Abstract
Non-small-cell lung cancer is the leading cause of cancer-related death worldwide1. Chemotherapies such as the topoisomerase II (TopoII) inhibitor etoposide effectively reduce disease in a minority of patients with this cancer2,3; therefore, alternative drug targets, including epigenetic enzymes, are under consideration for therapeutic intervention4. A promising potential epigenetic target is the methyltransferase EZH2, which in the context of the polycomb repressive complex 2 (PRC2) is well known to tri-methylate histone H3 at lysine 27 (H3K27me3) and elicit gene silencing5. Here we demonstrate that EZH2 inhibition has differential effects on the TopoII inhibitor response of non-small-cell lung cancers in vitro and in vivo. EGFR and BRG1 mutations are genetic biomarkers that predict enhanced sensitivity to TopoII inhibitor in response to EZH2 inhibition. BRG1 loss-of-function mutant tumours respond to EZH2 inhibition with increased S phase, anaphase bridging, apoptosis and TopoII inhibitor sensitivity. Conversely, EGFR and BRG1 wild-type tumours upregulate BRG1 in response to EZH2 inhibition and ultimately become more resistant to TopoII inhibitor. EGFR gain-of-function mutant tumours are also sensitive to dual EZH2 inhibition and TopoII inhibitor, because of genetic antagonism between EGFR and BRG1. These findings suggest an opportunity for precision medicine in the genetically complex disease of non-small-cell lung cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jemal, A. et al. Global cancer statistics. CA Cancer J. Clin. 61, 69–90 (2011)
Zornosa, C. et al. First-line systemic therapy practice patterns and concordance with NCCN guidelines for patients diagnosed with metastatic NSCLC treated at NCCN institutions. J. Natl. Compr. Canc. Netw. 10, 847–856 (2012)
Wang, L. et al. Randomized phase II study of concurrent cisplatin/etoposide or paclitaxel/carboplatin and thoracic radiotherapy in patients with stage III non-small cell lung cancer. Lung Cancer 77, 89–96 (2012)
Baylin, S. B. & Jones, P. A. A decade of exploring the cancer epigenome – biological and translational implications. Nature Rev. Cancer 11, 726–734 (2011)
Simon, J. A. & Lange, C. A. Roles of the EZH2 histone methyltransferase in cancer epigenetics. Mutat. Res. 647, 21–29 (2008)
Shedden, K. et al. Gene expression-based survival prediction in lung adenocarcinoma: a multi-site, blinded validation study. Nature Med. 14, 822–827 (2008)
Tan, J. et al. Pharmacologic disruption of Polycomb-repressive complex 2-mediated gene repression selectively induces apoptosis in cancer cells. Genes Dev. 21, 1050–1063 (2007)
Choudhury, S. R. et al. (-)-Epigallocatechin-3-gallate and DZNep reduce polycomb protein level via a proteasome-dependent mechanism in skin cancer cells. Carcinogenesis 32, 1525–1532 (2011)
McCabe, M. T. et al. EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature 492, 108–112 (2012)
Chou, T. & Talalay, P. Quantitative analysis of dose-effect relationships: the combined effects of multiple drugs of enzyme inhibitors. Adv. Enzyme Regul. 22, 27–55 (1984)
Deweese, J. E. & Osheroff, N. The DNA cleavage reaction of topoisomerase II: wolf in sheep’s clothing. Nucleic Acids Res. 37, 738–748 (2009)
Ji, H. et al. The impact of human EGFR kinase domain mutations on lung tumorigenesis and in vivo sensitivity to EGFR-targeted therapies. Cancer Cell 9, 485–495 (2006)
Jackson, E. L. et al. The differential effects of mutant p53 alleles on advanced murine lung cancer. Cancer Res. 65, 10280–10288 (2005)
Dykhuizen, E. C. et al. BAF complexes facilitate decatenation of DNA by topoisomerase II. Nature 497, 624–627 (2013)
Wilson, B. G. et al. Epigenetic antagonism between Polycomb and SWI/SNF complexes during oncogenic transformation. Cancer Cell 18, 316–328 (2010)
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012)
TCGA. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014)
ENCODE. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2013)
Hargreaves, D. C. & Crabtree, G. R. ATP-dependent chromatin remodeling: genetics, genomics and mechanisms. Cell Res. 21, 396–420 (2011)
Oike, T. et al. A synthetic lethality-based strategy to treat cancers harboring a genetic deficiency in the chromatin remodeling factor BRG1. Cancer Res. 73, 5508–5518 (2013)
Matsubara, D. et al. Lung cancer with loss of BRG1/BRM, shows epithelial mesenchymal transition phenotype and distinct histologic and genetic features. Cancer Sci. 104, 266–273 (2013)
Medina, P. P. et al. Frequent BRG1/SMARCA4-inactivating mutations in human lung cancer cell lines. Hum. Mutat. 29, 617–622 (2008)
Yamamoto, H. et al. PIK3CA mutations and copy number gains in human lung cancers. Cancer Res. 68, 6913–6921 (2008)
Bamford, S. et al. The COSMIC (Catalogue of Somatic Mutations in Cancer) database and website. Br. J. Cancer 91, 355–358 (2004)
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012)
Orimo, A. et al. Stromal fibroblasts present in invasive human breast carcinomas promote tumor growth and angiogenesis through elevated SDF-1/CXCL12 Secretion. Cell 121, 335–348 (2005)
Ramirez-Carrozzi, V. R. et al. Selective and antagonistic functions of SWI/SNF and Mi-2 nucleosome remodeling complexes during an inflammatory response. Genes Dev. 20, 282–296 (2006)
Xi, Q., He, W., Zhang, X. H.-F., Le, H.-V. & Massagué, J. Genome-wide impact of the BRG1 SWI/SNF chromatin remodeler on the transforming growth factor β transcriptional program. J. Biol. Chem. 283, 1146–1155 (2008)
Engelman, J. A. et al. Allelic dilution obscures detection of a biologically significant resistance mutation in EGFR-amplified lung cancer. J. Clin. Invest. 116, 2695–2706 (2006)
Fillmore, C. M. et al. Estrogen expands breast cancer stem-like cells through paracrine FGF/Tbx3 signaling. Proc. Natl Acad. Sci. USA 107, 21737–21742 (2010)
Zacharek, S. J. et al. Lung stem cell self-renewal relies on bmi1-dependent control of expression at imprinted loci. Cell Stem Cell 9, 272–281 (2010)
Chou, T.-C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70, 440–446 (2010)
Chou, T.-C. Theoretical basis, experimental design, and computerized simulation of synergism and antagonism in drug combination studies. Pharmacol. Rev. 58, 621–681 (2006)
Rhodes, D. R. et al. Oncomine 3.0: genes, pathways, and networks in a collection of 18,000 cancer gene expression profiles. Neoplasia 9, 166–180 (2007)
Beer, D. G. et al. Gene-expression profiles predict survival of patients with lung adenocarcinoma. Nature Med. 8, 816–824 (2002)
Garber, M. E. et al. Diversity of gene expression in adenocarcinoma of the lung. Proc. Natl Acad. Sci. USA 98, 13784–13789 (2001)
Gordon, G. J. et al. Translation of microarray data into clinically relevant cancer diagnostic tests using gene expression ratios in lung cancer and mesothelioma. Cancer Res. 62, 4963–4967 (2002)
Landi, M. T. et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS ONE 3, e1651 (2008)
Rohrbeck, A. et al. Gene expression profiling for molecular distinction and characterization of laser captured primary lung cancers. J. Transl. Med. 6, 69 (2008)
Su, A. I. et al. Molecular classification of human carcinomas by use of gene expression signatures. Cancer Res. 61, 7388–7393 (2001)
Yu, K. et al. A precisely regulated gene expression cassette potently modulates metastasis and survival in multiple solid cancers. PLoS Genet. 4, e1000129 (2008)
Lockwood, W. W. et al. DNA amplification is a ubiquitous mechanism of oncogene activation in lung and other cancers. Oncogene 27, 4615–4624 (2008)
Shankavaram, U. T. et al. Transcript and protein expression profiles of the NCI-60 cancer cell panel: an integromic microarray study. Mol. Cancer Ther. 6, 820–832 (2007)
Su, L.-J. et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics 8, 140 (2007)
Miyanaga, A. et al. Antitumor activity of histone deacetylase inhibitors in non-small cell lung cancer cells: development of a molecular predictive model. Mol. Cancer Ther. 7, 1923–1930 (2008)
Balko, J. et al. Gene expression patterns that predict sensitivity to epidermal growth factor receptor tyrosine kinase inhibitors in lung cancer cell lines and human lung tumors. BMC Genomics 7, 289 (2006)
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003)
Smyth, G. K., Michaud, J., l & Scott, H. S. Use of within-array replicate spots for assessing differential expression in microarray experiments. Bioinformatics 21, 2067–2075 (2005)
Hochberg, Y. & Benjamini, Y. More powerful procedures for multiple hypothesis testing. Stat. Med. 9, 811–818 (1990)
Li, C. & Wong, W. H. Model-based analysis of oligonucleotide arrays: Expression index computation and outlier detection. Proc. Natl Acad. Sci. USA 98, 31–36 (2001)
Curtis, S. J. et al. Primary tumor genotype is an important determinant in identification of lung cancer propagating cells. Cell Stem Cell 7, 127–133 (2010)
Acknowledgements
We thank the Kim laboratory, F. Luo, P. Louis, K. Harrington, X. Wang and J. Brainson for technical assistance and discussions, and J. Crabtree, D. Hargreaves, C. Kadoch, L. Zon, K. Cichowski, M. Enos, S. Orkin, A. Gutierrez and C. Roberts for discussions. This work was supported in part by the Ladies Auxiliary to the Veterans of Foreign Wars, PF-12-151-01-DMC from the American Cancer Society, and the Uniting Against Lung Cancer Young Investigator Award supported by Meryl Bralower (C.M.F.), Boston University Undergraduate Research Opportunities Program (P.T.D.), RO1 HL090136, U01 HL100402 RFA-HL-09-004, American Cancer Society Research Scholar Grant RSG-08-082-01-MGO, the V Foundation for Cancer Research, a Basil O’Conner March of Dimes Starter Award, the Harvard Stem Cell Institute, and the Lung Cancer Research Foundation (C.F.K.), the National Institutes of Health (NIH) grants CA122794, CA140594, CA163896, CA166480, CA154303 and CA120964 (K.K.W.), the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research (V.E.M.), and the NIH grant K08 CA163677 (P.S.H.).
Author information
Authors and Affiliations
Contributions
C.M.F., C.X., K.K.W. and C.F.K. designed the study; C.M.F., C.X., P.T.J., J.M.B. and Y.J.L. performed the experiments; S.P.R. cloned the EZH2 complementary DNA (cDNA) vector; H.Z. performed EZH2 immunohistochemistry, V.E.M. provided DZNep, P.S.H. analysed primary tumour sequencing data; K.K.W. allowed autochthonous mouse models studies in his laboratory; C.M.F. and C.F.K. wrote the manuscript with comments from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Verification of EZH2 as a potential target for NSCLC.
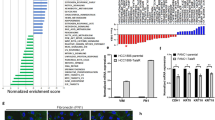
a, Survival of patients with lung adenocarcinoma in the Director’s Challenge data set. Samples were hierarchically clustered using the primary-tumour-generated EZH2 co-expression signature (Supplementary Table 1) into two risk groups. The Kaplan–Meier curve for the whole data set is shown (n = 416, P < 0.00001). b, Western blot was performed on whole-cell extracts from indicated lines for EZH2 and its catalytic mark H3K27me3; total histone H3 is shown as loading control. CR indicates a coding region targeting hairpin. c, RT-qPCR for average expression of EZH2 in the indicated cell lines after plating at equal density and treating for 4 days with indicated treatments. Each cell line is normalized to its shGFP control (n = 2 biological replicates).
Extended Data Figure 2 Pharmacological inhibition of EZH2 changes response of cells to TopoII inhibitors.
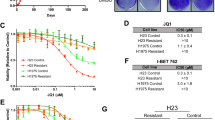
a, Western blot for EZH2 and H3K27me3 was performed on whole-cell extracts after administration of 1 µM DZNep for 4 days, 10 µM GSK126 for 4 days, 2 µM GSK126 for 9 days, or vehicle. Total histone H3 is shown as a loading control. b, The fold change in etoposide IC50 ± s.e.m. in response to DZNep is plotted (n = 3 biological replicates for H1975, H2030, HCC4006, A549, HCC2450, Calu1, H1650, H522, H2126, H1299, HCC15, H322, H2009, HCC95, H520, H460, Calu3, H2122, H23 and H3255; n = 4 biological replicates for PC9, H157, HCC827, Sw1573, Calu6 and H441; P < 0.02). Cell lines with mutations in BRG1 or EGFR are indicated. Note that the H23 cell line has a very late coding region mutation in BRG1 (K1533N) and is predicted to produce functional protein22, consistent with its protected phenotype in our assays. c, Fold change ± s.e.m. between vehicle-treated and indicated EZH2i-treated lines for etoposide IC50 is plotted (n = 3 biological replicates; P < 0.03, P < 0.01). d, Fold change in doxorubicin IC50 in response to DZNep (n = 2 biological replicates).
Extended Data Figure 3 Xenograft experiments confirm sensitized and protected phenotypes.
a, Representative image of mouse injected at four sites (arrows) with H23 tumour cells 12 days after cell injection. b, Representative images of mice injected at four sites with either H23 or H157 cells, and treated with indicated drugs, 35 days after cell injection. Palpable tumours that remain are indicated with arrows.
Extended Data Figure 4 Autochthonous mouse models confirm genotype specificity of dual EZH2i and TopoIIi.
a, Representative images of haematoxylin and eosin stained lung from EGFRT790M;L858R mice treated with indicated therapies for 4 weeks. Areas with tumours of similar sizes were chosen for comparison; scale bar, 200 µm. b, Histology from KrasG12D/+/p53Δ/Δ mouse lung tumours after 1 week of indicated treatments; top image is haematoxylin and eosin, bottom image is EZH2 immunohistochemistry; scale bar, 100 µm. c, Histology from EGFRT790M;L858R mouse lung tumours after 4 weeks of indicated treatments; top image is haematoxylin and eosin, centre image is phospho-EGFR immunohistochemistry and bottom image is EZH2 immunohistochemistry; scale bar, 100 µm.
Extended Data Figure 5 EZH2i modulates anaphase bridging differentially by genotype.
a, Representative images of nuclei undergoing a normal anaphase and of nuclei that scored positively for the presence of anaphase bridges. b, Average percentage of anaphase bridging ± s.e.m. in additional BRG1 WT H2009 and H441, BRG1 mutant H522, and EGFR mutant H1650 cell lines (n = 3 biological replicates for all except H1650 (n = 4); P < 0.05). c, Immunofluorescence on PC9 cultures showing increase in BRG1 staining in interphase nuclei in response to EZH2i while anaphase nuclei retain strong EGFR stain, representative of three biological replicates; scale bar, 30 µm.
Extended Data Figure 6 Cell cycle and apoptosis analysis of dual EZH2i- and TopoIIi-treated lines.
a, 7AAD cell cycle flow cytometry on cultures corresponding to each experiment shown in Fig. 3d. The average percentage S phase ± s.e.m. of each culture is plotted (n = 3 biological replicates for H460, H23, Calu6, PC9, HCC15, A549 and H522; n = 4 biological replicates for Sw1573, H441 and H157).
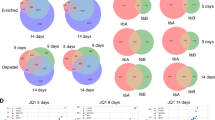
Extended Data Figure 7 EGFR and BRG1 negatively correlate in NSCLC.
a, Additional NSCLC cell lines with known EGFR and BRG1 mutations used to estimate mutually exclusivity of the two mutations. b, Venn diagram of differential gene expression overlap between cell lines of various genotypes. c, Average probe intensity ± s.e.m. of EGFR probe (201983_s_at) and EZH2 probe (203358_s_at) on the U133A Affymetrix array for cell lines with various EGFR and BRG1 mutational statuses (n = 6 per genotype, see Methods; P = 0.014).
Extended Data Figure 8 Modulation of EGFR and BRG1 influences sensitized and protected phenotypes.
a, RT-qPCR for average expression of BRG1, EGFR and EZH2 ± s.e.m. in the various indicated treated transduced cell lines (n = 3 biological replicates). b, For the indicated HCC15 and H460 stably transduced etoposide-treated cell lines, 7AAD flow cytometry was used to assess average changes in percentage S phase ± s.e.m. in response to DZNep (n = 3 biological replicates; P = 0.02, P < 0.001). c, Average percentage sub-G1 fractions ± s.e.m. of the indicated 4-day cultures were assessed during 7AAD cell cycle flow cytometry analysis. Critically, for these assays, the supernatant of each culture was retained and combined with the trypsinized adherent cells to reflect the total amount of apoptosis/necrosis in each culture accurately (n = 3 biological replicates; P = 0.03).
Extended Data Figure 9 Confirmation that BRG1 re-expression leads to formation of BAF complex.
a, Immunoprecipitation of BAF complex members from nuclear lysates of the (left) BRG1 mutant HCC15 shGFP control cell line and (right) the HCC15 line with BRG1 re-expressed shows that exogenously expressed Flag-tagged BRG1 does result in BRG1-containing BAF complex formation. The blot is representative of three biological replicates.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5. (PDF 389 kb)
Rights and permissions
About this article
Cite this article
Fillmore, C., Xu, C., Desai, P. et al. EZH2 inhibition sensitizes BRG1 and EGFR mutant lung tumours to TopoII inhibitors. Nature 520, 239–242 (2015). https://doi.org/10.1038/nature14122
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14122
This article is cited by
-
The innovative checkpoint inhibitors of lung adenocarcinoma, cg09897064 methylation and ZBP1 expression reduction, have implications for macrophage polarization and tumor growth in lung cancer
Journal of Translational Medicine (2024)
-
A rare Ewing-like small round cell tumor in prostate: a case report and literature review
Journal of Cancer Research and Clinical Oncology (2024)
-
The role of chromatin remodeler SMARCA4/BRG1 in brain cancers: a potential therapeutic target
Oncogene (2023)
-
Multifunctional Nanocarriers for Alzheimer’s Disease: Befriending the Barriers
Molecular Neurobiology (2023)
-
Effect of chromatin modifiers on the plasticity and immunogenicity of small-cell lung cancer
Experimental & Molecular Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.