Abstract
Epithelium folding is a basic morphogenetic event that is essential in transforming simple two-dimensional epithelial sheets into three-dimensional structures in both vertebrates and invertebrates1. Folding has been shown to rely on apical constriction2,3,4,5,6,7. The resulting cell-shape changes depend either on adherens junction basal shift2 or on a redistribution of myosin II3,4,5,7, which could be driven by mechanical signals8. Yet the initial cellular mechanisms that trigger and coordinate cell remodelling remain largely unknown. Here we unravel the active role of apoptotic cells in initiating morphogenesis, thus revealing a novel mechanism of epithelium folding. We show that, in a live developing tissue, apoptotic cells exert a transient pulling force upon the apical surface of the epithelium through a highly dynamic apico-basal myosin II cable. The apoptotic cells then induce a non-autonomous increase in tissue tension together with cortical myosin II apical stabilization in the surrounding tissue, eventually resulting in epithelium folding. Together our results, supported by a theoretical biophysical three-dimensional model, identify an apoptotic myosin-II-dependent signal as the initial signal leading to cell reorganization and tissue folding. This work further reveals that, far from being passively eliminated as generally assumed (for example, during digit individualization9), apoptotic cells actively influence their surroundings and trigger tissue remodelling through regulation of tissue tension.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Davidson, L. A. Epithelial machines that shape the embryo. Trends Cell Biol. 22, 82–87 (2012)
Wang, Y. C., Khan, Z., Kaschube, M. & Wieschaus, E. F. Differential positioning of adherens junctions is associated with initiation of epithelial folding. Nature 484, 390–393 (2012)
Martin, A. C., Kaschube, M. & Wieschaus, E. F. Pulsed contractions of an actin-myosin network drive apical constriction. Nature 457, 495–499 (2009)
Roh-Johnson, M. et al. Triggering a cell shape change by exploiting preexisting actomyosin contractions. Science 335, 1232–1235 (2012)
Sherrard, K., Robin, F., Lemaire, P. & Munro, E. Sequential activation of apical and basolateral contractility drives ascidian endoderm invagination. Curr. Biol. 20, 1499–1510 (2010)
He, B., Doubrovinski, K., Polyakov, O. & Wieschaus, E. Apical constriction drives tissue-scale hydrodynamic flow to mediate cell elongation. Nature 508, 392–396 (2014)
Brodu, V. & Casanova, J. The RhoGAP crossveinless-c links trachealess and EGFR signaling to cell shape remodeling in Drosophila tracheal invagination. Genes Dev. 20, 1817–1828 (2006)
Pouille, P. A., Ahmadi, P., Brunet, A. C. & Farge, E. Mechanical signals trigger myosin II redistribution and mesoderm invagination in Drosophila embryos. Sci. Signal. 2, ra16 (2009)
Montero, J. A. & Hurlé, J. M. Sculpturing digit shape by cell death. Apoptosis 15, 365–375 (2010)
Lohmann, I., McGinnis, N., Bodmer, M. & McGinnis, W. The Drosophila Hox gene deformed sculpts head morphology via direct regulation of the apoptosis activator reaper. Cell 110, 457–466 (2002)
Manjón, C., Sánchez-Herrero, E. & Suzanne, M. Sharp boundaries of Dpp signalling trigger local cell death required for Drosophila leg morphogenesis. Nature Cell Biol. 9, 57–63 (2007)
Yamaguchi, Y. et al. Live imaging of apoptosis in a novel transgenic mouse highlights its role in neural tube closure. J. Cell Biol. 195, 1047–1060 (2011)
Lubkov, V. & Bar-Sagi, D. E-cadherin-mediated cell coupling is required for apoptotic cell extrusion. Curr. Biol. 24, 868–874 (2014)
Fernandez-Gonzalez, R., Simoes, S. M., Röper, J. C., Eaton, S. & Zallen, J. A. Myosin II dynamics are regulated by tension in intercalating cells. Dev. Cell 17, 736–743 (2009)
Farhadifar, R., Röper, J. C., Aigouy, B., Eaton, S. & Jülicher, F. The influence of cell mechanics, cell-cell interactions, and proliferation on epithelial packing. Curr. Biol. 17, 2095–2104 (2007)
Rosenblatt, J., Raff, M. C. & Cramer, L. P. An epithelial cell destined for apoptosis signals its neighbors to extrude it by an actin- and myosin-dependent mechanism. Curr. Biol. 11, 1847–1857 (2001)
Toyama, Y., Peralta, X. G., Wells, A. R., Kiehart, D. P. & Edwards, G. S. Apoptotic force and tissue dynamics during Drosophila embryogenesis. Science 321, 1683–1686 (2008)
Yamaguchi, Y. & Miura, M. How to form and close the brain: insight into the mechanism of cranial neural tube closure in mammals. Cell. Mol. Life Sci. 70, 3171–3186 (2013)
Huang, J., Zhou, W., Dong, W., Watson, A. M. & Hong, Y. From the Cover: Directed, efficient, and versatile modifications of the Drosophila genome by genomic engineering. Proc. Natl Acad. Sci. USA 106, 8284–8289 (2009)
Oda, H. & Tsukita, S. Real-time imaging of cell-cell adherens junctions reveals that Drosophila mesoderm invagination begins with two phases of apical constriction of cells. J. Cell Sci. 114, 493–501 (2001)
Ishihara, S. & Sugimura, K. Bayesian inference of force dynamics during morphogenesis. J. Theor. Biol. 313, 201–211 (2012)
Takemoto, K., Nagai, T., Miyawaki, A. & Miura, M. Spatio-temporal activation of caspase revealed by indicator that is insensitive to environmental effects. J. Cell Biol. 160, 235–243 (2003)
Royou, A., Sullivan, W. & Karess, R. Cortical recruitment of nonmuscle myosin II in early syncytial Drosophila embryos: its role in nuclear axial expansion and its regulation by Cdc2 activity. J. Cell Biol. 158, 127–137 (2002)
Aldaz, S., Escudero, L. M. & Freeman, M. Live imaging of Drosophila imaginal disc development. Proc. Natl Acad. Sci. USA 107, 14217–14222 (2010)
Abrams, J. M., White, K., Fessler, L. I. & Steller, H. Programmed cell death during Drosophila embryogenesis. Development 117, 29–43 (1993)
Caserta, T. M., Smith, A. N., Gultice, A. D., Reedy, M. A. & Brown, T. L. Q-VD-OPh, a broad spectrum caspase inhibitor with potent antiapoptotic properties. Apoptosis 8, 345–352 (2003)
Guarner, A. et al. The zinc finger homeodomain-2 gene of Drosophila controls Notch targets and regulates apoptosis in the tarsal segments. Dev. Biol. 385, 350–365 (2014)
Peixoto, P. T. The graph-tool python library. figshare http://dx.doi.org/10.6084/m9.figshare.1164194 (2014)
van der Walt, S., Colbert, S. C. & Varoquaux, G. The NumPy Array: a structure for efficient numerical computation. Comput. Sci. Eng. 13, 22–30 (2011)
Acknowledgements
We thank C. Benassayag, E. Farge, Y. Gachet, T. Lecuit, P.-F. Lenne, F. Payre, E. Sanchez-Herrero, B. Sanson and S. Tournier for comments on the manuscript, T. de Paula Peixoto for providing the graph-tool library, D. Kiehart, H. Steller, X. Wang, R. Ward, BDSC and DSHB for stocks and reagents, the TRI platform for imaging facilities and M. Aguirrebengoa for helping us with statistics. The Suzanne laboratory is supported by grants from the Agence Nationale de la Recherche (ANR), Fondation de la Recherche et de l’Innovation Thérapeutique en Cancérologie (RITC) and the University of Toulouse.
Author information
Authors and Affiliations
Contributions
M.G., B.M. and M.S. have done the experiments in flies with the help of S.S. and A.G.; G.G. designed the simulation model. T.M. set up cell shape extraction, statistics and analysis in Matlab and performed the laser ablation experiments with the help of B.M.; M.S. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
The source code for the model is released under the GNU General Public Licence and is available on GitHub (https://github.com/glyg/leg-joint and http://dx.doi.org/10.5281/zenodo.13386).
Extended data figures and tables
Extended Data Figure 1 Spatio-temporal pattern of fold formation and hallmarks of apoptosis.
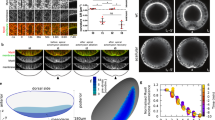
This figure is associated with Fig. 1. a, Time-lapse images and schematics of the distal region of a Dlg::GFP leg disc from pre-fold stage (WP) showing the progression of the t4-t5 fold (white arrowheads). Acridine orange was used to stain dying cells (green) and FM4-64 to stain membranes (red). Note the presence of apoptotic cells in the fold region (coloured in grey in the schematics below, n = 27). The t4-t5 domain is indicated by a black line on the schematics. b, Three-dimensional reconstruction of the distal region of Dlg::GFP pupal legs undergoing fold morphogenesis. The colour code indicates tissue depth. Images show legs at different stages of development, from WP to WP + 3 h. Throughout this study, we have focused on the t4-t5 fold, the only one that is exclusively formed during pupal stage and for which progression can be easily followed due to disc evagination. For each time point, top and bottom panels show dorsal and ventral views of leg discs respectively, and the t4-t5 domain is indicated by a white line. Note that the fold is initiated in the most ventral part of the leg, then progresses laterally (arrows) to end in the most dorsal part of the leg (n = 10 for WP, n = 7 for WP + 1 h, n = 10 for WP + 2 h, n = 10 for WP + 3 h). c, High magnification images from a time-lapse video during apoptosis showing caspase activity, revealed by the FRET construct SCAT3 (top), and the outline of cell membranes, revealed by FM4-64 staining (bottom) (n = 19). These images illustrate that the classical apoptotic stages, including shrinkage, blebbing (hollow arrowheads) and fragmentation (black arrowheads), are recapitulated in the developing Drosophila leg epithelium. Black outline (top) and red false-colour (bottom) highlight the apoptotic cell. Another apoptotic cell (outlined in white and coloured in pink) has also just turned on the apoptotic pathway. Note that in both cases the apoptotic pathway is turned on before visible morphological change.
Extended Data Figure 2 Adherens junctions and acto-myosin cytoskeleton dynamics in apoptotic cells.
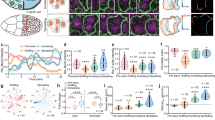
This figure is associated with Fig. 1. a, Time lapse images of a pupal leg disc expressing αCat-TagRFP in the ap domain (which encompasses the t4-t5 fold) and the FRET construct SCAT3 to reveal caspase activity and thus visualize apoptotic cells (n = 10). The apoptotic cell outline is visible on the sagittal section and represented on the scheme. The position of the sagittal section is indicated by a black line on the apical top view to the left. Note the reduction of the apical surface of the apoptotic cell (apical top views on the right). The apoptotic cell is highlighted in red on both panels. b, Acto-myosin cable (green arrowhead) observed in a single apoptotic cell from a pupal leg disc (n = 6). MRLC::GFP is in green, F-actin (stained with phalloidine) is blue and cleaved Dcp1 is red. c, Sagittal sections of an apoptotic cell (visualized with the FRET construct SCAT3) before and after fragmentation, illustrating the apoptotic adhesion peak (n = 27, red arrowhead) and its splitting up (hollow red arrowhead) when the apoptotic cell detaches at fragmentation. d, Time-lapse images of a MRLC::GFP pupal leg disc showing apical surface release upon apoptotic cell fragmentation (time-point 69 min, n = 12). Fragmentation is clearly visible in the z section (top) showing membrane staining of the apoptotic fragments with FM4-64 (white arrowheads) (the position of the z section is indicated by a black dotted line in the sagittal view). The myosin II cable is indicated by hollow open red arrowheads in sagittal sections. e, MRLC::GFP leg discs incubated in either DMSO (control) or Q-VD-OPH (cell death inhibition), showing the absence of the apico-basal myosin II cable in the absence of cell death (right, n = 0 out of 30). Arrowhead points out to the myosin II cable in the control (n = 14 out of 30). f, A single apoptotic cell (same cell shown in Fig. 1f, identified by GFP expression, red) generated by ectopic expression of reaper in the wing disc (from y,w,hs::flp; act-frt-y+-frt-Gal4, uas::GFP / uas::lifeactGFP; uas::rpr larvae). The green arrowhead points out to the myosin II cable stained by anti-phospho-Sqh/MRLC (green, n = 11). The dotted red line outlines the dying cell. g, Percentage of individual apoptotic cells (with or without myosin II activity) with apical deformation. This quantification is associated with Fig. 1f, g (n = 68 and n = 106, respectively). Genotype for ectopic cell death is y,w,hs::flp; act-frt-stop-frt::Gal4, uas::GFP / uas::lifeactGFP; uas::rpr and genotype for ectopic cell death without myosin II activity specifically in apoptotic cells is y,w,hs::flp; uas::DN-zip::GFP, uas::hid / act-frt-stop-frt::Gal4. Note that, in the latter condition, myosin II activity is maintained in the neighbouring cells. h, Sagittal sections close-ups of MRLC::GFP wing and antennal discs from 3rd instar larvae and a MRLC::GFP stage 11 embryo showing that an apico-basal structure of myosin II (green arrowheads) is formed in dying cells in each of these different tissues (n = 6, 7 and 7, respectively). Cleaved Dcp1 (activated caspase) is in red, E-Cad in blue (wing) and Arm/βCat in blue. The region shown in each close up is indicated by a red line on schematics on the left.
Extended Data Figure 3 Myosin II and tension dynamics during fold formation.
This figure is associated with Fig. 2. a, Close-up views of cells expressing MRLC::GFP, showing that, in the leg disc, myosin II is preferentially accumulated at the cortex (green) rather than at the medio-apical web (magenta). Relative intensity of myosin II at both levels is shown on separate panels (n = 22). b, Sagittal section of a leg disc from pre-fold stage (WP) showing that myosin II (visualized by MRLC::GFP construct, in green) co-localizes with E-Cad at adherens junctions (labelled in red) (n = 22). c, Quantification of myosin II::GFP (left) and F-actin (right) levels at individual junctions in fold (t4-t5) and segment (t4) domains of leg discs from pre-fold (young WP + 1 h) and late-fold (young WP + 4 h) stages (n = 144 and n = 112, respectively). Values are represented as mean values with error bars representing standard errors. The intensity of the signal in the fold has been normalized with the mean intensity in the segment domain. We used the non-parametric Wilcoxon rank sum test (also called Mann and Whitney test). d, e, Quantification of myosin II::GFP (left) and F-actin (right) levels per surface unit in fold (t4-t5) and segment (t4) domains of (d) leg discs from pre-fold (young WP + 1 h) and late-fold (young WP + 4 h) stages (n = 23 and n = 18 measurements respectively) or (e) in dissected discs cultured from pre-fold to mid-fold stage with either DMSO (control) or Q-VD-OPh (cell death inhibition) (n = 32 and n = 24 measurements respectively). Values are represented as mean values with error bars representing standard errors. The intensity of the signal in the fold has been normalized with the mean intensity in the segment domain. We used the non-parametric Wilcoxon rank sum test (also called Mann and Whitney test). f, Sagittal sections of pupal leg discs from mid-fold stage (WP + 2 h) of the following genotypes: Dll-Gal4[MD23] (ctl, n = 8), UAS-DIAP1;LP30-Gal4 (Diap1, n = 27), Dll-Gal4[MD23] / UAS-p35 (p35, n = 11) and Dll-Gal4[MD23] / UAS-p35; UAS-p35 (p35 x2, n = 15) (at early pupal stages, LP30-Gal4 and Dll::Gal4[MD23] show similar expression patterns, namely expression in the distal tibia and in all tarsal segments27). The stabilization of myosin II and F-actin in the t4-t5 fold observed in the control (red arrowheads) is reduced or absent when cell death is inhibited (open arrowheads point out to the t4-t5 domain in the context of cell death inhibition). Myosin II is detected using anti-Sqh/MRLC antibody. The fold domain is false-coloured in pink and the segment domain in yellow. g, Quantification of myosin II (using anti-sqh antibody, left) and F-actin (right) levels per surface unit in fold (t4-t5) and segment (t4) domains in leg discs from mid-fold stage (WP + 2 h) in control (DllGal4) and cell inhibition (Dll>p35x2) contexts (n = 25 and n = 28 measurements respectively). Values are represented as mean values with error bars representing standard errors. The intensity of the signal in the fold has been normalized with the mean intensity in the segment domain. We used the non-parametric Wilcoxon rank sum test (also called Mann and Whitney test). h, Laser ablation experiments of apical membranes in arm::GFP leg discs in the segment domain versus the fold domain where apoptosis takes place. Discs were dissected and cultured ex vivo from pre-fold stage (WP) to mid-fold stage. Note the increase in the length of vertex release in the fold domain compared to the segment domain (n = 4). i, Laser ablation experiments of apical membranes in the fold domain of arm::GFP leg discs incubated from pre-fold stage (WP) to mid-fold stage with either DMSO (control, n = 5) of Q-VD-OPH (cell death inhibition, n = 4). Note that vertex release in the fold is reduced in the absence of apoptosis. h, i, Right panels, graphs representing quantifications of the increase in distance between vertices following laser cut, revealing apoptosis-dependent increased cellular tension in the fold domain versus the segment domain. Examples of ablated cells before (green) and after (magenta) laser cut are shown on the left. Orange bars represent the region where the laser cut has been performed. Errors bars correspond to the standard error of the mean.
Extended Data Figure 4 Cell shape dynamics during fold formation.
This figure is associated with Fig. 2. a, Three-dimensional reconstructions and schematics of leg imaginal discs from pre-fold stage (WP) and mid-fold stage (WP + 2 h) stained with E-Cad in white and cleaved Dcp1 (revealing caspase activity) in red (n = 11 for each). High magnifications of the fold domain (surrounded in red) are shown on the right hand side of each panel. Arrowheads indicate apoptotic cells. b, Cell shape dynamics during fold formation (n = 8). c, Schematic of a pupal leg disc showing the apterous domain (in red) in the t4 tarsal segment and overlapping the t4-t5 fold (top). The bottom panel shows a typical result of automated cell outline extraction of a pupal leg disc double-stained for adherens junctions and the apterous domain. Cells from this domain have been subdivided into 12 sections along the proximo-distal axis (see Methods). d, Anisotropy, area and orientation of cells from DMSO (control, n = 7) or Q-VD-OPH (cell death inhibition, n = 8) leg discs from fold domain sections 10 and 11 (see Extended Data Fig. 4c) were quantified and values represented as box plot. Discs were dissected and incubated from pre-fold stage (WP) to mid-fold stage. We used the non-parametric Wilcoxon rank sum test (also called Mann and Whitney test). e, Three-dimensional reconstruction images of anti E-Cad stained pupal leg discs from mid-fold stage (WP + 2 h) (top left) of the following genotypes: LP30::Gal4 (control, ctl, n = 11), uas-DIAP1; LP30::Gal4 (Diap1, n = 22), Dll::Gal4[MD23] / uas-p35 (p35, n = 9) and Dll::Gal4[MD23] / uas-p35; uas-p35 (p35 x2, n = 8) (at early pupal stages, LP30-Gal4 and Dll::Gal4[MD23] show similar expression patterns, namely expression in the distal tibia and in all tarsal segments27). For each condition, cell outlines were extracted and anisotropy, apical surface area and orientation of cells from the fold domain were quantified and colour-coded. Note that when cell death is inhibited, anisotropy is reduced, apical surfaces are increased and cell preferential orientation is perturbed compared to the control situation.
Extended Data Figure 5 Three-dimensional epithelial cell model.
This figure is associated with Fig. 3. a, Each cell is represented as an apical surface delimited by apical junctions. Each cell interacts with its neighbours through the apical junctions at its borders. In the original work by Farhadifar et al.15, three interactions are considered: (1) the tension opposing the elongation of a particular junction edge, with energy increasing with edge length; (2) a contractility, with energy proportional to the cell perimeter squared, used to model cell constriction; (3) a surface elasticity bringing the apical cell area back to a preferred area. As in the original work, the model can produce cell division, types one and three transitions, to which we added apoptosis. Yet in our case, contrary to Jüllicher and colleagues work15, we must also take into account non planar modifications of the epithelial sheet. To this end, we modified the elastic area interaction to take into account a constrain on cell volume. The new interaction is termed volume elasticity and transmits contractions and dilations of the apical sheet along the apical-basal axis. The associated energy is proportional to the square of the difference between the current cell volume and a preferred volume. b, Average areas of cells as a function of the number of neighbouring cells in the epithelium before apoptosis, to be compared with Fig. 2g in Farhadifar et al.15. Our tissue shows a similar trend of growing area with the number of sides, in good quantitative agreement with Farhadifar et al.15. c, Distribution of the number of neighbouring cells (or, equivalently, of the number of cell sides), to be compared with Fig. 2f of Farhadifar et al.15; once again, we are in good quantitative agreement with their model. d, Ground state diagram of the vertex model, comparing two-dimensional and three-dimensional hexagonal network boundaries (we restricted ourselves to Λ > 0 and Γ > 0 regions). The black dot indicates the chosen values for the line tension and contractility parameters, which are the same as case I in the Farhadifar et al.15 article. e, Variation of the normalized energy of a regular epithelium comprised of identical hexagonal cells as a function of a scale factor δ. Plain lines, analytical calculus; dotted line, average cell energy for a cylindrical tissue of 32 cells in diameter per 29 cells long. A scaling of δ = 1 means that the cells are at their equilibrium volume in the absence of elasticity and contractility, and thus corresponds to the minimum of the blue lines (volume elasticity). Green lines correspond to line tension and yellow lines to contractility. The discrepancy between theoretical and computed values is due to the effect of cells lying at the border of the cylindrical epithelium.
Extended Data Figure 6 Effect of the apoptosis pattern on epithelium folding in silico.
This figure is associated with Fig. 3. a, Representation of the in silico cell death pattern. Note the similarity with the in vivo distribution of apoptosis observed in biological samples (Fig. 1b). Nonetheless, the representation is not exactly comparable since cell death pattern is represented relative to a developmental stage in the biological samples, whereas in the model, the cell death pattern is represented relatively to the number of dead cells generated by the theoretical simulation since the time scale is not taken into account in the model. Left, representation of the in silico distribution of apoptotic cells around the fold domain. b–g, For each panel, from left to right are represented (1) a scheme of the fold domain showing the pattern of apoptosis, (2) a three-dimensional representation showing whole tissue shape and (3) for each condition, the corresponding cell outlines extracted from three-dimensional simulations in which anisotropy, area and orientation of cells from the fold domain are colour-coded. b–f, In silico models showing whole tissue shape with an increasing number of apoptotic cells following the in vivo pattern of apoptosis. Note the gradual increase in anisotropy, gradual decrease of cell area and the preferential orientation with the gradual increase in the number of dying cells. The three-dimensional simulations in b and f are those presented in Fig. 3a and Fig. 3c, respectively. The three-dimensional simulation in f (framed in a blue rectangle in Extended Data Fig. 6, Extended Data Fig. 7 and Extended Data Fig. 8) corresponds to 30 apoptotic cells, with an apico-basal force of 1 Λ and an increase in apical contractility of 1 Γ. g, In silico model for a random pattern of apoptosis, with all other parameters similar to f. h, The mean value of radius, anisotropy, area and orientation of cells from the whole fold domain, defined as a ± 1 µm region around the fold centre of simulations b–g are represented by box plot. The number of cells considered is n = 33, 38, 45, 61 and 98 for 0, 5, 10, 20 and 30 cells, respectively. This number varies due to changes in cells density in the domain.
Extended Data Figure 7 Effect of apico-basal force intensity on epithelium folding in silico.
This figure is associated with Fig. 3. a–d, In silico models showing whole tissue shape following increasing values of apico-basal apoptotic force (30 apoptotic cells, apical contractility: 1 Γ). For each panel, from left to right are represented (1) a scheme of the strength of the apico-basal force applied, (2) a three-dimensional representation showing whole tissue shape and (3) for each condition, the corresponding cell outlines extracted from three-dimensional simulations in which anisotropy, area and orientation of cells from the fold domain are colour-coded. Note the gradual increase in anisotropy, gradual decrease of cell area and the preferential orientation with the gradual increase of the apico-basal force. e, The mean value of radius, anisotropy, area and orientation of cells from the whole fold domain of simulations a–d are represented by box plot. The Ø symbol corresponds to the condition in absence of apoptosis (from Extended Data Fig. 6b). The number of cells considered is n = 60, 69, 98 and 157 for Λ = 0, 0.5, 1 and 2, respectively.
Extended Data Figure 8 Effect of the gradual apical cell contractibility increase on epithelium folding in silico.
This figure is associated with Fig. 3. a–d, In silico models showing whole tissue shape with an apico-basal force of 1 Λ in 30 apoptotic cells and increasing values of apical contractility in neighbouring cells. For each panel, from left to right are represented (1) a scheme representing the gradual increase of contractility values applied in apoptotic neighbours, (2) a three-dimensional representation showing whole tissue shape and (3) for each condition, the corresponding cell outlines extracted from three-dimensional simulations in which cell anisotropy, area and orientation from the fold domain are colour-coded. Note the gradual increase in anisotropy, gradual decrease of cell area and the preferential orientation with the gradual increase of contractility. e, The mean value of radius, anisotropy, area and orientation of cells from the whole fold domain of simulations a–d are represented by box plot. The Ø symbol corresponds to the condition in absence of apoptosis (from Extended Data Fig. 6b). The number of cells considered is n = 31, 61, 98 and 137 for Γ = 0, 0.5, 1 and 2, respectively.
Extended Data Figure 9 Fold formation as a result of apoptotic forces.
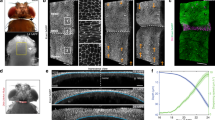
This figure is associated with Fig. 4. a–a′′, Schematics of wing discs depicting the pattern of apoptosis and the presence or absence of myosin II activity in apoptotic cells in each genetic context analysed in b–b′′′ and c–c′. Control (a, b, y,w,hs::flp; act-frt-y+-frt::Gal4, uas::GFP / uas::lifeactGFP; uas::rpr without clones), Ectopic cell death (a′, b′, b′′′, y,w,hs::flp; act-frt-y+-frt::Gal4, uas::GFP / uas::lifeactGFP; uas::rpr), and ectopic cell death without myosin II activity specifically in apoptotic cells (a′′, b′′, y,w,hs::flp; uas::DN-zip::GFP, uas::hid; act-frt-CD2-frt::Gal4). Note that, in the latter condition, myosin II activity is maintained in living cells. Wing discs were dissected from larvae heat shocked for 20 min at 38 °C. b–c′, For each panel, sagittal views and schematics of sagittal sections are shown (sagittal views correspond to the black dotted line indicated in a–a′′). b–b′′′, A high concentration of myosin II positive apoptotic cells is sufficient to induce a fold in a naive tissue (red arrowhead, b′, n = 11) as shown by the visualization of the wing disc apical surface stained with an anti-β-catenin antibody (compare b′ with b). Note that no ectopic fold is observed when only a low number of apoptotic events occur (b′′′) or when myosin II is inhibited in apoptotic cells (open arrowhead, n = 5 out of 6) (b′′). c–c′, F-actin accumulates in ectopic folds (red arrowhead) when apoptotic myosin II is active in ectopic dying cells (compare c′ with c, n = 3). d, Quantification of myosin II levels in the patch (ptc) domain (domain of cell death induction) in control (w; ptc::Gal4, uas::GFP; tub::G80[ts] / SM5-TM6B) ; ptc > rpr (w; ptc::Gal4, uas::GFP / uas::lifeactGFP; tub::G80[ts] / uas::rpr) and ptc > rpr+DN-MyoII (w; ptc::Gal4, uas::GFP / uas::DN-zip::GFP; tub::G80[ts] / uas::rpr) wing discs (control, n = 24; ptc > rpr, n = 28; ptc > rpr+DN-MyoII, n = 32). Values are represented as mean values with error bars representing standard errors. The intensity of the signal in the ptc domain has been normalized with the intensity in the anterior and posterior domains of the same disc (n.s. is for non-significant). We used the non-parametric Wilcoxon rank sum test (also called Mann and Whitney test). e–e′′, Wing disc close-ups (of wing discs shown in Fig. 4c) and schematics in the absence (e) or presence (e′ and e′′) of ectopic apoptosis in the ptc domain (red cells, false-coloured in red on the black and white images), with (e′) or without (e′′) myosin II activity in dying cells. e′, Note that we can distinguish two distinct pools of stabilized apical myosin II: “contractile ring myosin II” required for dying cell extrusion16 (blue arrows, purple in schematics) and “fold domain apical myosin II” stabilized in response to the apico-basal apoptotic force (red arrows, green in schematics). e′′, Note that, consistently with normal extrusion in this background, contractile ring myosin II is still present around apoptotic cells (blue arrows), whereas fold domain apical myosin II is absent. The star points at a dividing cell, further indicating that myosin II is still present in apoptotic cell neighbours.
Supplementary information
Cell death dynamics during fold morphogenesis
Time-lapse images of Dlg::GFP pre-fold leg disc showing t4-t5 fold progression. Cell membranes are in red (stained with FM4-64) and apoptotic cells in green (stained with acridine orange). Apoptotic cells at the tip of the leg disc correspond to the claw region and are not involved in fold formation. (MP4 3062 kb)
Adherens junction dynamics in an apoptotic cell
Time-lapse images of E-Cad-GFP (green) and alpha-catenin-TagRFP (red) fusion proteins in a pre-fold leg disc. The sagittal (XZ, bottom) and corresponding apical top (XY, top) views are shown. Note on sagittal sections that the apical surface of the epithelium is transiently bended towards the basal floor concomitantly to the appearance of an adhesion peak (20’). The apical surface then goes back to its original location as the apoptotic adhesion peak detaches and moves basally (30’ onwards). The apoptotic cell is identified by the shrinkage of its apical surface, visible on the apical top view (see also ExtData2a). The apoptotic cell is highlighted in red on the schematics on the right. (MP4 262 kb)
Myosin II dynamics in an apoptotic cell
Sagittal view highlighting Myosin II-GFP dynamics in an apoptotic cell identified by the shrinkage of its apical surface (not shown). Note that Myosin II accumulates apically (12’), then propagates towards the basal side (15’-18’) forming an apico-basal cable that precedes maximal pulling of the apical surface (21’). In following frames, the apical surface of the epithelium goes back to its original position as the apoptotic cell fragments (see also ExtData2d). (MP4 176 kb)
Cell shape changes at the vicinity of an apoptotic cell
Time-lapse images of an apoptotic cell (labelled in red) and of its direct and indirect neighbours (respectively dark and light blue), highlighting the correlation between the decrease in apoptotic cell apical surface and the stretching of neighbouring cells as well as the decrease in their apical surface. Adherens junctions are labelled with E-Cad::GFP. (MP4 143 kb)
Cell shape changes associated with epithelial folding in the leg
Time-lapse images of an E-Cad::GFP pre-fold leg disc highlighting cell shape changes associated with leg epithelium folding: progressively, cells stretch decreasing their apical surface. Cells of the future fold are false-coloured in blue. (Black dots on both sides of the fold domain correspond to cells of the nervous system). (MP4 348 kb)
Rights and permissions
About this article
Cite this article
Monier, B., Gettings, M., Gay, G. et al. Apico-basal forces exerted by apoptotic cells drive epithelium folding. Nature 518, 245–248 (2015). https://doi.org/10.1038/nature14152
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14152
This article is cited by
-
Homeotic compartment curvature and tension control spatiotemporal folding dynamics
Nature Communications (2023)
-
A role for Flower and cell death in controlling morphogen gradient scaling
Nature Cell Biology (2022)
-
Adhesion-mediated heterogeneous actin organization governs apoptotic cell extrusion
Nature Communications (2021)
-
The Actomyosin Cortex of Cells: A Thin Film of Active Matter
Journal of the Indian Institute of Science (2021)
-
Epithelial invagination by a vertical telescoping cell movement in mammalian salivary glands and teeth
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.