Abstract

Allelic differences between the two homologous chromosomes can affect the propensity of inheritance in humans; however, the extent of such differences in the human genome has yet to be fully explored. Here we delineate allelic chromatin modifications and transcriptomes among a broad set of human tissues, enabled by a chromosome-spanning haplotype reconstruction strategy1. The resulting large collection of haplotype-resolved epigenomic maps reveals extensive allelic biases in both chromatin state and transcription, which show considerable variation across tissues and between individuals, and allow us to investigate cis-regulatory relationships between genes and their control sequences. Analyses of histone modification maps also uncover intriguing characteristics of cis-regulatory elements and tissue-restricted activities of repetitive elements. The rich data sets described here will enhance our understanding of the mechanisms by which cis-regulatory elements control gene expression programs.
Similar content being viewed by others
Main
We performed chromatin immunoprecipitation followed by sequencing (ChIP-seq) to generate extensive data sets profiling 6 histone modifications across 16 human tissue types from four individual donors (181 data sets). In combination with previously published data sets2,3, we conducted in-depth analyses across 28 cell/tissue types, covering a wide spectrum of developmental states, including embryonic stem cells, early embryonic lineages and somatic primary tissue types representing all three germ layers (Fig. 1a) (protocols received approval from IRB/ESCRO and Mid-American Transplant Services, and research consent was obtained from families). The modifications demarcate active promoters (histone H3 lysine 4 trimethylation (H3K4me3) and H3 lysine 27 acetylation (H3K27ac)), active enhancers (H3 lysine 4 monomethylation (H3K4me1) and H3K27ac), transcribed gene bodies (H3 lysine 36 trimethylation (H3K36me3)) and silenced regions (H3K27 or H3 lysine 9 trimethylation (H3K27me3 and H3K9me3, respectively))4,5. We systematically identified cis-regulatory elements by employing a random-forest-based algorithm (RFECS)2,6, predicting a total of 292,495 enhancers (consisting of 175,912 strong enhancers with high H3K27ac enrichment) across representative samples of all 28 tissues types (Supplementary Table 1). We additionally identified 24,462 highly active promoters with strong H3K4me3 enrichment (see Supplementary Table 2). Subsequently, we defined tissue-restricted promoters (n = 10,396) and enhancers (n = 115,222) (Extended Data Fig. 1a). Consistent with previous studies7,8,9, enhancers appear more tissue-restricted than promoters and cluster along developmental lineages (Extended Data Fig. 1a, b). Moreover, tissue-restricted enhancers were enriched for putative binding motifs of particular transcription factors known to be important in maintaining the cell/tissue type’s identity and function10,11,12,13,14,15 (Extended Data Fig. 2).
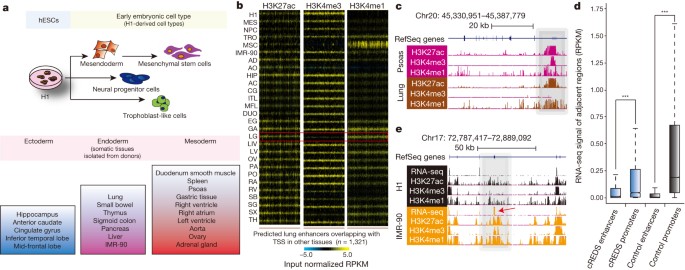
a, Schematic of the cell/tissue types profiled and their progression along developmental lineages. Samples include embryonic stem cells (H1), early embryonic lineages (mesendoderm cells (MES), neural progenitor cells (NPC), trophoblast-like cells (TRO) and mesenchymal stem cells (MSC)) and somatic primary tissues, representative of all three germ layers (ectoderm: hippocampus (HIP), anterior caudate (AC), cingulate gyrus (CG), inferior temporal lobe (ITL) and mid-frontal lobe (MFL); endoderm: lung (LG), small bowel (SB), thymus (TH), sigmoid colon (SG), pancreas (PA), liver (LIV) and IMR-90 fibroblasts; mesoderm: duodenum smooth muscle (DUO), spleen (SX), psoas (PO), gastric tissue (GA), right heart ventricle (RV), right heart atrium (RA), left heart ventricle (LV), aorta (AO), ovary (OV) and adrenal gland (AD)). b, Heat maps show H3K27ac, H3K4me3 and H3K4me1 enrichment (input normalized reads per kilobase per million mapped reads (RPKM)) at predicted lung enhancers (n = 1,321), which are defined as promoters in other tissues, across all 28 samples. The red box highlights the signatures in lung. c, A UCSC genome browser snapshot of a region on chromosome 20, showing the chromatin states of a cREDS element (grey shading) predicted as a promoter in psoas and an enhancer in lung. d, A box-plot of RNA-seq signals (RPKM) overlapping ±1 kb of cREDS enhancers, cREDS promoters, non-cREDS control enhancers and non-cREDS control promoters. ***P < 10 × 10−142, Wilcoxon test. e, RNA-seq and chromatin states of a cREDS element (grey shading) is shown for a region on chromosome 17 in H1 and IMR-90. Arrow indicates an alternative exon incorporated in IMR-90.
Recent studies showed that particular repetitive elements, such as endogenous retroviruses (ERVs), could participate in transcriptional regulation during mammalian development16,17,18. Given the representation of samples available, we systematically examined histone modifications at different classes of ERVs. While most are inactive, subsets, especially class I ERVs (ERV-I), are marked by H3K27ac in a tissue-restricted manner (Extended Data Fig. 3a and b). For instance, HERV-H element activities are restricted to human embryonic stem cells (hESCs) (Extended Data Fig. 3c, d). Furthermore, some ERVs carried marks of active promoters or enhancers (Extended Data Fig. 3d, e). We also observed that the LTR12C subfamily had substantial H3K27ac enrichment across different tissues (Extended Data Fig. 3e, f). Notably, the individual members appeared to be tissue restricted, suggesting that although the subfamily can be classified as non-tissue restrictively active, individual LTR12C elements were active only in distinct tissue/cell types (Extended Data Fig. 3e). Taken together, the data illustrate that human ERVs display precisely controlled patterns of activity in distinct tissues.
Intriguingly, 15.2% (n = 3,717) of strong promoters were also predicted as enhancers in other tissues, analogous to observations in mice, where intragenic enhancers act as promoters to produce cell-type-specific transcripts19. These sites possessed histone modification signatures of active enhancers in some tissue/cell types but were enriched with active promoter marks in others. We termed these sequences cis-regulatory elements with dynamic signatures (cREDS). For example, 1,321 cREDS enhancers showed enrichment of H3K27ac and H3K4me1 and a striking depletion of H3K4me3 in lung (Fig. 1b, c and Supplementary Table 3). However, the signature shifted to that of active promoters in other tissues (Fig. 1b, c). cREDS are also found in other cell/tissue types (Extended Data Fig. 4a). To determine whether cREDS have dual functions, we selected a subset of promoter-marked elements and validated their function with a luciferase reporter assay in hESCs. The majority (7 out of 10) showed promoter activity (Extended Data Fig. 4b). Similarly, 10 of 11 selected cREDS with enhancer signatures in hESCs also functioned as enhancers (Extended Data Fig. 4c). Additionally, subsets of enhancers previously validated in transgenic mice also possessed dynamic signatures (Extended Data Fig. 5)20. Furthermore, we selected two cREDS, predicted as enhancers in the left heart ventricle, with significant cap analysis of gene expression (CAGE) signal21, typical of active promoters (Extended Data Fig. 6a, b), and found that they possess heart-restricted enhancer activities in an in vivo zebrafish reporter assay (Extended Data Fig. 6c). Consistent with reporter activities, transcriptional properties (RNA-seq values based on reads per kilobase per million mapped reads (RPKM) within ±1 kb of the elements) of cREDS enhancers and promoters are similar to non-cREDS enhancers and promoters, respectively (Fig. 1d). Interestingly, when comparing isoform dynamics across H1 and IMR-90 RNA-seq data sets22 with cREDS identified between these two cell types we discovered that a subset of cREDS promoters was accompanied by creation of new transcripts and/or alternative exon usage (n = 99) (Fig. 1e), revealing a possible function whereby cREDS influence cell/tissue-specific transcript variants. Taken together, these data show that cREDS can potentially function as both promoters and enhancers in distinct cell types and fine-tune transcriptomes.
Reasoning that global analysis of allelic histone modification and gene expression patterns would elucidate mechanisms of long-range gene regulation by distal cis-regulatory elements, we re-analysed RNA-seq and ChIP-seq data sets by considering haplotype information. For this purpose, we applied HaploSeq1, which integrated genome sequencing with high-throughput chromatin conformation capture (Hi-C) data sets to derive chromosome-spanning haplotypes (see Supplementary Information). For four different tissue donors, we generated haplotypes spanning entire chromosomes with 99.5% completeness on average (the coverage of haplotype-resolved genomic regions) and average resolution (the coverage of phased heterozygous SNPs) ranging from 78% to 89% (Fig. 2a and Supplementary Tables 4 and 5). The accuracy of haplotype predictions was validated by the concordance with SNPs residing in the same paired-end sequencing reads. The concordance rates were 99.7% and 98.4% for H3K27ac ChIP-seq reads (described below) and RNA-seq reads, respectively, indicating high accuracy. We then re-analysed 36 mRNA-seq data sets from 18 tissues (including 16 tissues noted above with the addition of bladder and adipose tissue) and 187 ChIP-seq data sets for 6 histone modifications (Supplementary Table 6), from up to 4 individual donors, in a haplotype-resolved context.
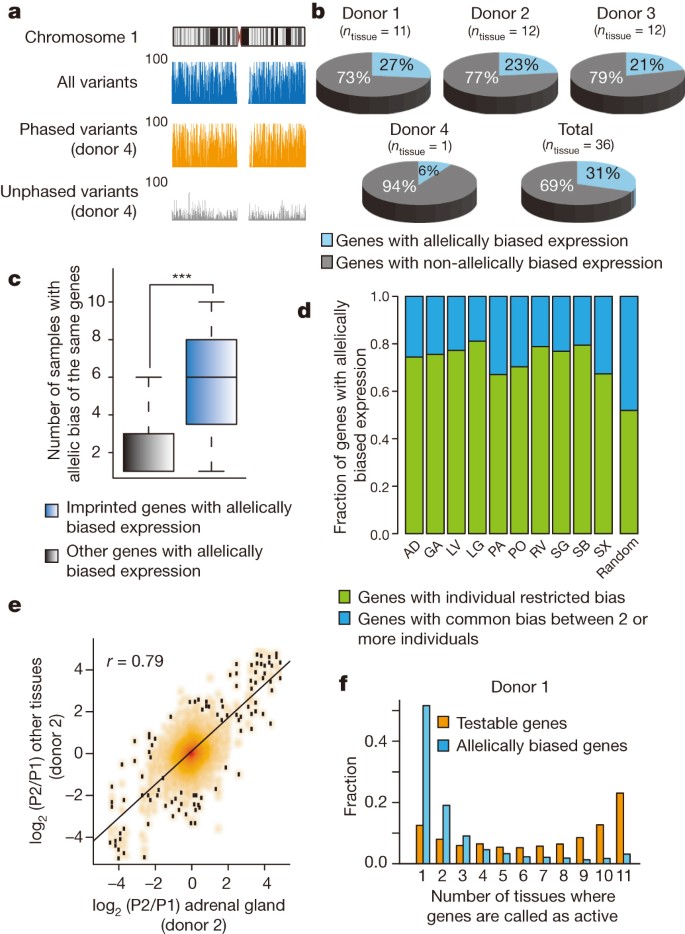
a, Genome browser snapshots illustrate completeness and resolution of haplotypes resolved in donor 4. The y axis indicates the number of variants within 100-kb windows. The density of all (blue), phased (orange) and unphased (grey) variants across chromosome 1 is shown. b, Proportion of genes with allelically biased expression among informative genes and the number of tissue samples derived from each donor (ntissue) are described. c, Box-plot illustrates occurrence of imprinted (n = 15) and other allelically biased genes, excluding pseudogenes (n = 2,334) across samples. ***P < 9.9 × 10−5, Kolmogorov–Smirnov (KS) test. d, Including only tissues with two or three equivalent samples derived from distinct donors (ntissue = 10), genes with allelic imbalances were defined as common between individuals (consistent bias among same tissue type from multiple donors) or as individual-restricted. Random control represents average from randomly selected samples (10,000 iterations). Abbreviations are defined in the legend to Fig. 1. e, Fold change of gene expressions between alleles (parental allele 1 (P1) and 2 (P2)) in adrenal gland from donor 2 (x axis) is compared to all other tissues from donor 2 (y axis). f, A histogram illustrates the proportions of allelically expressed genes in donor 1 (n = 1,375) defined in various numbers of tissues. The fraction of all testable genes or allelically expressed genes (y axis) is calculated for the number of tissues where they are identified as active (x axis) (P value <2.2 × 10−16, KS test).
Although widespread allelic imbalances in gene expression had been previously noted7,23,24,25, it remains unclear whether this phenomenon is consistent across distinct tissues and individuals, and the underlying mechanism remains undefined. To address the first point, we defined genes with allelically biased expression by means of mapping the RNA-seq reads in each tissue sample in a haplotype-resolved manner. We observed extensive allelically biased gene expression, ranging from 4% to 13% of all informative genes (>10 allelic read counts) in each tissue sample (false discovery rate (FDR) = 5%, Extended Data Fig. 7a, b). Comparatively, the proportion of allelically biased genes in individual tissue donors ranged from 6% to 23% of all informative genes, giving a combined total of 2,570 allelically biased genes (Fig. 2b and Supplementary Table 7). As a control, known imprinted genes (n = 15) showed common allelic biases across multiple samples (Fig. 2c) and donors (Extended Data Fig. 7c). Our data sets, representing the only collection of haplotype-resolved transcriptomes across an array of tissues from multiple individuals, allowed us to characterize allelic transcription across tissues and donors. While most genes with allelically biased expression demonstrate bias in multiple samples, approximately 75% exhibit statistically significant donor-specific bias (Fig. 2d and Extended Data Fig. 7d). This suggests a connection between sequence differences of individuals and allelically biased gene expression. In support of this model, genes frequently demonstrate consistent direction of allelic bias across multiple tissues of a given donor (Fig. 2e and Extended Data Fig. 7e). Interestingly, allelically biased genes were not restricted to the same tissue type across distinct donors. Rather, they were mostly specific to individual samples derived from each donor (Fig. 2f and Extended Data Fig. 7f), possibly resulting from differential levels of tissue-restricted transcription factors among different tissue samples.
As natural genetic variations can affect enhancer selection and function in mammalian cells26, we hypothesized that polymorphisms at cis-regulatory sequences underlie the widespread allelic transcriptional biases. We thus exploited the unique resource of 187 haplotype-resolved ChIP-seq data sets to analyse the state of cis-regulatory elements. We identified allelically biased marks at promoter regions (H3K27ac, H3K4me1, H3K4me3, H3K27me3 and H3K9me3) and transcribed gene bodies (H3K36me3) (see Supplementary Information). In support of our hypothesis, the allelic biases of gene expression strongly agreed with chromatin states of sequences at or near the genes (Fig. 3a, b and Extended Data Fig. 8a).
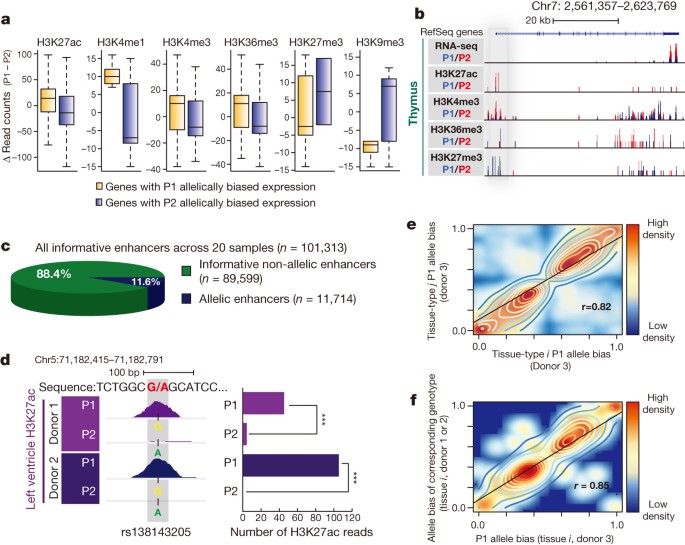
a, Box-plots present haplotype-resolved ChIP-seq reads at promoter or gene bodies (H3K27ac: n = 744, P = 10 × 10−14; H3K4me1: n = 32, P = 0.035; H3K4me3: n = 177, P = 0.0047; H3K27me3: n = 12, P = 0.43; H3K9me3: n = 27, P = 0.13; H3K36me3: n = 291, P = 4.3 × 10−6, KS test). b, Allelically biased gene expression of IQCE is concordant with chromatin marks at the promoter (grey) and gene body. c, Proportion of allelic (n = 11,714) and non-allelic (n = 89, 599) among all informative enhancers (n = 101,313) across 20 tissue samples. d, A snapshot showing a SNP (rs138143205) with H3K27ac bias towards the G allele in both left heart ventricle donors (left). Bar chart illustrates the number of H3K27ac reads corresponding to the P1 versus P2 alleles in both donors (right). ***P <10 × 10−19, binomial test. e, f, Scatter plots show strong correlation of the P1 allele bias of enhancer activities among two different tissue types from donor 3 (n = 4,427) (e) and among the P1 allele bias in donor 3 (x axis) and the allele bias of corresponding genotypes in donor 1 or 2 (y axis) at allelic enhancer in the same tissue type (f) (n = 447).
Furthermore, if allelic imbalances of enhancer activities indeed contribute to allelically biased gene expression, we expect that chromatin states at enhancers will be concordant with the expression of their targets. Therefore, we generated additional H3K27ac ChIP-seq data sets with deeper coverage and longer sequencing reads (for better delineation of alleles) for 14 of the previously analysed tissue samples and an additional 6 samples from independent donors (Supplementary Table 7). Of the informative enhancers (with >10 polymorphism-bearing sequence reads), 11.6% (n = 11,714, FDR = 1%) showed significant allelically biased H3K27ac enrichment in any tissue types (Fig. 3c and Supplementary Table 8). H3K27ac biases were validated by allele-specific ChIP-qPCR (Extended Data Fig. 8b). Interestingly, identical genotypes often yielded the same direction of biases in allelic enhancer activities (Fig. 3d). We further tested whether sequence variations are systematically associated with allelic H3K27ac, which reflects enhancer activities27. Indeed, H3K27ac biases were strongly correlated with specific genotypes, whereby given identical genotypes, this histone modification was biased to the same alleles, both across tissue types and individuals (Fig. 3d–f and Extended Data Fig. 9a). Furthering this finding, we analysed previously generated data sets from lymphoblastoid cell lines28 and found similar significant correlation of genotype and molecular phenotype of H3K27ac enrichment (Extended Data Fig. 9b). Taken together, these data reveal that extensive allelic imbalance events are associated with sequence variants in cis-regulatory elements.
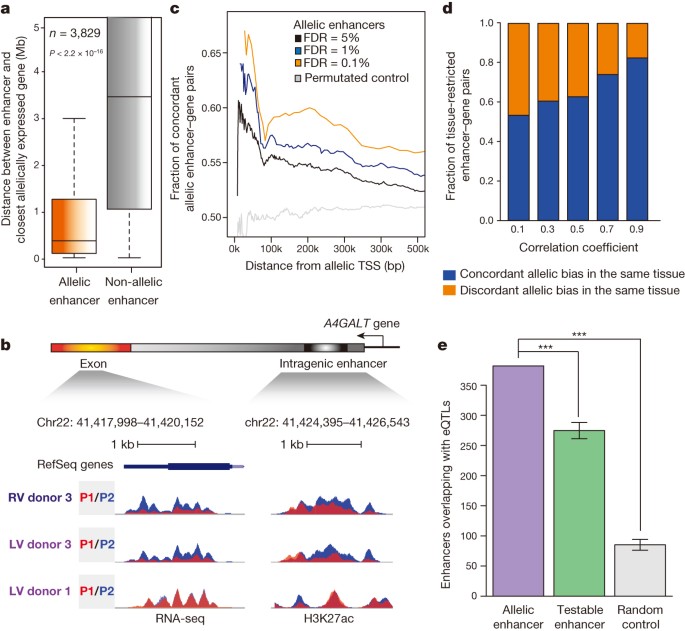
We discovered that allelic enhancers resided in significantly closer proximity to genes with allelically biased expression, as compared to non-allelic enhancers (Fig. 4a, b). We also observed examples where distinct tissues from the same donor showed similar allelic biases of gene expression and H3K27ac at enhancers (left ventricle and right ventricle from donor 3); however, the same tissue type derived from a different donor (left ventricle from donor 1) yielded no consistent patterns (Fig. 4b), supporting the hypothesis that allelically biased gene expression is driven by individual-specific genetic variation in enhancers. Indeed, within close proximity, the concordance between allelic enhancers and gene expression is significantly higher than permutated control enhancer/gene sets (Fig. 4c). Remarkably, 56% of allelic enhancer–gene pairs are greater than 300 kb apart (Extended Data Fig. 10a, b), the delineation of which was enabled by whole-chromosome-spanning haplotypes.
a, Average distance of allelic (5% FDR) and non-allelic enhancers to the closest allelically expressed gene is significantly different (n = 3,829, P < 2.2 × 10−16, KS test). b, Genome browser snapshots show an allelic enhancer within the intron of the allelically expressed A4GALT gene (P1, red; P2, blue) on chromosome 22 across three samples. c, Line plot presents the fraction of concordant allelic bias between allelically expressed genes and allelic enhancers in terms of distance. The allelic enhancer–gene pairs were defined with FDR cutoff values of 5% (n = 14,082) (black), 1% (n = 6,057) (blue) and 0.1% (n = 2,362) (orange). Permutated control of a set of enhancer–gene pairs was included (n = 14,082) (grey). Distance between allelically biased enhancer–gene pairs and fraction of concordant allelic bias are denoted by x and y axes, respectively (P < 2.2 × 10−16, KS test). d, Fractions of tissue-restricted enhancer–gene pairs (n = 3,106) (y axis) that show concordant (blue) or discordant (orange) allelic biases in the same tissue, are presented across a range of Pearson correlation coefficients (x axis) (P < 2.2 × 10−16, KS test, random permutated control concordant pairs = 50%). e, Overlap between eQTLs30 and allelic enhancers; testable enhancers or random control regions are shown. Error bars represent standard deviations. Testable enhancers and random control regions were generated 10,000 times with the same numbers as allelic enhancers. ***P < 10×10−5.
Similar to genes, many allelically biased enhancers are tissue restricted (Extended Data Fig. 10c). We reasoned that gene expression biases could result from tissue-restricted enhancer activities, evidenced by significant correlation between allelic enhancers and allelically expressed genes (Fig. 4d). Allelic enhancers also significantly overlapped with expression quantitative trait loci (eQTLs) (Fig. 4e), DNase I hypersensitivity QTLs and H3K27ac QTLs (Extended Data Fig. 10d), defined independently28,29,30, corroborating the functional roles of identified allelic enhancers on gene regulation. Taken together, these observations support a model whereby allelic biases of cis-regulatory element activities could be responsible for allelic gene expression.
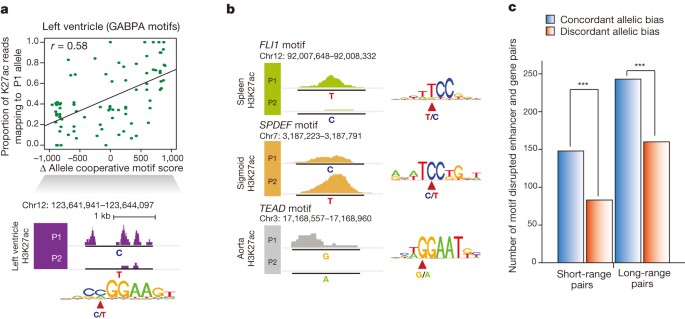
Finally, to elucidate further the mechanism by which allelically biased enhancer activities arise, we examined single nucleotide polymorphisms (SNPs) that potentially disrupt or weaken transcription factor binding motifs. We calculated changes in motif score between alleles (motif disruption score) at allelic enhancers and discovered 133 transcription factor motifs showing significant concordance between allelic reduction of enhancer activities and transcription factor motif disruption (Fig. 5a, b) (FDR = 10%, Supplementary Table 9). Moreover, genes with allelically biased expression were concordant with enhancer motif disruptions within close proximity (<20 kb) or displaying strong Hi-C interactions at longer distances (>20 kb) (Fig. 5c and Supplementary Information). Our results therefore suggest that genetic variations are probably responsible for allelic enhancer activities and consequently allelically biased gene expression.
a, Differential GABPA binding motif scores between two alleles (P1 − P2 motif scores) in left heart ventricle are correlated with the proportion of H3K27ac reads corresponding to the P1 allele (top). Values range from negative to positive, indicating P1 and P2 motif disruption, respectively. An example on chromosome 12 illustrating that P1, with a motif preserving C allele, has higher H3K27ac enrichment and that P2, with the motif disrupting T allele, has little H3K27ac enrichment (bottom). b, Three examples (FLI1 in spleen, SPDEF in sigmoid colon, and TEAD in aorta) of motif-disrupted enhancers demonstrate allelic biased activities. The variant location and genotypes of P1 and P2 alleles are marked in motif sequence. c, All possible motif disrupted enhancer–gene pairs within the indicated distance window are defined with concordant allelic bias (blue, gene–enhancer pairs with biases towards the same allele) or discordant allelic bias (red, gene–enhancer pairs with biases towards different alleles). Only thymus, left heart ventricle and aorta were considered due to the availability of Hi-C data. Short-range pairs are defined if any allelically expressed genes are located <20 kb away. ***P < 2.5 × 10−5, binomial test.
By generating chromosome-spanning haplotypes, we carried out a comprehensive survey of allelic chromatin state and gene expression. We found evidence for extensive allelically biased gene expression, which is connected to change in chromatin states at cis-regulatory elements, probably resulting from transcription factor binding disruption by sequence variations. These observations mirror findings in mice where allelic biases of cis-regulatory element activities could be responsible for allelic gene expression26, and demonstrate that such a phenomenon is probably widespread in the human genome. These observations shed light on the importance of considering genetic variants in understanding individual-specific gene regulation. Analyses of haplotype-resolved transcriptomes and epigenomes in additional individuals and tissues should further illuminate the role of sequence variations in defining individual-specific transcriptional programs and phenotypes.
References
Selvaraj, S., Dixon, J. R., Bansal, V. & Ren, B. Whole-genome haplotype reconstruction using proximity-ligation and shotgun sequencing. Nature Biotechnol. 31, 1111–1118 (2013)
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013)
Zhu, J. et al. Genome-wide chromatin state transitions associated with developmental and environmental cues. Cell 152, 642–654 (2013)
Rivera, C. M. & Ren, B. Mapping human epigenomes. Cell 155, 39–55 (2013)
Heintzman, N. D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nature Genet. 39, 311–318 (2007)
Rajagopal, N. et al. RFECS: a random-forest based algorithm for enhancer identification from chromatin state. PLOS Comput. Biol. 9, e1002968 (2013)
Fang, F. et al. Genomic landscape of human allele-specific DNA methylation. Proc. Natl Acad. Sci. USA 109, 7332–7337 (2012)
Stergachis, A. B. et al. Developmental fate and cellular maturity encoded in human regulatory DNA landscapes. Cell 154, 888–903 (2013)
Andersson, R. et al. An atlas of active enhancers across human cell types and tissues. Nature 507, 455–461 (2014)
Flandez, M. et al. Nr5a2 heterozygosity sensitises to, and cooperates with, inflammation in KRas(G12V)-driven pancreatic tumourigenesis. Gut 63, 647–655 (2014)
Hirai, H. et al. Involvement of Runx1 in the down-regulation of fetal liver kinase-1 expression during transition of endothelial cells to hematopoietic cells. Blood 106, 1948–1955 (2005)
Hwang, D. H. et al. Transplantation of human neural stem cells transduced with Olig2 transcription factor improves locomotor recovery and enhances myelination in the white matter of rat spinal cord following contusive injury. BMC Neurosci. 10, 117 (2009)
Jahan, I., Kersigo, J., Pan, N. & Fritzsch, B. Neurod1 regulates survival and formation of connections in mouse ear and brain. Cell Tissue Res. 341, 95–110 (2010)
Lee, C. S. et al. Loss of nuclear factor E2-related factor 1 in the brain leads to dysregulation of proteasome gene expression and neurodegeneration. Proc. Natl Acad. Sci. USA 108, 8408–8413 (2011)
Moya, M. et al. Foxa1 reduces lipid accumulation in human hepatocytes and is down-regulated in nonalcoholic fatty liver. PLoS ONE 7, e30014 (2012)
Kunarso, G. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nature Genet. 42, 631–634 (2010)
Xie, M. et al. DNA hypomethylation within specific transposable element families associates with tissue-specific enhancer landscape. Nature Genet. 45, 836–841 (2013)
Lu, X. et al. The retrovirus HERVH is a long noncoding RNA required for human embryonic stem cell identity. Nature Struct. Mol. Biol. 21, 423–425 (2014)
Kowalczyk, M. S. et al. Intragenic enhancers act as alternative promoters. Mol. Cell 45, 447–458 (2012)
Visel, A., Minovitsky, S., Dubchak, I. & Pennacchio, L. A. VISTA Enhancer Browser–a database of tissue-specific human enhancers. Nucleic Acids Res. 35, D88–D92 (2007)
The FANTOM Consortium and the RIKEN PMI and CLST (DGT) A promoter-level mammalian expression atlas. Nature 507, 462–470 (2014)
Trapnell, C. et al. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nature Biotechnol. 31, 46–53 (2013)
Gimelbrant, A., Hutchinson, J. N., Thompson, B. R. & Chess, A. Widespread monoallelic expression on human autosomes. Science 318, 1136–1140 (2007)
Kilpinen, H. et al. Coordinated effects of sequence variation on DNA binding, chromatin structure, and transcription. Science 342, 744–747 (2013)
Skelly, D. A., Johansson, M., Madeoy, J., Wakefield, J. & Akey, J. M. A powerful and flexible statistical framework for testing hypotheses of allele-specific gene expression from RNA-seq data. Genome Res. 21, 1728–1737 (2011)
Heinz, S. et al. Effect of natural genetic variation on enhancer selection and function. Nature 503, 487–492 (2013)
Creyghton, M. P. et al. Histone H3K27ac separates active from poised enhancers and predicts developmental state. Proc. Natl Acad. Sci. USA 107, 21931–21936 (2010)
McVicker, G. et al. Identification of genetic variants that affect histone modifications in human cells. Science 342, 747–749 (2013)
Kasowski, M. et al. Extensive variation in chromatin states across humans. Science 342, 750–752 (2013)
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511 (2013)
Acknowledgements
This work is supported by the NIH Epigenome Roadmap Project (U01 ES017166), CIRM RN2-00905-1, NIH ES017166, NSFC 91019016, NBRPC 2012CB316503 and NIH Fellowship Grants F32HL110473 and K99HL119617. We thank A. Kulkarni and J. Wu for help with processing RNA-seq data sets, and Y. He and M. Schultz for discussions regarding allelic analyses of RNA-seq data sets. We also thank members of the Ren laboratory for comments.
Author information
Authors and Affiliations
Contributions
D.L., W.X., J.R.E., N.C.C. and B.R. led the data production. I.J., N.R., M.Q.Z., J.R.E. and B.R. led the data analyses. I.J., N.R., S.S., F.Y., Y.Q., L.E., M.H., A.S. and P.R. conducted analyses. S.L. and Y.L. processed tissue samples. D.L., A.S., A.Y.L., C.-A.Y., S.K. and H.Y. produced data. D.L., I.J., N.R. and B.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.S. and B.R. are co-founders of Arima Genomics.
Extended data figures and tables
Extended Data Figure 1 Active enhancers cluster along developmental lineages.
a, Pie charts showing fractions of tissue-restricted and non-tissue-restricted strong enhancers and promoters. b, Hierarchical clustering with optimal leaf ordering based on all H3K27ac-marked highly active enhancers. Four major clusters are represented: early embryonic cell types (blue); a large set of meso/endoderm-derived tissues (dark green); a set consisting of ectoderm-derived brain tissues (red); and a small cluster of mesoderm cell lines (purple), which bridged the early embryonic lineages with the somatic tissues. Although TRO did not fall within any clusters, it shared the highest degree of similarity with the early embryonic cell lines. On a subsequent level, two clusters are seen separating endoderm-derived tissues (grey line) and mesoderm-derived tissues (green line). Heart tissues are denoted by yellow asterisks. c, Clustering of tissues by promoters’ histone acetylation status shows grouping of tissues that are of similar types but are less evident in germ-layer divisions than clustering of enhancers.
Extended Data Figure 2 Tissue-restricted enhancers are enriched for transcription factor motifs important for cell identity and/or function.
Significantly enriched motifs (P < 10 × 10−10) across all 28 tissues are divided into 29 clusters (method described in Supplementary Information). An overall P value is generated for the enrichment of each tissue for each cluster. The figure illustrates –log(P value) of (a) pancreas, (b) anterior caudate and (c) liver-restricted enhancer motif enrichment for the various clusters. For ease of visualization, any cluster with P values greater than 0.05 is denoted 0. Red highlighted text refers to a subset of motifs for transcription factors with literature support (see Supplementary Information) to have function in (a) the pancreas, (b) the brain and (c) the liver.
Extended Data Figure 3 Endogenous retroviruses (ERVs) are enriched for active cis-regulatory element marks in a tissue-restricted fashion.
a, A clustered heat map showing the H3K27ac enrichment (RPKM) of all mappable elements of the three classes of ERVs. b, Distribution of the Shannon entropy of H3K27ac across enhancers, promoters and three classes of ERVs is shown as a density curve, demonstrating that H3K27ac enrichment of ERVs is more tissue-restricted than promoters and slightly less than enhancers. c, Box-plots illustrating the H3K27ac enrichment of 127 mappable members of the HERV-H subfamily across all tissue/cell types. The enrichment in H1 hESCs is significantly higher than all other cell/tissues types (P < 1.4 × 10−9, Wilcoxon test). d, UCSC genome browser snapshots showing example of an HERV-H element harbouring H1-restricted active promoter marks, corresponding RNA-seq signal and H3K36me3 enrichment. Notably, this particular element has been annotated in Refseq as the ES cell related gene (ESRG), a human-specific long non-coding RNA gene. e, UCSC genome browser snapshots showing example of a LTR12C element harbouring TRO-restricted active enhancer chromatin marks. f, A matrix illustrating the average H3K27ac enrichment for subfamilies of class I ERVs across all cell and tissue types. LTR12C subfamily (green arrow) shows enrichment of H3K27ac across many distinct cell types and tissues.
Extended Data Figure 4 cREDS are enriched with dynamic histone mark signatures in different tissues and have putative cis-regulatory functions.
a, Heat maps showing the enrichment (RPKM) of the H3K27ac, H3K4me3 and H3K4me1 at MES-restricted enhancers (n = 650), which are predicted as promoters in other tissues, across all 28 samples. The red box highlights the histone modifications in MES cells. b, A schematic of the pGL3-enhancer vector used in these luciferase-reporter assays (top) and the activity of 10 selected cREDS with promoter signatures and a negative control region cloned 5′ to the reporter gene after transfection into H1 hESCs (bottom). Luciferase activity of each region is normalized to the average activity of the negative controls. c, A schematic of the pGL3 promoter vector used in these luciferase-reporter assays (top) and the activity of 11 selected cREDS with enhancer signatures and a negative control region cloned 3′ to the reporter gene after transfection into H1 hESCs (bottom). Luciferase activity of each region is normalized to averaged activity of negative control regions. Error bars reflect standard deviation between three technical replicates.
Extended Data Figure 5 VISTA validated enhancers also possess dynamic histone modification signatures across tissues.
Example screen shots of VISTA validated enhancers and the patterns of activity in vivo are displayed along with histone modification patterns in representative tissues (adapted from VISTA enhancer browser20).
Extended Data Figure 6 cREDS show enrichment of CAGE signal and putative enhancer functions in a zebrafish reporter assay.
a, UCSC genome browser screen shots show the two cREDS elements (grey shading) harbouring enhancer and promoter signatures in distinct tissue types. When compared to CAGE data sets from the FANTOM5 project, these elements show substantial overlap with transcript signals (red and blue signals indicate CAGE signal on the forward and reverse orientation, respectively). b, Selected cREDS (same elements as above) with enhancer marks in left ventricle show heart-restricted enhancer activity, as indicated by GFP expression, in 3 days post-fertilization (3 dpf) zebrafish larvae. In parallel, pT2MX negative control did not show any GFP expression. White arrow indicates location of the 3 dpf zebrafish heart. For enhancer 1, 13 out of the 38 surviving embryos showed similar patterns. For enhancer 2, 18 out of the 35 surviving embryos showed similar patterns. None of the 30 surviving embryos, injected with the control vector, showed any appreciable GFP signal in the heart. Scale bar, 50 μm.
Extended Data Figure 7 Identification of widespread allelically expressed genes.
a, Fraction of genes with allelically biased expression in each sample. y axis indicates number of samples and x axis indicates fraction of allelically biased genes among informative genes (more than 10 SNP-containing short reads). b, Distribution of fold change of allelically biased genes between P1 and P2 alleles. c, The occurrence of allelically biased imprinted and other genes is shown. x axis refers to the number of individual donors where corresponding allelically expressed genes are commonly detected. d, A density plot showing the fraction of sample-restricted genes with allelically biased expression (grey). Three tissue samples were randomly selected and sample-restricted allelically expressed genes were defined, which includes random variance effect. The random selection was repeated 10,000 times. The shaded blue box indicates the range of fractions of individual-restricted allele-biased genes in all analysed tissue types (n = 10). The fraction of sample-restricted allelically biased genes is lower than individual-restricted allele-biased genes in Fig. 2e. e, Fold change of allele-biased gene expression between two alleles is shown as scatter plot. x axis is for the fold changes in one randomly selected tissue in each donor and y axis is for the fold changes in all other remaining tissues in the corresponding donor. Allelic bias in one tissue is highly correlated with allelic bias in other tissues in the same individual. f, A histogram illustrates the proportions of allelically expressed genes in donor 2 (left) and 3 (right) defined in various numbers of tissues. The fraction of all testable genes or allelically expressed genes (y axis) is calculated for the number of tissues where they are called as active (x axis). The results indicate that the majority of allelically biased genes, as opposed to testable genes, are restricted to one or two tissue samples. KS test was performed between allele-biased genes and testable genes (P < 2.2 × 10−16).
Extended Data Figure 8 Allele-biased chromatin states.
a, Box-plots illustrating haplotype-resolved ChIP-seq signal enrichment on the two alleles at promoter regions. The P1 or P2 allele-biased promoter regions were defined by H3K27ac signals and then H3K4m1, H3K4me3 and H3K9me3 signals were presented for the corresponding promoter regions. All chromatin states are consistent according to the allele-biased H3K27ac patterns. KS test was performed for P value calculation. b, Allelically biased enhancers were tested in thymus from donor 1 and pancreas from donors 2 and 3. H3K27ac enrichment was tested by allele-specific ChIP-qPCR. Two control enhancers were included and showed to have no allelic biases in thymus or pancreas from donor 2 (top right and bottom left, respectively).
Extended Data Figure 9 Putative enhancers with identical genotypes in different individuals exhibit similar biases in histone acetylation.
a, Scatter plots of P1 allele-biased enhancer activities for pairwise comparison of allele-biased enhancers in donor 1 (n = 85) and donor 2 (n = 4,427). x and y axes indicate P1 allele bias. b, Scatter plot of reference allele bias of enhancer activities for pairwise comparison of allele-biased enhancers in all tissues from all three donors and lymphoblastoid data sets obtained from a previous study28 (n = 309).
Extended Data Figure 10 Analyses of concordant allelically biased gene–enhancer pairs.
a, Frequency of allelically expressed genes according to the distance between concordantly allele-biased enhancer–gene pairs. Blue bars represent data obtained from whole-chromosome-spanning haplotype blocks while green bars represent data obtained from simulated 300-kb haplotype blocks. 56% of enhancer–gene pairs are more than 300 kb apart. b, Accumulation curve showing fraction of allelically biased genes that have at least one concordantly allelic enhancer within a given distance (x axis). Up to 83% of allelically expressed genes are within 2 Mb of a concordantly biased allelic enhancer. c, The frequency of allele-biased enhancers in donors 1, 2 and 3. y axis indicates fraction of enhancers and x axis indicates frequency of allelically biased enhancers. KS test was performed between allele-biased enhancers and testable enhancers. d, Bar plots presenting the number of enhancers overlapping with DHS QTLs and H3K27ac QTLs for allelic enhancers, testable enhancers, and random control regions. ***P < 10 × 10−5.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Figure 1, Supplementary Tables 4 – 9 (see separate excel files for Supplementary Tables 1-3) and Supplementary References. (PDF 728 kb)
Supplementary Table 1
This table shows all prediced enhancers across 28 tissues/cell-types. (XLSX 6403 kb)
Supplementary Table 2
This table shows all predicted promoters with strong H3K4me3 across single sample of all 28 tissues/cell-types. (XLS 1931 kb)
Supplementary Table 3
This table contains a list of cREDS elements defined across all tissues. (XLSX 198 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported licence. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Leung, D., Jung, I., Rajagopal, N. et al. Integrative analysis of haplotype-resolved epigenomes across human tissues. Nature 518, 350–354 (2015). https://doi.org/10.1038/nature14217
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14217
This article is cited by
-
A distant global control region is essential for normal expression of anterior HOXA genes during mouse and human craniofacial development
Nature Communications (2024)
-
Prioritization of risk genes in colorectal cancer by integrative analysis of multi-omics data and gene networks
Science China Life Sciences (2024)
-
Genetic impacts on DNA methylation help elucidate regulatory genomic processes
Genome Biology (2023)
-
Crosstalk between RNA m6A and DNA methylation regulates transposable element chromatin activation and cell fate in human pluripotent stem cells
Nature Genetics (2023)
-
De novo and inherited variants in coding and regulatory regions in genetic cardiomyopathies
Human Genomics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.