Abstract
Higher-order chromatin structure is emerging as an important regulator of gene expression. Although dynamic chromatin structures have been identified in the genome, the full scope of chromatin dynamics during mammalian development and lineage specification remains to be determined. By mapping genome-wide chromatin interactions in human embryonic stem (ES) cells and four human ES-cell-derived lineages, we uncover extensive chromatin reorganization during lineage specification. We observe that although self-associating chromatin domains are stable during differentiation, chromatin interactions both within and between domains change in a striking manner, altering 36% of active and inactive chromosomal compartments throughout the genome. By integrating chromatin interaction maps with haplotype-resolved epigenome and transcriptome data sets, we find widespread allelic bias in gene expression correlated with allele-biased chromatin states of linked promoters and distal enhancers. Our results therefore provide a global view of chromatin dynamics and a resource for studying long-range control of gene expression in distinct human cell lineages.
Similar content being viewed by others
Main

Three-dimensional genome organization is increasingly considered an important regulator of gene expression1,2,3,4. Recent high-throughput studies of chromatin structure have begun to shed light on the global organization of our genome4,5,6,7,8,9,10. For instance, we and others recently discovered that interphase chromosomes are partitioned into megabase-sized topological domains and smaller sub-domains (also known as topologically associated domains or TADs)6,7,8,9. These TADs form the basis for higher-level structures referred to as the ‘A’ and ‘B’ compartments5,6. The A and B compartments are closely linked to other functional partitions of the genome, such as early or late DNA replication timing and nuclear lamina association11,12. Despite these advances, our understanding of the dynamic nature of chromatin architecture across human cell types and its effect on cellular identity is incomplete. Here we analyse genome-wide higher-order chromatin interactions in H1 human ES cells and four human ES-cell-derived lineages, mesendoderm (ME), mesenchymal stem (MS) cells, neural progenitor (NP) cells and trophoblast-like (TB) cells13. These lineages represent extra-embryonic and embryonic lineages at early stages of development and have been extensively characterized by the Epigenome Roadmap project13, with data sets including mRNA-seq, ChIP-seq for 13-24 histone modifications, base-resolution methylC-seq and DNaseI hypersensitivity (DHS) in each lineage13,14. As such, this experimental system provides an opportunity to compare variability in higher-order chromatin structure with underlying gene expression and chromatin state in a genome-wide manner. Further, using a newly developed method to phase two parental alleles into chromosome-span haplotypes from high-resolution chromosome conformation capture (Hi-C) data15, we have phased the H1 genome to allow for analysis of allele-specific activity and chromatin structure. This represents the most extensive data set generated to date, to our knowledge, for the analysis of higher-order chromatin structure, allele-specific chromatin structure and state, and allele-specific gene expression.
Data generation and validation
We performed Hi-C experiments5 in two biological replicates in H1 human ES cells and each of the four H1-derived lineages, generating a total of 3.85-billion unique read pairs (Supplementary Table 1). We normalized the intrinsic biases in Hi-C data16, and confirmed the high reproducibility and accuracy of our Hi-C data sets using several metrics (Extended Data Fig. 1a–d, Supplementary Information and Supplementary Table 2).
Extensive A/B compartment switching
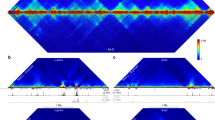
Hi-C interaction maps provide information on multiple hierarchical levels of genome organization4. Previous studies demonstrated that the genome is organized into A and B compartments, containing relatively active and inactive regions, respectively5,11. Currently, it is unclear if the A and B compartments change during differentiation and how this relates to lineage specification. We observe a large degree of spatial plasticity in the arrangement of the A/B compartments across cell types, with 36% of the genome switching compartments in at least one of the lineages analysed (Methods; Fig. 1a and Extended Data Fig. 2a–c). Many of the A/B compartment transitions are lineage-restricted (Fig. 1b). Notably, there appears to be a large expansion of the B compartment upon differentiation of human ES cells to MS cells or in IMR90 fibroblasts. These two cell types have previously been shown to undergo an expansion of repressive heterochromatin modifications during differentiation13,17. In this regard, there appears to be a similar redistribution of the spatial organization of their genomes as well. We observe that the regions that change their A/B compartment status typically correspond to a single or series of TADs (Fig. 1a, c and Extended Data Fig. 2d, e), suggesting that TADs are the units of dynamic alterations in chromosome compartments. Consistent with previous studies of individual loci18,19,20, we found that genes that change from compartment A to B tend to show reduced expression, whereas genes that change from B to A tend to show higher expression (Fig. 1d). In addition, lineage-restricted compartment A regions tend to include more lineage-restricted genes compared to other regions (Extended Data Fig. 3a). Although statistically significant, the overall patterns of change in expression are subtle. Reasoning that this modest correlation may be due to the possibility that only a subset of genes may be affected by compartment changes, although most genes remain unaffected, we identified a subset of 718 genes with co-variation between gene expression and compartment switching (Fig. 1e, Extended Data Fig. 3b, c, and Methods). These genes were enriched for low CpG content promoters (21.8% versus 15.6% for non-concordant genes, P value 8 × 10−11, Fisher’s exact test), and several significant Gene Ontology (GO) terms, most notably related to extracellular proteins and extracellular matrix (Supplementary Table 3). Taken together, these results indicate that at a global level, there is a high degree of plasticity in the A and B compartments, yet relatively subtle corresponding changes in gene expression, indicating that the A and B compartments have a contributory but not deterministic role in determining cell-type-specific patterns of gene expression.
a, First principal component (PC1) values and Hi-C interaction heat maps in H1 ES cells and H1-derived lineages. PC1 values are used to determine the A/B compartment status of a given region, where positive PC1 values represent A compartment regions (blue), and negative values represent B compartment regions (yellow). Dashed lines indicate TAD boundaries in ES cells. b, K-means clustering (k = 20) of PC1 values for 40-kb regions of the genome that change A/B compartment status in at least one lineage. c, K-means clustering of PC1 values surrounding TAD boundaries (‘b’ denotes boundary location). d, Distribution of fold-change in gene expression for genes that change compartment status (‘A to B’ or ‘B to A’) or that remain the same (‘stable’) upon differentiation (P < 2.2 × 10−16, P values by Wilcoxon test; whiskers correspond to interquartile range). e, Genome browser for two genes of which one (OTX2) shows concordance between expression and PC1 values, whereas a second (TMEM260) does not.
Domain-level chromatin dynamics
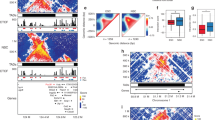
We next examined higher-order chromatin structure at a sub-chromosomal scale. Previous studies indicated that chromosomes are composed of cell-type-invariant TADs6,8. Across the six lineages analysed in this study, we observe that although the positioning of TADs remains stable between cell types (Fig. 2a), numerous changes in chromatin structure occur within domains. We observed a phenomenon that within some domains, a large portion of the interactions appears to increase or decrease across the entire domain between cell types (Fig. 2b). This suggests that a subset of TADs in a given lineage undergo concerted, domain-wide changes in interaction frequency. Hundreds of TADs underwent such alterations in each lineage (Fig. 2b and Extended Data Fig. 3d), with the changes in interaction frequency correlated positively with active marks such as DHS, H3K27ac and with CTCF binding, and negatively correlated with repressive chromatin modifications such as H3K27me3 and H3K9me3 (Fig. 2c, see Methods for details). TADs that have a concerted increase in intra-domain interaction frequency tend to shift from the B to A compartments, while domains that have a concerted decrease in interaction frequency tend to shift from A to B (Extended Data Fig. 3e, f). Consistent with the changes in chromatin state activity, genes within domains that have increased intra-domain interaction frequency tend to be upregulated, while genes within domains that decrease intra-domain interaction frequency tend to be downregulated (Extended Data Fig. 3g, h).
a, Chromatin interaction heat maps in H1 lineages and IMR90 fibroblasts. Also shown are domain calls in ES cells and the directionality index (DI) in each lineage. b, Changes in interaction frequency between ES and MS cells. Regions with higher interaction frequency in ES cells are shown in blue, while regions with higher interaction frequency in MS cells are shown in yellow. TADs having a concerted increase or decrease in intra-domain interaction frequency are labelled yellow or blue, respectively, with the fraction of the domain showing increased or decreased interaction frequency listed. Domains that do not show a concerted change are shown in grey. c, Boxplots of Pearson correlations coefficients between interaction frequency changes and chromatin mark changes across TADs for each chromosome (n = 23). Whiskers correspond to the highest and lowest points within 1.5× the interquartile range. d, Classification accuracy of the Random Forest model in predicting whether a bin increases or decreases in interaction frequency (n = 768,793), tested on 10 randomly selected subsets of Hi-C data. Accuracy was also checked using actual data (blue), circularized permutation (green) and a random permutation (yellow) of the data. As expected, randomly permuting the data yields 50% accuracy. Accuracy was also assessed considering the top 30, 40, 50% or all predictions based on vote frequency difference (error bars show the standard deviation of accuracies from the 10 randomly selected data subsets). e, Ranked chromatin features shown according to importance in classification as boxplots of the mean decrease in Gini index from 10 randomly selected data subsets. Whiskers correspond to the highest and lowest points within 1.5× the interquartile range.
Chromatin state and dynamic interactions
In order to understand the relationship between chromatin dynamics and other genomic and epigenomic features, we performed integrative analysis of the Hi-C data along with the histone modifications, DHS, and CTCF binding data in the six lineages. Specifically, we asked if particular chromatin state patterns predict changes in chromatin interaction frequency. We divided the genome into 40-kb bins and computed changes in chromatin features in each bin upon differentiation. We then built a Random Forest classification model based on chromatin features to classify local interacting bins as having either increased or decreased interaction frequency (see Methods for details). The model was able to classify regions of the genome that increased or decreased interaction frequency with 73% accuracy (Fig. 2d, 100% graph; Extended Data Fig. 4a), which increased to over 80% when we consider only the highest confidence predictions as based on the vote frequency difference (Fig. 2d, 30% graph). The Random Forest model not only indicates that chromatin state features provide information on changes in interaction frequency, it also allows us to determine which chromatin marks are most predictive. Specifically, the ‘mean decrease’ of the Gini index for each chromatin mark indicates the importance of a given feature during classification. In this regard, we found that change in H3K4me1 density is the most important feature in predicting changes in long-range chromatin interactions (Fig. 2e and Extended Data Fig. 4b, c). As H3K4me1 is present mostly at poised or active enhancers21,22, and as enhancers are known to engage in looping interactions that exist in a cell-type-specific manner23, these results suggest that enhancer dynamics may play a role in regulating local interaction changes during lineage specification. Consistent with this hypothesis, 40-kb regions with increased interaction frequency tend to have increased enhancer density (Extended Data Fig. 4d, e).
Allele-specific chromatin organization
Normal diploid human cells contain two copies of each chromosome. The collection of variants on a given parental chromosome (also known as the parental haplotype) can be used to determine functional differences between two homologous chromosomes. Previous studies have revealed substantial differences between alleles in gene expression, DNA methylation, and chromatin states24,25,26,27,28,29. Apart from studies of individual loci in the genome30,31,32, little is known about the variability in higher-order chromatin structure between homologous chromosomes. Recent work from our laboratory15 has demonstrated that Hi-C data can be re-purposed to reconstruct chromosome-span haplotypes, which allows for the study of chromatin state and gene expression as a true diploid. We generated chromosome-span haplotypes incorporating ∼93.5% of all heterozygous variants for H1 from a combination of Hi-C data sets, whole genome sequencing, and local conditional phasing15 (Fig. 3a). We observe a high level of concordance among the predicted haplotypes and paired sequence reads from data sets with ‘long insert’ sizes (Extended Data Fig. 5a), indicating that the reconstructed haplotypes are of high quality. Next, we re-analysed data sets from Hi-C, mRNA-seq, ChIP-seq, methylC-seq, and DNase-seq experiments and determined from which parental haplotype each sequence read was derived (arbitrarily termed the ‘p1’ and ‘p2’ allele, as we cannot determine which is the maternal or paternal copy from sequence information alone) (Fig. 3b and Extended Data Fig. 5b).
a, Variants per megabase for all (green), phased (orange) and unphased variants (purple) along chromosome 1. The inset zooms in on a ∼1 Mb region, where the presence of a variant at each base is indicated by a value of 1. b, Genome browser image of allele specific chromatin features and strand-specific mRNA sequencing. c, Genome browser image of PC1 values along chromosome 2 for the p1 and p2 allele. d, Allele specific compartment A/B patterns and mRNA-seq surrounding the imprinted ZDBF2 gene. e, Boxplots of the difference between alleles of PC1 values. Regions with imprinted genes (P = 0.003) and allelic genes (P = 0.002) have more variable PC1 values (Kolmogorov–Smirnov (KS) test). Whiskers correspond to the highest and lowest points within 1.5× the interquartile range. f, Similar to e, but for regions with differential allelic chromatin activity (the number of allelic biased variants per 200-kb bin). Regions in the top 0.1% of differential allelic activities (orange) show greater differences in PC1 values compared other regions (P = 1.6 × 10−8 and P = 0.0015, respectively, KS test).
From the haplotype-resolved A and B compartment patterns across the p1 and p2 alleles in each lineage, we found that homologous chromosomes have highly similar A/B compartment patterns (Fig. 3c and Extended Data Fig. 5c–e), with only 0.6–2.3% of the genome having different A/B compartments between alleles in any given cell type (Extended Data Fig. 5f). Notably, rare regions of the genome do show changes in A/B compartment status between alleles (Fig. 3d), but are not enriched for either allele-biased or known imprinted genes (Extended Data Fig. 5g, h). On the contrary, regions of the genome containing allele-biased or imprinted genes have a subtle but statistically significant increase in the variability of A/B compartment scores between alleles (Fig. 3e). Likewise, the genomic regions with allelic chromatin states have greater variability in A/B compartment scores (Fig. 3f). This indicates that although most allele-biased and imprinted genes do not have differential compartment status between alleles, there may be subtle differences in higher-order chromatin structure between homologous chromosomes at allele-biased regions, reflecting their underlying allele biases in activity. Lastly, similar to A/B compartment patterns, topological domain patterns appear consistent between alleles (Extended Data Fig. 6a, b). Together, these results suggest that the global folding patterns of homologous chromosomes are highly similar.
Allelic imbalances in gene expression
Previous studies of allele-resolved gene expression have identified widespread imbalances in gene expression between different alleles24,25,26,27,33. However, it remains unclear to what degree allele-biased gene expression varies among different lineages of a single individual. To address this, we re-analysed haplotype-resolved mRNA-seq data and identified allelic biases in gene expression across the five H1 lineages. A total of 1,787 genes showed allelic bias in gene expression in one or more lineages studied here, representing ∼24% of all testable genes (false discovery rate (FDR) 10%, Fig. 4a). Most allelic differences in expression are not ‘on/off’ events, but instead reflect biases in the level of expression from each allele (Fig. 4b). Further, allele-biased genes include both lineage-specific and constitutively expressed genes (Extended Data Fig. 6c, d), and patterns of allelic bias can also be constitutive or cell-type variable (Fig. 4c, d). Only in rare cases do genes switch expression from one allele to the other between cell types.
a, Proportion of genes with detectable allelic expression with statistically significant allelic bias. b, Density plot of the absolute value of the fold change in expression (log2) between alleles. c, Heat map showing k-means (k = 20) clustering of the allelic expression ratios (log2) at genes with constitutively testable expression (a minimum of 10 reads in each lineage). d, Genome browser image of variable allelic expression of the PARP9 gene. e, Fraction of imprinted genes among allele-biased genes and other genes. (P = 4.4 × 10−5, Fisher’s exact test). f, Fraction of allele-biased genes that are known imprinted genes. g, Cumulative density plot of distances from variants to the nearest allele-specific gene. Allele specific variants are defined using histone acetylation, H3K9me3, H3K27me3, DHS and H3K4me3 (n = 3,920, P < 2.2 × 10−16, KS test). h, Number of allele-biased genes showing consistent allele specific chromatin states in their promoter regions. Active variants are defined by H3K4me3, DHS or histone acetylation. Inactive promoter variants are defined by DNA methylation and H3K9me3/27me3. i, Genome browser image of mRNA-seq and chromatin features surrounding the TDG gene.
As expected, genes subject to genomic imprinting are enriched among genes with allelic biases in expression (Fig. 4e), though these represent ∼1% of allele-biased genes (Fig. 4f). Although imprinted genes often occur in clusters, the majority of allele-biased gene expression is not clustered in the genome (Extended Data Fig. 6e). Taken together, these data suggest that most instances of allele-biased gene expression are due to mechanisms other than genomic imprinting. One possible regulatory mechanism that could give rise to allele-biased expression would be allelic bias in activity of cis-regulatory elements near these genes. Indeed, regions of the genome that show allele bias in histone acetylation, histone methylation, CTCF binding, and DHS are closer to allele-biased genes than randomly selected genomic regions (Fig. 4g). Furthermore, allelic gene expression is strongly correlated with DNA methylation or chromatin modification state at promoters (Fig. 4h, i). Of the 247 genes that contain heterozygous variants in their promoter regions and display biased transcription in at least one lineage, a majority exhibit allele-biased chromatin modifications or DNA methylation at the promoter (Fig. 4h). Interestingly, 29% of the testable genes that have allele-biased expression show no evidence of allelic bias in chromatin state or DNA methylation at the promoter (Fig. 4h), raising the possibility that elements outside of promoters may be responsible for the allelic gene expression.
We identified 726, 969, and 5,769 allelic enhancers13 that showed allele bias in histone acetylation, DHS, and DNA methylation, respectively (Fig. 5a). We observed a general concordance in allelic biases between enhancers exhibiting allelic histone acetylation and enhancers showing allelic DHS (Fig. 5a). However, we observe only modest concordance between DHS or acetylation defined enhancers with those identified based on allelic DNA methylation (Fig. 5a). This may reflect greater power in identifying differentially methylated regions between the two alleles. Alternatively, this may reflect the presence of ‘poised’ enhancers, where there is not a strict relationship between differences in DNA methylation and enhancer or DHS state34,35. Enhancers with allele-biased acetylation are generally located closer to genes that also show allele-biased expression when compared with enhancers that lack allele bias (Fig. 5b and Extended Data Fig. 6f). A majority (66%) of the 640 allelic genes that display strong Hi-C interactions with allelic enhancers also show concordant allelic activity between the enhancer and promoter (Fig. 5c, Extended Data Fig. 7, and Methods). Additionally, enhancer-gene pairs linked by relatively strong Hi-C interactions show greater correlation between allelic enhancer activity and allelic gene expression compared with pairs linked by weaker Hi-C interactions (Fig. 5d). To test if allelic enhancers indeed form specific contacts with allele-biased genes, we performed 4C-seq31,36 with 6 allele-biased enhancers and identified that 4 out of these 6 allelic enhancers showed specific 4C interactions with a nearby allele-biased gene (Fig. 5e, Extended Data Fig. 8 and Supplementary Table 4). Taken together, our results strongly support that allele-biased enhancer activity is a possible mechanism underlying allele-biased gene expression.
a, Enrichment of acetylation (top row), DHS (middle) and DNA methylation (bottom) at enhancers defined as allelic by acetylation (left column), DHS (middle), or DNA methylation (right). The active allele is in blue, inactive allele in red. b, The distance between allelic genes and enhancers as defined by allelic histone acetylation (purple) compared with randomly selected enhancers (grey). Random enhancers were selected to match the read coverage of allele-biased enhancers. c, Number of allele specific genes linked to concordantly biased allele specific enhancers. Genes linked by ‘long-range enhancers’ are defined using Hi-C interaction frequencies, whereas ‘short-range enhancers’ are defined as any enhancer less than 20 kb from a genes transcription start site. d, Boxplots of the Pearson correlation coefficients between allelic gene-enhancer pairs defined by acetylation (left, n = 1,388) or DHS (right, n = 1,601). Gene-enhancer pairs are grouped into strongly interacting (top 30%), weakly interacting (bottom 30%), and intermediately interacting pairs (others) based on Hi-C interaction frequency (P values using Welch’s t-test). Whiskers correspond to the highest and lowest points within 1.5× the interquartile range. e, Normalized 4C-seq interaction frequencies near the HAPLN1 gene. The 4C-seq bait region is in an allele-biased enhancer near the 3′ end of the EDIL3 gene. Specific interactions called by the LOWESS regression model are shown in black as ‘bait interacting regions’ (BIRs). f, Allele-biased expression of the two alleles of the HAPLN1 gene, histone acetylation levels at the nearby interacting allele-biased enhancer and allele resolved 4C-seq data (4C-seq P value from t-test, n = 2 for p1 allele, n = 2 for p2 allele).
To determine if part of the mechanism of regulation by allele-biased enhancers also involved allelic chromatin looping between distal enhancers and putative target genes, we tested for the presence of allele-biased Hi-C reads at allele-biased enhancers throughout the H1 genome by aggregating all Hi-C reads between allelic enhancers and the promoters of nearby allelic genes. We observed that alleles containing enhancer activity generally have higher numbers of chromatin interactions with the target promoters (Extended Data Fig. 9a). This result is confirmed by re-analysis of previous high-resolution 4C-seq results31. Two loci (HAPLN1 and MAN1C1) show a similar trend between allele bias in enhancer–promoter interactions with the allelic enhancer acetylation and gene expression levels (Fig. 5f and Extended Data Fig. 9), though the trend in the allelic 4C-seq does not meet statistical significance. The remaining two loci (FAM65B, PXK) appear to have nearly equal interaction frequencies with the target promoters. Taken together, these results suggest that the allele-biased enhancers can impart allele-biased gene expression either through stable higher-order DNA looping between the two alleles or through potential allele-specific enhancer–promoter interactions.
Discussion
We have presented genome-wide chromatin interaction maps in H1 human ES cells and four H1-derived lineages. We observed dynamic reorganization of higher-order chromatin structure during ES cell differentiation at multiple hierarchical scales. We found extensive switching between the A and B compartments during ES cell differentiation, and observed that distinct subsets of genes have concordant A/B compartments status and expression levels. In this regard, these results are similar to what has been seen with nuclear lamina tethering studies20,37,38,39, where the expression of only a subset of genes is affected by compartment changes, while other genes remain unaffected. Changes in compartment status may influence the accessibility of genomic regions to transcription factors or other regulatory proteins, which may be particularly important for certain subsets of genes.
In addition, we have observed local alterations in chromatin interaction frequency within TADs. These local changes are best predicted by changes in levels of H3K4me1 and the density of enhancer elements. This is in agreement with recent 5C studies demonstrating that cell-type specific interaction regions are enriched for Smc1, mediator, and transcription factor binding sites7. Taken together, these results suggest that enhancer elements likely play an important role in shaping local higher-order chromatin structure throughout the genome. In addition, by analysing patterns of chromatin interactions on each parental allele, we observe relatively minor global changes in higher-order chromatin structure between alleles.
The chromatin interaction maps generated in this study also allowed the reconstruction of chromosome-span haplotypes for the H1 genome. This data set represents one of the first studies of allele-biased expression across multiple cell types of a single individual, as well as analysis of chromatin state at the linked cis regulatory elements. Our data set will serve as a valuable tool for the community to better understand the gene regulatory networks controlling pluripotency and differentiation of human embryonic stem cells.
Methods
Cell culture and previous data sets analysed
H1 human ES cells and H1-derived cells were cultured as previously described13. ChIP-seq experiments for CTCF were performed using previously published methods and antibodies13,40. Hi-C libraries were generated as previously described5. Two biological replicates of Hi-C data were generated for each lineages in order to assess the reproducibility of the data. Hi-C and ChIP-seq libraries were sequenced on the Illumina Hi-Seq 2000 and Hi-Seq 2500 platforms. mRNA-seq, ChIP-seq for histone modifications and methylC-seq data sets have been previously published13. DNase-seq experiments have been previously described elsewhere14.
Sequence read alignment
The following description applies for the alignment of DNA methylation, ChIP-seq and DNase-seq data sets. Single-end sequencing data was mapped to a variant masked reference genome (hg18) using Novoalign. Unmapped and non-uniquely mapping reads were removed, and PCR duplicate reads were removed with Picard. Reads were processed with the Genome Analysis Toolkit (GATK)41. Specifically, reads underwent indel recalibration and variant realignment. Lastly, reads that overlapped with variant loci were split into the ‘p1’ and ‘p2’ allele according to whether the base in each sequencing read matched the sequence from either the p1 or the p2 alleles.
For Hi-C data sets, read pairs were mapped independently to the variant masked genome using Novoalign. Reads were then manually paired using in house scripts. Non-uniquely mapping, unmapped reads and PCR duplicate read pairs were removed. Reads pairs were then split into single reads and processed through the same GATK pipeline described above including indel re-alignment and variant recalibration. Finally, read pairs were manually re-paired using in house scripts.
For mRNA-seq, we mapped the paired-end data to a variant masked reference. We used Useq software to first process the variant masked genome to create a splice junction reference. Reads were then mapped to the Useq processed reference genome using Novoalign. Lastly, we converted the read alignment locations from the Useq processed genome back to hg18 coordinates using Useq.
Whole-genome sequencing, genotyping and haplotyping
Whole genome sequencing (WGS) data for the H1 genome was downloaded from the Sequence Read Archive database (SRA049981). Reads were mapped to the hg18 reference genome using Novoalign. Unmapped and non-uniquely mapping reads were removed using in house scripts. PCR duplicate reads were removed using Picard. The data was processed through the Genome Analysis Toolkit (GATK) best practices guidelines. We performed indel recalibration, variant realignment, variant calling using the Unified Genotyper, and variant recalibration.
Haplotyping was performed using the previously described HaploSeq method15. Briefly, Hi-C reads from each of the H1 derived lineages were used as input sequencing into the HapCUT software42 in order to generate haplotype predictions. For final haplotype calls, Hi-C data was combined with WGS mate-pair data for the H1 genome. HapCUT generates several ‘blocks’ for each chromosome. The vast majority of variants on each chromosome are in the ‘most variants phased’ (MVP) block. The MVP block for each chromosome was used as a ‘seed haplotype’ for local conditional phasing using population sequencing data from the 1000 genomes project using the Beagle v.4.0 software43. This generates two haplotypes for each chromosome, one for the maternal allele and one for the paternal allele. As we do not have information regarding the parent of origin in the H1 genome, we arbitrarily define each allele as the p1 or p2 allele (p1 and p2 for parent 1 and 2, respectively). The p1 and p2 allele for different chromosomes are not necessarily derived from the same parent, as this information is only accessible if the sequence of H1’s parents were also available.
Haplotype alignment bias
Although we mapped the ChIP-seq, DNase-seq, Hi-C and DNA methylation data sets to a variant masked genome, we recognize that there could still be local alignment biases favouring a given allele. To account for this, we performed a two-step filtering process. First, we generated simulated reads that span each position surrounding a variant location in the genome. SNPs and indels that showed >5% and >10% biases, respectively, were excluded from all downstream analyses, as these variants show an inherent mapping bias. Second, for each variant in the genome, we calculated the coverage over the variant based on WGS data. Based on the WGS data, we expect each variant to have near equal coverage between the two alleles. Any variant that had sequencing coverage greater than 3 standard deviations above the mean for each haplotype along a chromosome was excluded, as were variants that showed a Benjamini corrected binomial P value of ≤0.05 when comparing the WGS read coverage on each allele. Lastly, analysis of allele-biased coverage at a SNP level can be very sensitive to genotyping errors, in particular if a homozygous variant is erroneously called as heterozygous. To account for this we made a null hypothesis that all called heterozygous variants were actually homozygous. We excluded any heterozygous variant with a GATK derived genotype P value of greater than 0.05 (after Benjamini correction). This excluded roughly 2% of all heterozygous SNPs in the genome as having genome sequencing coverage that could be expected for a homozygous variant.
Estimation of random collision events in Hi-C data
We estimated random collision events by calculating the intermolecular ligation rate between a nuclear chromosome (chrN) and the mitochondrial chromosome (chrM). The interacting space between chrN and chrM can be defined by multiplying (roughly 16 kb per chrM × number of chrM per chrN) and (roughly 6.16 Gb per diploid nucleus). The number of chrM per chrN was calculated from ChIP-seq input sequencing data.
Number of chrM per chrN = Number of read counts for chrM/number of read counts for chrN × 6.16 Gb per chrN × 16 kb per chrM.
The number of random collision events between any given two loci (40-kb bin size) was estimated as following.
Number of random collision events per 40 kb2 = number of intermolecular interactions between chrN and chrM/interacting space between chrN and chrM × 40 kb2.
The estimated random collision events are summarized in Supplementary Table 2.
Topological domain calling
We systemically identified topological domains based on the directionality index (DI) score and a Hidden Markov Model (HMM) as previously described6. The number of identified topological domains across human genome was 2,468, 2,489, 2,202, 2,144 and 2,407 for ES, ME, MS, NP and TB cells, respectively. According to the topological domain patterns, genomes were partitioned into domains, boundaries and unstructured regions as previously described.
Identification of A and B compartments
Identification of A and B compartments was performed conceptually similarly to what has been previously described5, though with several modifications. We used the normalized 40-kb interaction matrices for each cell type and calculated the expected interaction frequency between two 40-kb bins given the distance separating them in the genome. We used a sliding window approach with a bin size of 400 kb and a step size of 40-kb to generate an observed/expected matrix. The observed frequency was the sum of all observed interaction frequencies of the 40-kb bins making up the larger 400-kb bin. Likewise, the expected frequency was the sum of the expected frequencies of each of the 40-kb bins making up the larger 400 kb bin. This value was used to generate the observed/expected. This was then converted to a Pearson correlation matrix and subsequently used for principal component analysis as previously described5. Specifically, we used the ‘cov’ function in R to generate a covariance matrix from the Pearson correlation matrix, and then we used the ‘eigen’ function in R to generate Eigen vectors and Eigen values from the covariance matrix. The first principal component for each chromosome was used to identify regions of the genome as belonging to either the A or B compartment. The direction of the Eigen values is arbitrary, and therefore positive values were set to ‘A’ and negative values were set to ‘B’ based on their association with gene density.
To identify regions of the genome that switched A/B compartment status with differentiation, we first identified regions with statistically significant variability in PC1 values across all cell types using ANOVA. Second, we considered only regions where both biological replicates showed changes in PC1 values from positive to negative or vice versa. This allowed us to define the 36% of the genome that changes compartment status in at least one lineage.
Identification of genes with concordant expression and A/B compartment status
To define genes with concordant changes in expression and compartment status, we calculated the covariance between the vector of the log2 of gene expression values and vector of PC1 values for each gene across the six lineages analysed. We use this calculated covariance as a metric to quantitatively define ‘concordance’. To calculate a P value for the covariance for each gene, we compared these observed covariance values to a random background distribution. The background distribution was generated by randomly shuffling the vector of log2 of gene expression for each gene and then calculating the covariance between the random gene expression vector and the PC1 values. This was repeated 1,000 times for each gene, and a rank-based P value could then be calculated for the observed covariance values. These genes were shown to be enriched for low CpG content promoters, which is defined here by an observed/expected CpG content of <0.35. GO terms analysis of this subset of genes was performed using the DAVID GO terms website.
Identification of A and B compartments in each allele
Identification of A and B compartments in each allele was performed similarly as described in the above section, though with several modifications. Due to the low density of Hi-C interaction frequencies in each allele, we used a sliding window approach with a bin size of 1-Mb and a step size of 200-kb to generate an observed/expected matrix. The first principal component in each allele was used to identify regions of the genome as belonging to either the A or B compartment. The direction of the Eigen values is arbitrary, and therefore the direction was determined according to the correlation coefficient values with the PC1 values generated in the above section.
Changes in intra-domain interaction frequency
To compute the change in interaction frequency between cell types, we first merged the Hi-C data between two replicates for each cell type. The merged, normalized interaction matrices were quantile normalized between all lineages to accommodate for differences in frequency strictly due to sequencing depth. The differences between cell types were computed by simply subtracting the interaction frequency of each bin Iij of ES cells from the differentiated cell types (as shown in Fig. 2b).
To assess for concerted domain-wide changes in interaction frequency, we calculated two values for each domain: the fraction of interacting bins in the domain that showed an increase in interaction frequency and the fraction of bins that showed a decrease in interaction frequency. To compare these numbers to what would be expected at random, we calculated the same two values for each domain where the bins of the domain where made up of randomly selected intra-domain interacting bins from throughout the genome, keeping the portion of bins in each domain separated by a given genomic distance constant. This randomization was performed 10,000 times for each domain. At random, each domain on average had roughly 50% of bins that increased in interaction frequency and 50% that decreased in interaction frequency. By seeing deviations from these expected values, we could assess for ‘concerted’ changes in interaction frequency. We assigned a rank-based P value of the degree of ‘concertedness’ for each domain by comparing the actual observed portion of the domain that was either increased or decreased in interaction frequency with what was observed at random for each domain. These P values were adjusted for multiple testing using Benjamini correction, and we considered any domain as having undergone a concerted change if the final corrected P value was less than 0.001 (0.1% FDR).
Changes in intra-domain interaction frequency between alleles
Domain-wide interaction frequency differences between alleles were calculated by using the same approach described in the above section. If the domain-wide average interaction frequency difference between alleles was significantly more than randomized data (P value 0.001), the corresponding domains are considered as having allele specific domain-wide interaction frequency changes.
Correlation coefficient between domain-wide interaction frequency changes and modification changes
The domain-wide correlations shown in Fig. 2c between changes in interaction frequency and various chromatin marks were calculated as follows. For each domain, the intra-domain interaction frequency differences between ES cells and each differentiated lineage was calculated for each 40-kb interacting bin of the domain (where we define a single ‘interacting bin’ as being formed by the interaction of two underlying 40-kb genomic bins). These values were considered as the first vector for the correlations. The vector of histone modification values was calculated as follows. For each 40-kb interacting bin, the enrichment of a given chromatin mark in the two 40-kb bins that compose the interaction was averaged. The average enrichment was then multiplied by a weight proportional to the genomic distance between the two 40-kb bins. This weight was based on the global average of Hi-C interaction frequencies from six lineages analysed between loci separated by a given genomic distance. The two vectors were used to calculate a Pearson correlation in each chromosome, which reflects how change in domain-wide interaction frequency correlates with domain-wide chromatin mark changes.
The Random Forest classification model
We built a Random Forest model to better understand which chromatin modifications may be most predictive of changes in interaction frequency between any two given loci. The aim of the Random Forest model was to classify 40-kb interacting bins as either increased or decreased in interaction frequency given information about the enrichment of various chromatin marks, DHS and CTCF binding sites. The utility of the Random Forest model is twofold: first, by assessing the accuracy of the model using observed data, we can learn whether the information supplied to the model (in this case the chromatin state, DHS and CTCF data) is predictive of the outcome, namely changes in interaction frequency. The second powerful aspect of the model is that it allows us to assess which input data supplied to the model is most informative, allowing us to determine which chromatin state features may be most predictive of changes in higher-order chromatin structure.
The model was built as follows: 40-kb interacting bins in the genome were classified into two groups, ones that increased in interaction frequency, and ones that decreased in interaction frequency. These changes were defined if the 40-kb based interaction frequencies increased or decreased more than twofold in the differentiated lineage compared to those in H1 ES cells. We only considered interacting bins separated by less than 2 Mb. We added a pseudocount value to the average interaction frequencies when we calculate fold changes to allow for comparison of zero values. The resulting criteria yielded 768,793 interacting bins as either losses or gains. Chromatin state changes of H3K4me1, H3K4me3, H3K9me3, H3K27ac, H3K27me3, H3K36me3, DHS signal, and CTCF were also calculated. For each 40-kb bin, RPKM values for each chromatin mark were calculated. Fold changes of RPKM values were calculated by comparing with RPKM values in H1 ES cells. Those 8 chromatin marks were assigned to each interacting region, thus for each interaction (consisting of two interacting 40-kb bins) we can construct a 1 by 16 feature vector.
Using those feature vectors, we built a classification model between gain and loss of interactions using a Random Forest R package with default parameters except for specifying the model to use 500 trees. The performance was measured according to two criteria, the out-of-back error rate achieved from the Random forest model and the tenfold cross validation. We compared out-of-back error rate to tenfold cross validation and observed very similar results (shown in Extended Data Fig. 4a).
As a final result, the Random Forest model gives vote frequencies for predicting whether a given interaction is increased or decreased. The difference in vote frequency between the two states reflects the confidence of the model in a given prediction, with a larger vote frequency difference indicating a higher degree of confidence. The sum of the vote frequencies is equal to 1. As an example, in the case where the model could not predict any changes in interaction frequency, the vote frequencies would be expected to both equal 0.5. If the vote frequency for the ‘loss of interaction’ class was greater than 0.5, the interacting bin would be classified as having undergone a loss of interaction. Likewise, if the ‘gain of interaction’ vote frequency was greater than 0.5, the bin was classified as a gain of interaction. Again, the difference in vote frequencies between the two classes reflects the degree of confidence of the model in a given prediction.
When we built the classification model, the balance for the number of inputs between two classes is important. If the model includes more gain of interaction features rather than loss of interaction features, the model is more likely trained to predict a gain of interactions. To avoid this issue, we randomly selected the same number of gain of interaction and loss of interaction feature vectors while building the classification model.
The Random Forest model also provides a measure of the importance of each variable during classification as the ‘mean decrease’ metric of the Gini index. For a given variable, higher the mean decrease in Gini index, the more important the variable is during classification.
Identification of allelic biased genes, enhancers and SNPs
Allelic genes
We considered the two replicates of mRNA-seq data and used a negative binomial distribution (10% FDR) to calculate significantly biased genes between the two alleles, where genes are defined by merging isoforms (from RefSeq). We used the edgeR software package in R for calculating the P values.
Allelic SNPs
We estimated if a SNP is allele-biased on different types of readouts. In particular, we used ChIP-seq, DHS, and CTCF data sets independently to obtain readouts of each SNP between the two alleles. We then used a binomial statistic (with an expectation P = 0.5) to identify significantly biased SNPs for a given data set. FDR was based on 1,000 random permutations.
Differential methylation among alleles (DMRs)
Bisulfite sequencing reads were mapped using Novoalign methylation aligner to an H1 variant masked hg18 reference genome. Duplicated and poorly mapped reads were removed, and the reads that contain SNPs were retained for downstream analyses. Reads were then assigned to either the p1 or the p2 allele on the basis of the SNPs present in each read. During this assignment, certain SNPs could not be resolved between the two alleles because of considerations of bisulfite conversion. Specifically, when a SNP is C/T (or listed as A/G on the reverse strand), the conversion of methyl-C to T by bisulfite will make it impossible to distinguish whether a given read is a methylated cytosine from one allele or a thymidine from the other allele. In these cases, these SNPs were excluded from distinguishing from which allele a given read was derived. After resolving into each allele, CpGs were called and nearby CpG were merged (within 100 bp). Of note, in instances where a SNP contains a cytosine, it would be impossible to distinguish whether a difference between two alleles is due to the polymorphism or due to the change in methylation. As such, any position in the genome with a SNP was excluded from our calculation of the percentage methylation over a given window. We called ASM in each of these CpGs using Fisher’s exact test with 10% FDR after multiple testing correction as a threshold for significance. We randomly shuffled the methylation and unmethylation values for a given haplotype (for a CpG) and used these random estimates to obtain FDR.
Allelic enhancers
To study allele bias at enhancers, we first calculated the combined coverage of whole genome sequencing data and bisulfite sequencing (without regard for methylation status). Any enhancer where one of the two alleles contained less than 35% of the total allele resolved reads at the enhancer was excluded as having an inherent bias in mapping between the two alleles. To systematically study allelic enhancers, we combined several enhancer marks to obtain a combined acetylation bam file. This combined bam file gives us the required coverage in an allelic context to perform an in-depth analyses. In particular, we combined data from H4K8ac, H4K91ac, H2BK120ac, H3K18ac, H3K23ac, H3K27ac, H3K4ac, H2AK5ac and H3K9ac marks. Using this combined bam file, we examined allelic SNPs described as above. For evaluating allelic enhancers, we obtained readout for enhancers defined in ref. 13 (± 2.5 kb from enhancer peaks) between the two alleles. Then we used binomial to obtain significance at an FDR of 10%, as evaluated by the random permutation analyses (1,000 permutations). The same analysis was used to call allele-biased enhancers based on DHS data. For the analysis of allele bias in DNA methylation at enhancers, we considered any enhancer as having allele-biased DNA methylation if at least one ASM bin overlapped with the enhancer. If more than one bin of ASM overlapped an enhancer, we checked to see whether the patterns of ASM were concurrent between all bins. If there were divergent patterns between ASM bins at an enhancer, these enhancers were excluded.
Distance of allelic enhancers to allelic genes
We compared the distance between allele-biased enhancers, as identified by histone acetylation levels with randomly selected enhancers, to test the hypothesis that if allele-biased enhancers regulate allele-biased genes, they should generally be closer to allele-biased genes than should randomly chosen enhancers (Fig. 5b). This analysis was complicated by the fact that the rates of heterozygous SNPs near allele-biased genes are higher than for non-allele-biased genes in the genome (Extended Data Fig. 6f). This creates a situation of possible ascertainment bias, owing to the fact that enhancers near allele-biased genes will therefore tend to have slightly higher allele-resolved read coverage as compared with randomly chosen enhancers throughout the genome. To account for this, when comparing the distance of allele-biased enhancers to allele-biased genes with randomly chosen enhancers, we selected random enhancers to match the coverage profile of allele-biased enhancers. This was accomplished by binning all enhancers into increments of 50 sequencing reads, from 0 to 49, 50 to 99, etc, up to 1,700 reads. For each identified allele-biased enhancer, we selected 100 random enhancers from the same coverage bin. This limits the effects of local variation in heterozygosity rates throughout the genome on the likelihood of identifying allele-biased enhancers near allele-biased genes. As such, the results in Fig. 5b are probably not due to the possibility of having greater statistical power for calling allele-biased enhancers near allele-biased genes (because of greater heterozygosity rates and higher numbers of allele-resolved reads).
Enhancers, gene expression levels, lineage-specific genes, housekeeping genes and imprinting genes
The enhancer regions were defined as previously described44. Briefly, enhancer chromatin signatures were trained for p300 binding sites in H1 ES cells using RFECS algorithm based on H3K4me1, H3K4me3 and H3K27ac signals at 100-bp bin size. Next, these modification signals in all cell lines were tested to predict enhancers. The predicted enhancers that overlap with H3K4me3 peaks or within 2.5 kb of the transcription start site were removed. Enhancers were merged from all cell types if they are located close to each other (<2 kb) by taking the midpoint at the centre of the new enhancer.
For the gene list, gene expression levels, housekeeping genes and lineage-specific genes we used the same data set as described in ref. 13. For imprinting genes, we obtained known imprinted genes downloaded from publicly available imprinting gene database (http://www.geneimprint.com/).
Linking between allelically expressed genes and allele-biased promoter activities
To investigate how many allele-biased gene promoter activities are consistent with allelic gene expression levels, first we selected allelic genes that contain at least one allelic SNP in their promoter regions (1.5 kb upstream and downstream from transcription start site). We only considered allelic SNPs defined by DHS, H3K4me3, histone acetylation, combined H3K9me3 and H3K27me3, and DNA methylation because the signatures of those chromatin marks at the promoter regions are well defined. If promoters are marked by allelic SNPs from H3K9me3/H3K27me3 or DNA methylation and the allelic gene expression levels are consistent with the allele-biased promoter activities, the genes can be explained by allelic repressive marks. If promoters are marked by allelic SNP from histone acetylations, H3K4me3, and DHS, and allelic gene expression levels are consistent with the allele-biased promoter activities, the genes can be explained by allelic active marks.
Identification of enhancer–promoter interactions
To investigate the linking between allelic genes and allelic enhancers we first defined enhancer-promoter interactions using Hi-C interaction frequency data. Hi-C interaction frequencies were calculated in terms of 5-kb windows and normalized using HiCNorm. After that, we considered all pairs of promoters and enhancers in each chromosome. Promoter regions were fixed as ± 5 kb surrounding transcriptional start sites and enhancer regions were defined by using different window sizes as: 5 kb, 10 kb, 20 kb, 30 kb, 40 kb, 50 kb, 75 kb, 100 kb, 300 kb and 500 kb surrounding the centre of each enhancer (Extended Data Fig. 7). The interaction frequencies between a promoter and an enhancer at a certain window size were calculated as (interaction frequency / window size of an enhancer) × 5 kb. Final interaction scores were defined as summation of interaction frequencies between promoter and enhancer with multiple window sizes. To calculate significance of each enhancer–promoter interaction, we generated a random interaction frequency score by randomly permutated interaction frequencies between the promoter and enhancer in each window size. The distribution of random interaction frequency scores was fit to Weibull distribution and then P values of the interaction frequency in each enhancer–promoter pair were calculated. At a given P value cutoff, we defined enhancer–promoter interactions. At a P value cutoff of 1 × 10−3, more than 80% of interactions are reproducible between two biological replicates (Extended Data Fig. 7b). By taking this P value cutoff, we defined 339,761, 354,529, 319,169, 158,453, 250,495, and 210,010 enhancer–promoter interactions for ES, ME, MS, NP, TB and IMR90 cell lines between 103,982 enhancers and 18,532 promoter regions. These enhancer–promoter interaction numbers can be changed according to cutoff P values.
Comparison of enhancer–promoter interaction with other experiments
To validate predicted enhancer–promoter interactions we compared the interaction frequency scores to 5C scores and DNaseI quantitative trait loci (dsQTL) information. 5C is a chromosome conformation capture (3C)-based approach to measure the interactions of all versus all targeted regions. For the H1 ES cell lines, we downloaded previously generated 5C data23 and compared this to our interaction frequency between enhancers and promoters. We observe very strong correlative patterns between 5C and our interaction frequency scores (Extended Data Fig. 1b). We can also observe that interacting pairs tend to show higher 5C scores compared to non-interacting pairs.
We also compared interaction frequency scores to dsQTL relationships. dsQTL data provide functional relationships between DHS and their target promoters based on QTL information45. We calculated enhancer–promoter interactions scores again for all pairs of DHS and promoter regions based on dsQTL data. Interactions defined by dsQTL were considered as target relationships, otherwise off-target relationships. According to interaction distance, enhancer–promoter interaction scores were calculated between target and off-target gene relationships. We observe that target gene relationships tend to have higher interaction frequency scores (Extended Data Fig. 7d).
Pearson correlation coefficient between allele-biased gene-enhancer pairs
We calculated Pearson correlation coefficients between allele-biased gene-enhancer pairs. First we generate 1 by 10 vectors for each allelic gene and allelic enhancer using the data from H1 human ES cells and the four H1-derived lineages. For each lineage, we assigned log2 (p2 allele read counts / p1 allele read counts) and log2 (p1 allele read counts / p2 allele read counts) values as allelic bias information. After constructing two 1 by 10 vectors for both allelic gene and allelic enhancer, we calculated the Pearson correlation coefficient between them.
Identification of allelic Hi-C interactions
We investigated allelic Hi-C interactions for allelic gene-enhancer pairs. We considered allelic interactions between 10-kb surrounding regions for both the transcription start site and enhancer, respectively. Many of allelic gene-enhancer pairs do not have any allelic interactions, but allelic gene-enhancer pairs show concordance if they are connected by allelic Hi-C interactions.
4C-seq experiments and data analysis
4C seq was performed essentially as described previously36. Six bait regions were chosen at allele-biased enhancer elements containing SNPs that would allow for performance of allele specific 4C-seq, as has been previously described31. Primers were designed such that the first read of a paired-end read would sequence the primer sequence derived from the bait region and read into the target region of interest. The second read in the pair would read a portion of the bait region containing the SNP of interest (see Extended Data Fig. 8 for a diagram of the experimental strategy). The primers were designed to include the Illumina adaptor sequences necessary for sequencing as well as the presence of barcodes derived from Illumina’s TruSeq adaptors that allowed for multiplexing of 4C-seq reactions. We used two 4 base cutters, NlaIII and DpnII, for the first or second restriction enzyme digestion, depending on the locus in question (See Supplementary Table 4). 4C-seq templates were prepared as previously described36. 16 PCR reactions using 200 ng of 4C template were performed for 30 cycles for each bait region and pooled together. The PCR reactions underwent a final purification step using AMPure beads (Beckman-Coulter) according to the manufacturer’s instructions (using a bead-to-sample ratio of 1.8). The concentrations of each 4C library were calculated using the KAPA qPCR system using a standard curve. The libraries were then combined and spiked in with other non-4C sequencing libraries for sequencing on the Illumina Hi-Seq 2500 machine.
Sequence reads were processed as follows. For each read, the first and second sequencing reads were checked to identify the presence of the primer sequences and any expected portion of the bait region. Any sequence with greater than 20% mismatches to the expected bait region was discarded. The reads were trimmed such that each read was represented as a 36-mer, with 20 bp derived from the bait region and the subsequent 16 bp, presumably containing the target region of interest. Based on the SNP identified in the second sequencing read derived from the bait region, each of these files were split into allele specific 4C-seq FASTQ files for further analysis.
4C-seq data was mapped to a version of the hg18 genome with known SNPs in the H1 genome masked to N, similar to other the strategy of mapping other sequence read data sets performed in this study. Custom indexes for this H1-masked hg18 genome were built using the 4cseqpipe “-build_re_db” command. The reads were mapped using the 4Cseqpipe software “-map” command to custom built indexes. Normalized contact intensities were derived using the 4seqpipe “-nearcis” command for a 1-Mb region upstream and downstream of the bait locus. We then took the normalized fragment level interaction frequency tables and removed any fragments where a SNP either could create or disrupt a potential restriction enzyme site between the two alleles. In addition, given the short sequencing read length, any fragment with an insertion or deletion mapping within 16 bp of the fragment end was removed. These final filtered sets of normalized fragment level interaction frequencies were then processed using a sliding window approach with the window size of 5 kb and step size of 1 kb using the average fragment interaction frequency over the 5-kb window. These sliding interaction frequency files were then quantile normalized across all replicates in order for comparison between experiments using the “normalize.quantiles.robust” function (with use.median = TRUE) in the “preprocessCore” library in R. For display purposes, the average of two replicates was converted to bedGraph format and displayed in the UCSC genome browser.
To identify regions that showed specific interactions with the bait region controlling for the genomic distance between loci, we developed a LOWESS regression model. We pooled the sliding window interaction frequency files from each of the 4C-seq replicates and performed LOWESS regression in R with the function “lowess” (with f = 0.01) on the log10 transformed interaction frequencies controlling for the distance between the bait and potential interaction locus. We considered any region as showing ‘specific’ interactions if it showed an increase in interaction frequency greater than 2.5-fold over expected given the distance between the bait and target loci. These were considered to be the bait interacting regions.
To test for any allelic bias in 4C-seq interaction frequencies, the average normalized fragment level interaction frequency was calculated for each allele of each replicate over the bait interacting regions nearest to the transcription start site (TSS) of the putative target gene. A t-test was performed using these average values (n = 2 for each allele) to determine statistical significance.
The primers used for each 4C-seq experiment are listed in Supplementary Table 5 (please see Supplementary Table 4 for information regarding the bait regions for each experiment). In Supplementary Table 5, the Illumina barcode adaptors are shown in red, with the region matching the bait region shown in blue. There is also an additional variable region in the Illumina TruSeq adaptors that has been incorporated and shown in green. The phosphorothioate bond is indicated by an asterisk.
References
Smallwood, A. & Ren, B. Genome organization and long-range regulation of gene expression by enhancers. Curr. Opin. Cell Biol. 25, 387–394 (2013)
Phillips, J. E. & Corces, V. G. CTCF: master weaver of the genome. Cell 137, 1194–1211 (2009)
Lettice, L. A. et al. Disruption of a long-range cis-acting regulator for Shh causes preaxial polydactyly. Proc. Natl Acad. Sci. USA 99, 7548–7553 (2002)
Gorkin, D. U., Leung, D. & Ren, B. The 3D genome in transcriptional regulation and pluripotency. Cell Stem Cell 14, 762–775 (2014)
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012)
Phillips-Cremins, J. E. et al. Architectural protein subclasses shape 3D organization of genomes during lineage commitment. Cell 153, 1281–1295 (2013)
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012)
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012)
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014)
Ryba, T. et al. Evolutionarily conserved replication timing profiles predict long-range chromatin interactions and distinguish closely related cell types. Genome Res. 20, 761–770 (2010)
Peric-Hupkes, D. et al. Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol. Cell 38, 603–613 (2010)
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013)
Maurano, M. T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012)
Selvaraj, S., Dixon, J. R., Bansal, V. & Ren, B. Whole-genome haplotype reconstruction using proximity-ligation and shotgun sequencing. Nature Biotechnol. 31, 1111–1118 (2013)
Hu, M. et al. HiCNorm: removing biases in Hi-C data via Poisson regression. Bioinformatics 28, 3131–3133 (2012)
Hawkins, R. D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6, 479–491 (2010)
Brown, K. E. et al. Association of transcriptionally silent genes with Ikaros complexes at centromeric heterochromatin. Cell 91, 845–854 (1997)
Kosak, S. T. et al. Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 296, 158–162 (2002)
Holwerda, S. & de Laat, W. Chromatin loops, gene positioning, and gene expression. Front. Genet. 3, 217 (2012)
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature Genet. 39, 311–318 (2007)
Heintzman, N. D. et al. Histone modifications at human enhancers reflect global cell-type-specific gene expression. Nature 459, 108–112 (2009)
Sanyal, A., Lajoie, B. R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012)
Heinz, S. et al. Effect of natural genetic variation on enhancer selection and function. Nature 503, 487–492 (2013)
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013)
Kasowski, M. et al. Extensive variation in chromatin states across humans. Science 342, 750–752 (2013)
Kilpinen, H. et al. Coordinated effects of sequence variation on DNA binding, chromatin structure, and transcription. Science 342, 744–747 (2013)
Kuleshov, V. et al. Whole-genome haplotyping using long reads and statistical methods. Nature Biotechnol. 32, 261–266 (2014)
Xie, W. et al. Base-resolution analyses of sequence and parent-of-origin dependent DNA methylation in the mouse genome. Cell 148, 816–831 (2012)
Splinter, E. et al. The inactive X chromosome adopts a unique three-dimensional conformation that is dependent on Xist RNA. Genes Dev. 25, 1371–1383 (2011)
Holwerda, S. J. et al. Allelic exclusion of the immunoglobulin heavy chain locus is independent of its nuclear localization in mature B cells. Nucleic Acids Res. 41, 6905–6916 (2013)
de Wit, E. et al. The pluripotent genome in three dimensions is shaped around pluripotency factors. Nature 501, 227–231 (2013)
Gimelbrant, A., Hutchinson, J. N., Thompson, B. R. & Chess, A. Widespread monoallelic expression on human autosomes. Science 318, 1136–1140 (2007)
Hon, G. C. et al. Epigenetic memory at embryonic enhancers identified in DNA methylation maps from adult mouse tissues. Nature Genet. 45, 1198–1206 (2013)
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012)
van de Werken, H. J. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nature Methods 9, 969–972 (2012)
Reddy, K. L., Zullo, J. M., Bertolino, E. & Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 452, 243–247 (2008)
Finlan, L. E. et al. Recruitment to the nuclear periphery can alter expression of genes in human cells. PLoS Genet. 4, e1000039 (2008)
Kumaran, R. I. & Spector, D. L. A genetic locus targeted to the nuclear periphery in living cells maintains its transcriptional competence. J. Cell Biol. 180, 51–65 (2008)
Kim, T. H. et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell 128, 1231–1245 (2007)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
Bansal, V. & Bafna, V. HapCUT: an efficient and accurate algorithm for the haplotype assembly problem. Bioinformatics 24, i153–i159 (2008)
Browning, B. L. & Browning, S. R. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 84, 210–223 (2009)
Rajagopal, N. et al. RFECS: a random-forest based algorithm for enhancer identification from chromatin state. PLOS Comput. Biol. 9, e1002968 (2013)
Degner, J. F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012)
Acknowledgements
We thank the members of the Ren laboratory for support and suggestions throughout the course of this work. This work is funded in part by the Ludwig Institute for Cancer Research, the NIH Roadmap Epigenome Project (U01 ES017166 and R01 ES024984), and the California Institute of Regenerative Medicine (RN2-00905). Y.D. is supported by a postdoctoral fellowship from the Human Frontier Science Program (LT000576/2014-L).
Author information
Authors and Affiliations
Contributions
J.R.D., I.J., S.S., and B.R. conceived the study. J.R.D. performed Hi-C and 4C experiments with assistance from A.K. J.R.D., I.J., and S.S. performed the data analysis. A.Y.L. performed ChIP-seq experiments for CTCF with assistance from Z.Y. J.E.A.-B. performed ES cell culture, differentiation and sample collection. N.R. and W.X. provided expertise and assistance in data analysis. Y.S., Y.D., V.V.L. J.L., H.Z., J.R.E., and J.T. provided insights and support for the design and execution of the project. J.R.D. and I.J. prepared the manuscript with assistance from S.S. and B.R.
Corresponding author
Ethics declarations
Competing interests
J.R.D., S.S. and B.R. are co-founders of Arima Genomics.
Extended data figures and tables
Extended Data Figure 1 Reproducibility of Hi-C data.
a, Scatter plots showing the correlation of Hi-C interaction frequencies between two biological replicates for H1 ES cells and H1-derived lineages. The Pearson correlation coefficient between replicates is shown in each plot. b, Heat maps showing interaction frequencies of Hi-C and 5C data over the chromosome 7 ENCODE loci. Pearson correlation coefficients between heat maps are shown together. The correlation coefficients between Hi-C data and 5C data (PCC 0.72, 0.71, 0.58, 0.57) are similar to the correlation coefficient between two biological replicates of 5C data (PCC 0.68). c, Bar plots showing the fraction of topological domain boundaries that are reproducible between biological replicates over H1 and H1-derived lineages (x axis). d, Scatter plot showing the PC1 values derived from compartment A/B analysis between biological replicates. PC1 values are used to determine the A and B compartments in each cell type. The Pearson correlation coefficient is shown in each graph.
Extended Data Figure 2 A/B compartments changes are concordant with topological domain boundaries.
a, Genome browser image of the A/B compartments determined using the previously described 1-Mb bin algorithm (1-Mb track) compared with the 40-kb sliding window approach used in our analysis (40-kb sliding window track). b, Pie-charts demonstrating the fraction of the genome in the A (blue) or B (yellow) compartments in each of the six lineages studied. Shown in black are regions with a PC1 of zero, often corresponding to centromeric and telomeric regions of the chromosomes. c, Percentage of the genome that changes A/B compartment upon differentiation of ES cells into each of the five differentiated lineages. d, Cumulative density plot of the distance between topological domain boundaries and transition points between the A and B compartments. The red line represents the observed distances and the grey line represents distances for randomly generated topological domain boundaries. Domain boundaries are closer to A/B compartment transitions when compared with random (P value 2.2 × 10−16, Wilcoxon rank sum test). e, K-means clustering of PC1 values in human ES cells and differentiated lineages surrounding topological domain boundaries. Similar to Fig. 1c, domain boundaries correspond to the transition points between the A/B compartments, and changes in A/B compartments that occur during differentiation tend occur at domain boundaries. Regions that stay as A or B compartment are termed stable A or stable B. Regions that stay as A/B compartment switching are labelled as stable switch. Regions where the boundary becomes a new switching point for the A/B compartment are labelled new switch. Regions that previously were A/B compartment switching but are no longer after differentiation are labelled switch loss. Regions that entirely switch from A to B or vice versa are labelled as switch A/B.
Extended Data Figure 3 A/B compartment changes and gene expression.
a, Fraction of lineage-restricted genes in lineage-restricted compartment A regions and other remaining regions. If only one or two cell lines are assigned as compartment A across the six lineages, the region is defined as a lineage-restricted compartment A region. For all six lineages, lineage-restricted genes tend to be enriched in lineage-restricted compartment A regions compared to other genomic regions (P values <0.05, Fisher’s exact test). b, Empirical cumulative density plot of covariance values between gene expression and PC1 score across the 6 lineages analysed. Shown in red are the observed covariance values, while in grey are covariance values calculated after randomly shuffling the vector of gene expression values for each lineage. The slight shift of the red curve to the right indicates that the observed data has a subset of genes with higher covariance values than would be expected at random, indicating that a subset of genes have concordant gene expression and PC1 values. c, Histogram of P values for the covariance between gene expression and PC1 values for each gene. To calculate the P value, a random background distribution of covariance values was generated by calculating the covariance between the PC1 values and a randomly shuffling of the vector of gene expression values for each gene. This shuffling was performed 1,000 times. The actual observed covariance can then be assigned a rank based P value given the random background distribution for that gene. The plot shows that a subset of genes is enriched for having low P values, consistent with the idea that a subset of genes shows concordant gene expression and compartment status. d, Pie charts showing the fraction of domains that are identified as having a concerted increase (yellow) or decrease (blue) in intra-domain interaction frequency between H1 human ES cells and the five lineages analysed. e, Relationship between A/B compartment and intra-domain interactions. 2 × 2 tables for each lineage (or for all lineages combined, labelled as total) for domains that show a concerted increase or decrease in interaction frequency and whether they show a change from A to B or B to A compartments. Domains are considered to undergo a compartment change if >80% of individual bins within the domain change compartment. Odds ratio (OR) for each lineage and the total are listed, as are P values for the association (Fisher’s exact test). f, Box plots showing the average difference of intra-domain interaction frequencies between H1 human ES cells and the five lineages analysed. Regions that change from compartment B to A (blue) tend to show increased intra-domain interaction frequencies compared to regions that change from compartment A to B (orange). P values are less than 2.2 × 10−16 for all lineages (KS test). g, Box plots of the fold-change in gene expression of genes located in domains that have a significant increase (+), decrease (−), or no change (0) in intra-domain interaction frequency between ES cells and each of the differentiated lineages specified. The fold-change in expression is the log2 of the expression of a gene in the differentiated cells over ES cells (P values from Wilcoxon rank-sum test). h, Box plots showing Z-scores of intra-domain interactions between lineage-restricted genes and other remaining genes. The average intra-domain interaction frequency was calculated for each domain in six lineages analysed and converted to a Z-score. The Z-score of each gene was assigned by the Z-score of corresponding domain that includes the gene. Lineage-restricted genes tend to reside in domains with higher Z-scores compared to other remaining genes. The P values were less than 1 × 10−4 from the KS test.
Extended Data Figure 4 Random Forest model to predict Hi-C interaction changes.
a, Comparison of the classification accuracy between tenfold cross validation and the out-of-back (OOB) error rates. The two methods show similar classification accuracies at each vote frequency threshold. b, Heat map showing the difference of normalized interaction frequencies between H1 and MS cells. The boxes indicate the regions with relatively strong higher interaction frequencies in H1 ES cells. H3K4me1 and CTCF ChIP-seq signals are also shown together. H3K4me1 ChIP-seq signals are highly correlated with changes of interaction frequency. c, Similar to Fig. 2e, ranked Gini index of different chromatin features of the Random Forest model when using randomized data. The red line represents the centre of the Gini index. d, Box plots demonstrating the difference in the number of enhancers in each 40-kb bin that undergoes a gain of interactions (GOI) or loss of interactions (LOI) upon differentiation. We observe that regions that are involved gain of interactions tend to contain more enhancers in differentiated lineages compared to H1 cells. We only considered regions containing more than 10 enhancers in total for H1 and differentiated cell lines (P value < 2.2 × 10−16, KS test). e, Histogram showing the fraction of interactions classified as GOI (orange) or LOI (blue) according to the difference of the number of lineage-specific enhancers between differentiated lineages and H1 ES cells. The regions with more lineage-specific enhancers in differentiated lineages are enriched by gain of interactions.
Extended Data Figure 5 Allele-specific chromatin structure.
a, Validation of haplotypes by (i) RNA-sequencing (i) and whole-genome sequencing (WGS) (ii). Shown in dark blue is the percentage of reads connecting variants in the same predicted haplotype, while in light blue is the percentage of reads connecting variants predicted to be on different haplotypes. b, Inset labelled with an asterisk is from Fig. 3b showing DHS sequencing reads over a SNP upstream from the SNRPN gene, demonstrating how different chromatin features can be assigned to a given haplotype. c, Scatter plots showing the correlation coefficient of PC1 values obtained from compartment A/B analysis between the two biological replicates for each allele. Despite the reduction in reads when Hi-C data are split into two alleles, the PC1 scores were highly reproducible between replicates. d, Shown is a genome browser image of PC1 values in chromosome 2 for the p1 allele, p2 allele, and for all Hi-C reads without resolving the two alleles. PC1 scores are highly consistent, suggesting that homologous chromosomes fold in highly similar patterns. e, Scatter plot of PC1 values between the p1 and p2 alleles in H1 and H1-derived lineages. The Pearson correlation coefficient value is 0.96. f, Fraction of the genome that shows changes in A/B compartment status across alleles. For ES, ME, MS, NP and TB cells, 1.3%, 0.6%, 1.9%, 2.3% and 1.6% of total genomic regions shows allelic compartment A/B patterns, respectively. g, Percentage of allele-biased (purple) or non-allele-biased (grey) genes that have different A/B compartment status in each lineage. Only in ES cells is there a significant association between allele-biased genes and regions with variable A/B compartment between alleles (Fisher’s exact test). h, Similar to g, but showing the association between imprinted genes and changes in A/B compartment between alleles. No lineage shows a significant association.
Extended Data Figure 6 Domain-wide structural changes and allele-biased genes.
a, Scatterplot showing domain ‘intactness’ between the p1 and p2 alleles. Domain intactness is defined as the log2 ratio of the total number of intra-domain interactions versus total number of inter-domain interactions for each topological domain. The highly correlated domain intactness scores between the p1 and p2 alleles support the similar topological domain patterns between two homologous chromosomes. b, Pie charts showing the fraction of domains that are identified as having a concerted p1 allele specific increase (blue) or p2 allele specific increase (yellow) in interaction frequency. Grey in the pie charts indicates the fraction of domains that do not show allele specific patterns compared to the random model (P value cutoff is 0.001). c, Heat map showing K-means clustering (k = 12) of gene expression levels of allele-biased genes across each of the five H1 lineages. The expression levels are shown as the fold-change of expression in each lineage relative to the average expression level across each of the five lineages. Allele-biased genes consist of both cell-type specific and constitutively expressed genes. d, Fraction of housekeeping genes and lineage-restricted genes that show allele-biased expression. There is no statistically significant enrichment between allele-biased genes (orange) and non-allele-biased genes (blue) among housekeeping or lineage-restricted genes. e, Empirical cumulative density plot of the distance between each allele-biased gene and the nearest allele-biased gene (purple) as compared with randomly chosen genes (yellow). The difference from an allele-biased gene to the nearest allele-biased gene is less than what would be expected at random (P = 0.0482, Wilcoxon rank sum test), however, the difference is subtle, indicating that most allele-biased expression does not occur in clusters. f, Rate of heterozygous SNPs near both allele-biased (gold) and non-allele-biased (black) genes. See Supplementary Information for further details.
Extended Data Figure 7 Identification of enhancer–promoter interactions.
a, Shown is the Hi-C interaction frequency between the IDO1 gene promoter regions and ± 1 Mb surrounding regions. Each entry in the heat map of Hi-C inter. freq. indicates Hi-C interaction frequency between the promoter and the surrounding regions. Each row indicates the Hi-C interaction frequencies for a given window size. The heat map of random permutation was generated by randomizing each row in Hi-C interaction frequency. The sum (Hi-C) and sum (random) indicate summation of Hi-C interaction frequencies for each 5-kb window. Predicted enhancers, H3K27ac, RNA-seq, and RefSeq gene information are shown together. b, Box plots showing the reproducibility of predicted enhancer–promoter interactions between two biological replicates for each chromosome with different P value cutoffs. If the P value is less than 0.001, the reproducibility between replicates is over 80%. c, Distribution of 5C signals between interacting pairs (interaction) and non-interacting pairs (others) defined by Hi-C interaction frequency score with different P value cutoffs. Interaction pairs defined by Hi-C interactions are also strongly enriched by 5C signals at both P value cutoffs (n = 11,461 for 1 × 10−3 and n = 1,841 for 1 × 10−4). d, Relationship between Hi-C interaction frequency scores and dsQTL target-gene pairs according to distance between gene and its target DHS regions. Target-gene relationships tend to show higher Hi-C interaction frequency scores compared to off-target-gene relationships.
Extended Data Figure 8 4C-seq between allelic enhancers and allelic genes.
a, Diagram of experimental design for 4C-seq and allelic 4C-seq. The orange region depicts the 4C bait locus, and the green region is the interacting target region. Primers containing the Illumina adaptor sequences and a bait-specific sequence are used for inverse PCR of the target region. Barcodes based on the Illumina TruSeq adaptors are incorporated into the primer sequences to allow for multiplexing. The second primer will read a sequence from the bait region with a SNP that determines the allele from with the bait was derived. b, Pearson correlation coefficients between replicates for each of the loci tested. Also shown is the Pearson correlation coefficient between replicates after log-transformation of the interaction frequency. c, Scatter plot of LOWESS regression of 4C-seq data. The x axis shows the genomic distance between the bait region and the putative target region. The y axis is the log10 of the quantile normalized interaction frequencies. LOWESS was performed to generate an expected interaction frequency at each genomic distance (green line). A cut off of 2.5-fold over expected (shown in the red dashed line) is used to determine if a region shows specific interactions, so-called bait interacting regions (BIRs). d, Normalized 4C-seq interaction frequencies surrounding a bait region located in an allelic enhancer near the FAM65B gene. The location of the bait is labelled as 4C bait. Regions with significant interactions according to the LOWESS regression model are labelled as black lines in the track marked bait interacting regions. Shown to the right is the level of mRNA-seq data for each allele of the FAM65B gene, the level of histone acetylation at the allelic enhancer bait region. Significance for mRNA-seq data was calculated using the edgeR software package in R. Acetylation P values were calculated using a binomial test. e, Similar to d, but for a 4C seq bait located in the MAN1C1 gene. f, Similar to d, but for an allelic enhancer located in the PXK gene. g, Similar to d, but for an enhancer located in near the GCLM gene. Of note, this allele-biased enhancer forms no specific contacts with any allelic genes. h, Similar to d, but for an enhancer located near the MT2A gene. There are no specific interactions between the allelic enhancer and the MT2A gene. There are specific interactions between the allelic enhancer and the MT1H and MT1G genes. However, neither gene has an exonic SNP and therefore we cannot determine if these genes have allele-biased expression.
Extended Data Figure 9 4C-seq interacting regions from allelic enhancers.
a, Allelic Hi-C interaction reads shown for allelic gene-enhancer pairs defined using either allelic histone acetylation, DHS or DNA methylation. Odds ratios (OR) and P values (Fisher’s exact test) are shown. For enhancers defined by histone acetylation and the pooled set of enhancers, a statistically significant association between allele-biased Hi-C reads and allele-biased enhancer activity is observed. b, Normalized 4C-seq interaction frequencies surrounding a bait region located in an allelic enhancer near the HAPLN1 gene. The blue line shows the interaction frequency for the p1 allele and the red line shows interaction frequencies for the p2 allele. The shaded regions represent 95% confidence intervals for the interaction frequency. Shown to the right are the allele-specific normalized 4C interaction frequencies for each allele. 4C-seq interaction frequencies for each allele were computed over the significant bait interacting regions nearest to the target gene TSS. Significance testing for allelic 4C-seq data was performed by t-test (n = 2 for each allele). Black bars below the plot indicate regions identified as bait-interacting regions (BIRs). Of note, the panel to the right is the same as that found in Fig. 5f. c, Similar to b, but for an allelic enhancer located in the MAN1C1 gene. d, Similar to b, but for a 4C seq bait located in the FAM65B gene. e, Similar to b, but for an enhancer located in near the PXK gene.
Supplementary information
Supplementary Data
This file contains Supplementary Table 1. (XLSX 47 kb)
Supplementary Data
This file contains Supplementary Table 2. (XLSX 47 kb)
Supplementary Data
This file contains Supplementary Table 3. (XLSX 43 kb)
Supplementary Data
This file contains Supplementary Table 4. (XLSX 44 kb)
Supplementary Data
This file contains Supplementary Table 5. (XLSX 65 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Dixon, J., Jung, I., Selvaraj, S. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015). https://doi.org/10.1038/nature14222
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14222
This article is cited by
-
Cell-type differential targeting of SETDB1 prevents aberrant CTCF binding, chromatin looping, and cis-regulatory interactions
Nature Communications (2024)
-
3D Enhancer–promoter networks provide predictive features for gene expression and coregulation in early embryonic lineages
Nature Structural & Molecular Biology (2024)
-
DiffDomain enables identification of structurally reorganized topologically associating domains
Nature Communications (2024)
-
Baf-mediated transcriptional regulation of teashirt is essential for the development of neural progenitor cell lineages
Experimental & Molecular Medicine (2024)
-
Strong interactions between highly dynamic lamina-associated domains and the nuclear envelope stabilize the 3D architecture of Drosophila interphase chromatin
Epigenetics & Chromatin (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.