to the first analysis step. Next, we specifically decided to exclude more gradual changes in histone modifications and restricted the set of differentially enriched regions to those that were above background in one but not the other condition in each of the pairwise comparisons: ES cell vs dMS, ES cell vs dEN, ES cell vs dME and ES cell vs dEC. Based on these differential analysis results, we then binarized our ChIP-seq histone modification enrichment matrix. Next, we used this binarized matrix to assign each tile one of 10 states, now also incorporating DNA methylation data. The states were defined as follows (see below for details) with their order recapitulating their precedence: H3K4me3&H3K27me3, H3K4me3, H3K27me3&H3K4me1, H3K27ac, H3K4me1, H3K27me3, unmethylated region (UMR, where 0% ≤ UMR ≤ 10% methylation), intermediate methylated region (IMR, where 10%<IMR ≤ 60% methylation), highly methylated region (HMR, where 60%<HMR ≤ 100% methylation), none (no detectable histone modification enrichment or DNA methylation data for a given 1-kb tile).
Super-enhancer analysis
Using chromatin data, we defined super-enhancers as in refs 24, 25. Briefly, we used MACS36 peak calling algorithm (default settings, except –p parameter was set to 1e−9) to detect enrichments in H3K27Ac ChIP-seq data for each cell type. Peaks were then merged if they were within a distance of 12.5 kb. We then ranked the stitched H3K27Ac enriched regions based on the normalized, background-subtracted average read density (in units of reads-per-million-mapped per bp of stitched region). The cutoff for classifying super-enhancers was defined as refs 24, 25, or the point where a line with a slope 1 is tangent to the curve of normalized region signal versus region ranking. The same procedure was used to define H3K4me1 super-enhancers per cell type.
We also used this procedure to find super-enhancers within a more inclusive set of parameters (MACS parameter –p set to 1e−5 instead of 1e−9 and stitching distance set to 5 kb instead of 12.5 kb), but found no differences in our conclusions (Supplementary Table 5). We also found no difference when using other cut-offs for defining super-enhancers (top 250, top 500, top 1,000, and top 2,000 enhancer regions, Supplementary Table 5), and found that using a fixed threshold had the advantage of uniformity between cell types in the enrichment analysis. Finally, excluding all enriched regions within 2,500 kb of TSSs also led to highly similar results and did not change our conclusions.
Chromatin states versus super-enhancers
H3K27Ac chromatin states are 1-kb genomic tiles that are significantly enriched for H3K27Ac over whole cell extract (WCE) and not enriched for other chromatin marks of higher priority. These regions are the ones displayed in the chromatin states maps that happen to fall into stitched H3K27Ac super-enhancers. For an extended H3K27Ac region to be classified as a super-enhancer, it must be enriched in H3K27Ac read density relative to all other H3K27Ac enhancer regions (not relative to WCE) for a given cell type.
TF enrichment analysis
We assessed the significance of overlap in TF binding and regions merged within super-enhancers by using the hypergeometric distribution. For each cell type, we only used TF peak regions in that cell type and super-enhancers as defined by chromatin data for that cell type. We used the same approach for measuring the TF binding enrichment at poised enhancers, or regions enriched for H3K4me1 and H3K27me3 histone modifications29. For chromatin state transition analysis, we defined the initial state as ES cells and the next cellular state as dMS or one of the three germ layers (dEN, dME, and dEC).
We then carried out TF enrichment analysis using MNChIP binding data per cell type and different epigenetic state transitions into that cell type. P values were again calculated using the hypergeometric distribution, and were subsequently corrected for multiple hypothesis testing. This analysis was used for both chromatin state transitions and DNA methylation state transitions. For Fig. 6b, we identified all differentially methylated 1-kb tiles in the genome (mean methylation difference ≥ 0.15) between ES cells and the three germ layers. In addition, we also identified regions that transitioned from an HMR state to an H3K27Ac state, termed regions that lose methylation and gain H3K27Ac. We then carried out the enrichment analysis for TF binding in these regions as described above.
Heat maps and composite plots
Heat maps were generated for regions −1 kb to 1 kb from the centre of each merged TF peak, using bins of size 50 bp. ChIP occupancy was normalized to sequencing depth as described above. Binding events for two or three ChIP-seq experiments were merged before heat map generation using Homer, as described above. ChIP-seq composite plots were generated for regions −5 kb to 5 kb from the centre of each TF peak, using bins of size 200 bp. Signal was normalized to sequencing depth, where 1 represents the mean ChIP occupancy at regions furthest from the peaks. DNA methylation composite plots were generated for regions −2 kb to 2 kb from the centre of each TF peak, using bins of size 100 bp. Mean methylation was calculated by averaging of the methylation ratio at all unique CpGs within a given bin, excluding bins with no CpGs. P values for composite plots were calculated between two samples (for example, KD and control) by finding the normalized histone mark enrichment or normalized methylation level for each sample at 300 bp regions centred around each TF peak, and then using the paired t-test. Using region size of 1 kb or 600 bp led to the same biological conclusions. Reduced-representation bisulfite sequencing captured only 1,897 of the 42,477 GATA4 bound regions in dEN and 2,331 of 35,842 GATA4 bound regions in dME with sufficient CpG methylation coverage; hence only these regions were used for the composite plots in Fig. 6f, Extended Data Fig. 10e, and associated P value calculations.
References
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005)
Young, R. A. Control of the embryonic stem cell state. Cell 144, 940–954 (2011)
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nature Genet. 42, 631–634 (2010)
Villar, D., Flicek, P. & Odom, D. T. Evolution of transcription factor binding in metazoans - mechanisms and functional implications. Nature Rev. Genet. 15, 221–233 (2014)
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nature Rev. Genet. 14, 204–220 (2013)
Cantone, I. & Fisher, A. G. Epigenetic programming and reprogramming during development. Nature Struct. Mol. Biol. 20, 282–289 (2013)
Ziller, M. J. et al. Charting a dynamic DNA methylation landscape of the human genome. Nature 500, 477–481 (2013)
Stadler, M. B. et al. DNA-binding factors shape the mouse methylome at distal regulatory regions. Nature 480, 490–495 (2011)
Gifford, C. A. et al. Transcriptional and epigenetic dynamics during specification of human embryonic stem cells. Cell 153, 1149–1163 (2013)
Lara-Astiaso, D. et al. Immunogenetics. Chromatin state dynamics during blood formation. Science 345, 943–949 (2014)
Teo, A. K. et al. Pluripotency factors regulate definitive endoderm specification through eomesodermin. Genes Dev. 25, 238–250 (2011)
Thomson, M. et al. Pluripotency factors in embryonic stem cells regulate differentiation into germ layers. Cell 145, 875–889 (2011)
Lee, G., Chambers, S. M., Tomishima, M. J. & Studer, L. Derivation of neural crest cells from human pluripotent stem cells. Nature Protocols 5, 688–701 (2010)
Hay, D. C. et al. Highly efficient differentiation of hESCs to functional hepatic endoderm requires ActivinA and Wnt3a signaling. Proc. Natl Acad. Sci. USA 105, 12301–12306 (2008)
Evseenko, D. et al. Mapping the first stages of mesoderm commitment during differentiation of human embryonic stem cells. Proc. Natl Acad. Sci. USA 107, 13742–13747 (2010)
Roadmap Epigenomics Consortium et al.Integrative analysis of 111 reference human epigenomes. Nature http://dx.doi.org/10.1038/nature14248 (this issue)
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013)
Henikoff, J. G., Belsky, J. A., Krassovsky, K., MacAlpine, D. M. & Henikoff, S. Epigenome characterization at single base-pair resolution. Proc. Natl Acad. Sci. USA 108, 18318–18323 (2011)
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012)
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013)
Harbison, C. T. et al. Transcriptional regulatory code of a eukaryotic genome. Nature 431, 99–104 (2004)
Wang, H. et al. Widespread plasticity in CTCF occupancy linked to DNA methylation. Genome Res. 22, 1680–1688 (2012)
Chambers, S. M. et al. Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nature Biotechnol. 27, 275–280 (2009)
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013)
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013)
Buecker, C. et al. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 14, 838–853 (2014)
Pikkarainen, S., Tokola, H., Kerkela, R. & Ruskoaho, H. GATA transcription factors in the developing and adult heart. Cardiovasc. Res. 63, 196–207 (2004)
Vernay, B. et al. Otx2 regulates subtype specification and neurogenesis in the midbrain. J. Neurosci. 25, 4856–4867 (2005)
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011)
Zaret, K. S. Genetic programming of liver and pancreas progenitors: lessons for stem-cell differentiation. Nature Rev. Genet. 9, 329–340 (2008)
Pouponnot, C., Jayaraman, L. & Massagué, J. Physical and functional interaction of SMADs and p300/CBP. J. Biol. Chem. 273, 22865–22868 (1998)
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008)
Zaret, K. S. & Carroll, J. S. Pioneer transcription factors: establishing competence for gene expression. Genes Dev. 25, 2227–2241 (2011)
Novershtern, N. et al. Densely interconnected transcriptional circuits control cell states in human hematopoiesis. Cell 144, 296–309 (2011)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Pickrell, J. K., Gaffney, D. J., Gilad, Y. & Pritchard, J. K. False positive peaks in ChIP-seq and other sequencing-based functional assays caused by unannotated high copy number regions. Bioinformatics 27, 2144–2146 (2011)
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010)
Bailey, T. L. & Machanick, P. Inferring direct DNA binding from ChIP-seq. Nucleic Acids Res. 40, e128 (2012)
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994)
Mikkelsen, T. S. et al. Comparative epigenomic analysis of murine and human adipogenesis. Cell 143, 156–169 (2010)
Bao, Y., Vinciotti, V., Wit, E. & ‘t Hoen, P. A. C. Accounting for immunoprecipitation efficiencies in the statistical analysis of ChIP-seq data. BMC Bioinformatics 14, 169 (2013)
Pauklin, S. & Vallier, L. The cell-cycle state of stem cells determines cell fate propensity. Cell 155, 135–147 (2013)
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012)
You, J. S. et al. OCT4 establishes and maintains nucleosome-depleted regions that provide additional layers of epigenetic regulation of its target genes. Proc. Natl Acad. Sci. USA 108, 14497–14502 (2011)
Mullen, A. C. et al. Master transcription factors determine cell-type-specific responses to TGF-beta signaling. Cell 147, 565–576 (2011)
Morsli, H. et al. Otx1 and Otx2 activities are required for the normal development of the mouse inner ear. Development 126, 2335–2343 (1999)
Greber, B. et al. FGF signalling inhibits neural induction in human embryonic stem cells. EMBO J. 30, 4874–4884 (2011)
Pérez-Losada, J. et al. Zinc-finger transcription factor Slug contributes to the function of the stem cell factor c-kit signaling pathway. Blood 100, 1274–1286 (2002)
Acknowledgements
We would like to thank all members of the Meissner laboratory for their support and feedback. We also thank F. Kelley and other members of the Broad Technology Labs and Sequencing Platform as well as J. Doench and members of the Genetic Perturbation Platform at the Broad Institute. We would like to thank L. Gaffney for graphical support. This work was supported by the NIH Common Fund (U01ES017155), NIGMS (P01GM099117), NHGRI (P50HG006193) and the New York Stem Cell Foundation. A.M.T. was supported by NIH Ruth L. Kirschstein NRSA fellowship 5F32DK095537. A.M. is a New York Stem Cell Foundation Robertson Investigator.
Author information
Authors and Affiliations
Contributions
A.M.T. and A.M. designed and conceived the study. A.M.T. performed the experiments and all analysis, H.G. generated libraries with supervision from A.G., V.A. performed cell culture, M.J.Z. helped with data processing and analysis, J.D. performed experiments, I.A. provided experimental advice, A.M.T. and A.M. interpreted the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 MNase ChIP-seq (MNChIP-seq) performance compared to sonication based ChIP-seq.
a, Venn diagram (top) and corresponding heat maps (bottom) show high reproducibility of CTCF binding between biological replicates in ES cells using MNChIP-seq. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the two replicates, where 10 = regions bound in replicate 1, 01 = bound in replicate 2, and 11 = bound in both. b, Venn diagram (top) and corresponding heat maps (bottom) show high reproducibility of NANOG binding between biological replicates in ES cells using MNChIP-seq. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the two replicates, where 10 = regions bound in replicate 1, 01 = bound in replicate 2, 11 = bound in both. c, Venn diagram (top) and corresponding heat maps (bottom) show high reproducibility of NANOG binding between biological replicates in ES cells using sonication ChIP-seq. Reproducibility of NANOG binding using sonication based ChIP-seq is similar to reproducibility using MNChIP-seq. 10 = regions bound in replicate 1, 01 = bound in replicate 2, 11 = bound in both. d, Venn diagram (top) and corresponding heat maps (bottom) show a higher sensitivity for capturing NANOG binding sites in ES cells using MNChIP-seq. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the two replicates, where 10 = regions bound in sonication ChIP-seq replicate, 01 = bound in MNChIP-seq replicate, 11 = bound in both replicates. e, Heat maps show high reproducibility of GATA4 binding in both dME (left) and dEN (right) using MNChIP-seq. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the two replicates, where 10 = regions bound in replicate 1, 01 = regions bound in replicate 2, 11 = regions bound in both. f, Top: number of significant binding peaks in our data set is comparable to that of 1,410 ENCODE TF ChIP-seq profiles (all currently available with matching peak and .bam files at UCSC). Bottom: the level of enrichment over background, as quantified by percentage of reads in peaks, is approximately 1.5 times less than that of the ENCODE TF binding data. ENCODE data was collected in cell types where the factors are known to be active; therefore, for this comparison we excluded all TF binding profiles from time points where the factors are not expressed and expected to be active (middle column).
Extended Data Figure 2 Motif analysis.
88% (28/32) of factors significantly associate with their known DNA binding motif (P < 10−75). De novo motif discovery for these factors confirms their known motifs, which provides further validation for the antibody specificity. For SRF, REX1, STAT3, and TAL1 the motifs did not match the database motifs. To be conservative, we excluded these factors from further analyses. For the remaining six factors, (POL2, SALL4, T, NR5A2, THAP11, TRIM28) we did not find a reliable DNA-binding motif in the database of 1,887 motifs combining TRANSFAC and Jolma et al. data sets20.
Extended Data Figure 3 Examples of TF binding dynamics across several loci.
a, Binding dynamics for a number of selected TFs in the four differentiated cell types versus ES cells (temporal) and in dEN versus dME (cross-lineage). b, Normalized TF binding of NANOG, EOMES, GATA4, and SMAD1 shows distinct and germ layer specific regulation of the GATA6 locus. c, Normalized TF binding at the HAND1 locus shows very static binding for NANOG between cell types, somewhat dynamic binding of OTX2 in dEN and dEC, and more dynamic binding of GATA4 in dEN and dME. Purple boxes upstream of HAND1 mark long domains of H3K27Ac, which are highly enriched for GATA4 and SMAD1 binding in dME (bottom tracks). d, Normalized MNChIP-seq binding of multiple factors across different cell types show strong enrichments over whole cell extract (WCE) control (bottom track). The high similarity in CTCF binding between cell types suggests that chromatin loops, nuclear lamina interactions, and chromatin boundaries regulated by CTCF are largely preserved during early human ES cell differentiation.
Extended Data Figure 4 Venn diagrams and heat maps highlighting different TF binding dynamics in human ES cells and their derivatives.
a, Heat maps show that CTCF binding overlaps highly in dEN and dME. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the two cell types, where 10 = regions bound in dME, 01 = bound in dEN, 11 = bound in both. b, Heat maps show that NANOG binding (left) is static in ES cells and dEN and suppressed in dME. In contrast, GATA4 binding (middle) is highly dynamic between dEN and dME and enhanced in the germ layers relative to ES cells. Finally, OTX2 binding (right) is dynamic in dEN and dEC relative to ES cells, but suppressed in dME. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the three conditions, where regions 100, 010, 001, 110, 101, 111 are defined in legend on bottom right (panel f). c, Venn diagrams (top) and corresponding heat maps (bottom) show the binding dynamics of SOX2 (left) and OCT4 (right). Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the three conditions, where regions 100, 010, 001, 110, 101, 111 are defined in legend in panel f. d, Venn diagram (top) and heat maps (bottom) show that TCF4 binding is temporally static in dMS and dEN (left) and suppressed in dME and dEC relative to dEN (right). e, Heat maps show that SMAD4 predominantly binds to unique regions in the three germ layers. f, Heat maps show that EOMES binding is enhanced from ES cells to dMS and dynamic in dEN. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the three conditions, where regions 100, 010, 001, 110, 101, 111 are defined in legend on the right.
Extended Data Figure 5 Heat maps of GATA4 and OTX2 co-binding relationship with SMAD1/4 in germ layers.
a, Venn diagrams (top) and corresponding heat maps (bottom) show that overlap in binding between GATA4 and SMAD1 is smaller in dEN (left) than in dME (right). Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the three conditions, where 10 = regions bound by factor 1, 01 = regions bound by factor 2, 11 = regions bound by both factors. Regions were considered co-bound if peaks for both factors occurred within distance d, set to 1000 bp for most analyses. Decreasing the distance d for dME to 500 bp has little effect. Setting d to 200 bp and 100 bp decreases co-bound peaks in dME by about 25% and 50%, respectively. b, Heat maps show that overlap in binding between OTX2 and SMAD1 is higher in dEN (left) than in dEC (right). Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the three conditions, where 10 = regions bound by factor 1, 01 = regions bound by factor 2, 11 = regions bound by both factors. c, Venn diagrams (top) and corresponding heat maps (bottom) show that the overlap in binding between GATA4 and SMAD4 is greater in dME than in dEN. Heat maps display normalized binding occupancy averaged using 50 bp bins. Regions are centred on the merged binding peaks for the three conditions, where 10 = regions bound by factor 1, 01 = regions bound by factor 2, 11 = regions bound by both factors.
Extended Data Figure 6 Extended H3K27Ac domains in the germ layers.
Related to Fig. 4. a, Venn diagram shows limited overlap between the top 500 extended H3K27Ac domains between cell types. b, Left, alternative lineage chromatin states of stitched dMS H3K27Ac super-enhancers (n = 698, merging 3,441 1-kb regions shown as rows in the heat map). Chromatin states (see Supplementary Information for detailed definitions of “extended H3K27Ac domains” and “H3K27Ac chromatin states”) that are displayed in the panel are defined in the legend (bottom left, HMR = highly methylated region). Center: corresponding binding of the most enriched TFs in dMS. Black bars indicate TF binding. Right, corresponding binding of selected factors in ES cells. c, Genome browser tracks for H3K27Ac across all cell types and normalized TF binding in selected cell types for EOMES, T, FOXA1/2, GATA4, and SOX17 over the EOMES locus. Grey bars highlight regions where TF binding is present in ES cells and at later stages in differentiation, suggesting that these loci are primed for binding by these factors in ES cells. Although we cannot distinguish whether this happens in all cells or just a subpopulation, it is tempting to speculate that this binding occurs in the subset of cells in G1, which is the population that is most responsive to differentiation cues43. This would also be in line with DNase I footprint studies that reported usage of EOMES DNA-binding sites in human ES cells44. d, Left, alternative lineage chromatin states of stitched H3K27Ac super-enhancers in ES cells (n = 1,052, merging 4,191 1-kb regions shown as rows in the heat map). Chromatin states that are displayed in the panel are explained in the legend in panel b (bottom left, HMR = highly methylated region). Centre, corresponding binding of the most enriched TFs in ES cells. Black bars indicate TF binding. OSN and OTX2 are the most enriched factors. Interestingly, OTX2 was recently shown to play an important role in the mouse naive to primed pluripotent state transition, a cellular state considered to be similar to human ES cells26. Right, corresponding binding data for T, EOMES, and SALL4 in dMS, showing that these key dMS regulators are present at many of these super-enhancers in the next stage of differentiation. e, Left, alternative lineage chromatin states of stitched H3K27Ac super-enhancers in dEN (n = 1,152, merging 4,051 1-kb regions shown as rows in the heatmap). States are defined as in panel b. Centre, corresponding binding of the most enriched TFs in dEN. Black bars indicate TF binding. Right, corresponding binding of selected TFs in ES cells shows that these factors occupy many of these regions in the undifferentiated state. Despite the fact that H3K27Ac domains are highly unique in the different cell types, we note that OSN, OTX2 and SMAD1 binding in undifferentiated ES cells is observed before the other factors that will mediate the transition to super-enhancer status in the three germ layers (e–h, right panels). Similarly, as noted above, regulators of super-enhancers in the germ layers also associate with these regions already in the pluripotent state. This might suggests that TF binding at germ layer specific H3K27Ac domains in the ES cells could be involved or necessary for the future handoff. Possible roles could include active regulatory binding or a way to simply mark super-enhancers; alternatively, it could also provide an active protection from silencing by the highly expressed DNA methylation machinery. In this context it is worth noting that OSN binding in the undifferentiated cells is depleted in a subset of super-enhancers that are highly methylated (e–h, bottom right) suggesting a possible binding sensitivity to DNA methylation, which has been reported for OCT4 (ref. 45). f, Left, alternative lineage chromatin states of dME H3K27Ac super-enhancers (n = 1,129, merging 4,717 1-kb regions shown as rows in the heat map). States are defined as in panel b. Centre, corresponding binding of the most enriched TFs in dME. GATA4 and SMAD1 are the most highly enriched factors at dME super-enhancers. Globally, GATA4 also interacts significantly with SMAD1 and SMAD4 in dME (hypergeometric P < 10−300) but less so in dEN (Fig. 3a, Extended Data Fig. 5c). This suggests that GATA4 interacts with SMAD1/4 at genomic targets and specifically at super-enhancers to act as a possible key regulator of the transition from pluripotent to a mesodermal state in response to BMP signalling. Recent studies have reported that master regulators in various cell types interact with TFs downstream of key signalling pathways in a similar manner46. Black bars indicate TF binding. Right, corresponding binding of selected factors in ES cells. g, Left, alternative lineage chromatin states of stitched H3K27Ac super-enhancers in dEC (n = 506, merging 908 1-kb regions shown as rows in the heat map). States are defined as in panel b. Centre, corresponding binding of the most enriched TFs in dEC. Black bars indicate TF binding. Right, corresponding binding of selected TFs in ES cells shows that these factors occupy many of these regions in the undifferentiated state. OTX2 is known to play important roles in brain, craniofacial, and sensory organ development28,47,48. In mice, Otx2 is required from E10.5 onward to regulate neuronal subtype identity and neurogenesis in the midbrain28, and inhibition of FGF signalling upregulates OTX2 and subsequently induces the neuroectodermal regulator PAX6 (ref. 48). Complementing these previous studies, our results suggest that it may play a central role in mediating the transition from pluripotency to early ectoderm. Interestingly, in dEC OTX2 does not globally associate with SMAD1 outside of super-enhancers to the same degree as in dEN (Fig. 3a). Taken together, we observe differential co-binding between SMAD1 and GATA4 or OTX2 in the respective germ layers that is linked to differential signalling, which may guide the remodelling of the associated chromatin. h, Left, alternative lineage chromatin states of the top 3,000, 1-kb-long H3K27Ac enhancers in dEC, showing a comparable number of genomic regions as in the other cell types. States are defined as in panel b. Centre, corresponding binding of OTX2 and SMAD1 in dEC shows a higher enrichment for these factors at H3K27Ac enhancers than when only surveying the top 908 1-kb regions (panel d). Black bars indicate TF binding. Right, corresponding binding of selected TFs in ES cells shows that these factors occupy many of these regions in the undifferentiated state.
Extended Data Figure 7 Quantification of cell sorting in dME on GATA4 binding and enrichment analysis.
a, Left, Venn diagrams (top) and heat maps (bottom) show that the overlap in binding between GATA4 ChIP-seq in sorted CD56+ cells and unsorted dME cells is very similar. In particular, the unique binding sites in unsorted cells (y axis label 01) also show visible but less significant binding in sorted cells, arguing that unsorted cells do not add many false positive peaks. Conversely, unique binding sites in sorted cells (y axis label 10) show that less than half of these sites are truly unique, or with no detectable binding in unsorted cells. Right, Venn diagrams (top) and heat maps (bottom) shows the overlap in binding between two GATA4 ChIP-seq replicates in unsorted dME populations. The overlap in binding between replicates using unsorted cells is similar to the overlap in binding between sorted and unsorted cells shown on the left. b, Left, alternative lineage chromatin states of dME H3K27Ac super-enhancers (n = 1,129, merging 4,717 1-kb regions shown as rows in the heat map). States are defined in legend (top right, HMR = highly methylated region). Centre, corresponding binding of GATA4 in sorted CD56+ cells, and two unsorted dME replicates (dME1 and dME2). Black bars indicate TF binding. Right, enrichment P values (–log10) for GATA4 binding at H3K27Ac super-enhancers are slightly more significant (hypergeometric P < 10−300) for unsorted cells than for sorted cells (hypergeometric P < 10−225). This shows that the conclusions for GATA4 in dME are largely unaffected by cell sorting. Moreover, since our enrichment analysis compares overlaps of binding at thousands of sites, this comparison argues that the analysis is in general robust to using unsorted cell populations. c, Enrichment P values (–log10) for the overlap in TF binding and regions that gain or lose DNA methylation relative to ES cells (see Supplementary Information). Possible transition states are defined at the top. Heat maps display the enrichment of GATA4 binding in sorted CD56+ cells, and two unsorted dME replicates (dME1 and dME2). Unsorted cells have similar enrichment P values (hypergeometric P < 10−300) to sorted cells (hypergeometric P < 10−300). This shows that the methylation conclusions for GATA4 in dME are largely unaffected by cell sorting and again argues that our enrichment analysis is robust to using unsorted cell populations.
Extended Data Figure 8 Regulation of poised enhancers and other epigenetic state transitions.
Related to Fig. 5. a, Left, alternative lineage chromatin states of dEN H3K4me1 super-enhancers (n = 309, merging 760 1-kb regions shown as rows in the heat map). Chromatin states that are displayed in the panel are explained in the legend (bottom right, HMR = highly methylated region). Right, corresponding binding of the most enriched TFs in dEN, where the black bars indicate TF binding. b, Left, alternative lineage chromatin states of the top 2,000 1-kb-long dEN H3K4me1 enhancers in dEN (shown as rows in the heat map). Chromatin states that are displayed in the panel are defined in the legend in panel a (bottom right, HMR = highly methylated region). Right, corresponding binding of the most enriched TFs in dEN, where the black bars indicate TF binding. Increasing the number of regions displayed shows a higher enrichment for dEN factors at H3K4me1 enhancers than when only surveying the top 760 1-kb regions in panel a. c, Hypergeometric P values (–log10) for the most significant overlaps between all poised putative enhancers (H3K27me3 and H3K4me1) and each TF’s binding profile in the respective cell type. Enrichment P values for dEN and dME (right column) are lower than in ES cells, which is likely the result of the overall smaller number of poised enhancers in those two germ layers. The scale is therefore adjusted for dEN and dME as shown in the respective P value legends. In ES cells, we find that poised enhancers are highly enriched for binding by OSN, OTX2, TCF4 and SMAD1 in the pluripotent state (c, d). In dMS, we see the same regulators along with T, EOMES, and LEF1 are present at poised enhancers (c, d, centre). In contrast, poised enhancers in dEN show strong enrichment for PRDM1 and many of the regulators mentioned above (c, d, right). Lastly, in dME, we find enrichment for SNAI2, which is known for its activity in mesoderm including blood development49. d, Summary table of hypergeometric P values (–log10) displaying the most significant overlaps between the top 500 poised enhancers (H2K27me3 and H3K4me1) and each TF’s binding profile within a given cell type (ES cells, left; dMS, centre; dEC, bottom centre; dEN and dME, right). Enrichment P values are more comparable between ES cells and the germ layers, since we compare TF binding with the same number of poised enhancers (500) in each cell type. The results are consistent with c, showing that the same factors are most enriched as when comparing to all poised enhancers. e, Table of hypergeometric P values (–log10) in overlap between TF binding and regions with different chromatin state transitions (relative to ES cells) within each germ layer (dEN, top; dME, middle; dEC, bottom; see Supplementary Information). Possible epigenetic state transitions are shown on top and states are defined in legend on the top left. Globally, we find a much stronger enrichment for gain of H3K4me1 in dEN than in dME, particularly for the endoderm factors present at the most methylated H3K4me1 domains. Conversely, in dME we find a strong association between remodelling of H3K27Ac and the dME factors that reside at H3K27Ac genomic regions. In concordance with this global trend, GATA4 is associated with dynamics of H3K4me1 in dEN and H3K27Ac in dME. f, Probability (y axis) of the best match to a given motif (SMAD1 and GATA4) occurring at a given position at regions centred on SMAD1 binding in dME (top) and dEN (bottom). This probability is based only on regions that contain at least one match with score greater than the minimum score defined for this motif by the default settings in Centrimo39. The position of the best GATA4 DNA binding sites (red) are more centrally enriched (P < 10−241, Centrimo39) at SMAD1 ChIP-seq peaks in dME (top) than in dEN (bottom).
Extended Data Figure 9 GATA4 knockdown experiments in dEN and dME.
a, Experimental design and data collected for the GATA4 knockdown (KD) and control experiments in dEN and dME (see Supplementary Information for details). b, Heat maps of GATA4 normalized occupancy at GATA4 targets (columns) in control and KD cell lines at corresponding genomic regions. GATA4 occupies very similar loci in control and KD cell lines in dME. c, Heat maps of GATA4 normalized occupancy at GATA4 targets (columns) in control and KD cell lines at corresponding genomic regions. GATA4 occupies very similar loci in control and KD cell lines in dEN. d, Venn diagram of dME H3K27Ac super-enhancers detected using H3K27Ac data in shControl and 3 shGATA4 KD cell lines. Super-enhancers in the shGATA4 KD lines 1, 2, and 3 overlap with super-enhancers in the shControl cell line at a much higher rate than different cell types in Fig. 4b. e, Pairwise rate of overlap between super-enhancers detected using different H3K27Ac ChIP experiments. Super-enhancers in the shGATA4 KD lines 1, 2, and 3 overlap with super-enhancers in the shControl cell line at a rate of 51.6%, 52.7%, and 47.4% (left-most bars). In comparison, the KD replicates overlap with one another at a rate of 58.8%, 59.3%, and 52.3%, and wild-type dME replicates overlap at a rate of 61.2% (middle bars). This shows that the GATA4 KD does not affect the genomic location of dME H3K27Ac domains greatly, as the overlap in domains between control and KD cell lines is only slightly lower than between H3K27Ac ChIP-seq replicates in dME. In contrast, the number of super-enhancers in common between different cell types ES cell, dEN, and dME is much lower at 14.3%, 13.7%, and 16.7% (right-most bars). Percentages are calculated relative to the experiment with fewer super-enhancers detected. f, Normalized SMAD1 (top) and H3K27Ac (bottom) mean occupancy is lower in dME for the shRNA KD lines versus control lines at SMAD1 sites both far from (distance > 1 kb, left panel) and near (distance ≤ 1 kb, right panel) from GATA4 binding (see Supplementary Information for details). The smaller decrease in occupancy away from GATA4 binding may be due to indirect effects, such as lower SMAD1 expression or co-binding with other unknown TFs.
Extended Data Figure 10 TF binding associates with specific loss of DNA methylation in dEN.
Related to Fig. 6. a, Top, genome browser tracks for H3K4me3 and H3K27me3 across four of the cell types over the SOX17 locus, zooming out on the region shown in Fig. 6a. Bottom, whole genome bisulfite sequencing (WGBS)-based CpG methylation measurements. Specific loss of DNA methylation in dEN and associated chromatin remodelling to a poised state (H3K4me3 and H3K27me3) occurs 240 kb upstream of SOX17, which coincides with loss of H3K27me3 and gain of H3K4me3 mark near the SOX17 gene. b, Top, genome browser tracks for selected TFs in different cell types upstream of SOX17. Bottom, WGBS-based CpG methylation measurements, where each rectangle represents a single CpG. Specific loss of DNA methylation in dEN coincides with specific binding of several endoderm factors. OTX2 and NANOG also bind nearby this region in ES cells. c, WGBS-based average CpG methylation level of 100-bp tiles over FOXA1 bound dEN targets in ES cells and the three germ layers shows a specific depletion of DNA methylation in dEN. d, WGBS-based average CpG methylation level of 100-bp tiles over FOXA2 bound dEN targets in ES cells and the three germ layers shows a specific depletion of DNA methylation in dEN. e, Distributions of mean DNA methylation difference in dEN between GATA4 KD and control cell lines at 1-kb regions centred on dEN GATA4 targets (left, P < 10−10, paired t-test) and at all 1-kb regions in the genome (right, P = 1, paired t-test).
Supplementary information
Supplementary Table 1
Gene expression levels in FPKM for the RNA-seq data generated for the mesendoderm (dMS) cell type in this study. (XLSX 3244 kb)
Supplementary Table 2
This table includes information for each factor (rows) in the study, including a brief description of why it was included (last column). In addition, the table includes detailed information for all antibodies used in this study, along with references that report the use and specificity of this antibody. TFs were selected through literature and computationally based on germ layer specific gene expression patterns (Fig. 1c and Supplementary Information), focusing on TFs that are relevant for the endoderm and mesoderm populations, since the additional dMS intermediate affords higher temporal resolution to capture early rewiring dynamics as well as factors involved in the lineage bifurcation. The set includes key developmental regulators as well as transcriptional effector proteins of the TGF-β (SMAD2/3/4), BMP (SMAD1/4), and WNT (TCF4, LEF1) signaling pathways, which play a critical role during the establishment of the three germ layers13,15. (XLSX 57 kb)
Supplementary Table 3
Uniquely aligned reads, number of binding events (peaks), and enrichment over background (% reads in peaks) for all ChIP-seq experiments (rows) generated in this study. (XLSX 57 kb)
Supplementary Table 4
Coordinates and annotation of the nearest gene (Homer) for extended H3K27Ac domains (super-enhancers) detected in each cell type (ESC, dMS, dEN, dME, and dEC displayed as separate sheets) and for extended H3K4me1 domains in dEN. (XLSX 808 kb)
Supplementary Table 5
TF binding enrichment analysis at super-enhancers detected using different thresholds. TF binding enrichment (–log10 P values) at super-enhancers defined as in24,25, using a variation of the published method with less stringent parameters (MACS parameter –p set to 1e−5 instead of 1e−9 and stitching distance set to 5kb instead of 12.5kb), and using several other thresholds (top 250, top 500, top 1000, and top 2000 enhancer regions). Different choices for super-enhancer threshold are displayed as different columns and cell types as different sheets. For each cell type, TF enrichments at super-enhancers are highly similar arguing that the binding enrichment analysis is robust to the choice of threshold for classifying super-enhancers. We also repeated this analysis for H3K4me1 super-enhancers in dEN in sheet “dEN_K4me1”. Finally, sheet “dENspecific_K4me1” compares TF binding enrichment analysis of all H3K4me1 regions, dEN-specific regions relative to all 5 cell types, and dEN-specific regions within the three germ layers. Spearman correlation of the non-zero enrichment values at dEN specific regions versus dEN H3K4me1 domains are shown at the bottom row of this sheet. (XLSX 71 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Tsankov, A., Gu, H., Akopian, V. et al. Transcription factor binding dynamics during human ES cell differentiation. Nature 518, 344–349 (2015). https://doi.org/10.1038/nature14233
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14233
This article is cited by
-
HAND factors regulate cardiac lineage commitment and differentiation from human pluripotent stem cells
Stem Cell Research & Therapy (2024)
-
Co-effects of m6A and chromatin accessibility dynamics in the regulation of cardiomyocyte differentiation
Epigenetics & Chromatin (2023)
-
H3.3 contributes to chromatin accessibility and transcription factor binding at promoter-proximal regulatory elements in embryonic stem cells
Genome Biology (2023)
-
H3K36 methylation maintains cell identity by regulating opposing lineage programmes
Nature Cell Biology (2023)
-
Cellular population dynamics shape the route to human pluripotency
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.