Abstract
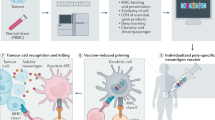
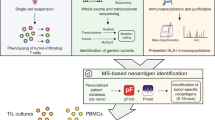
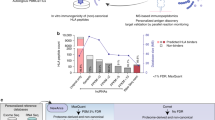
Human tumours typically harbour a remarkable number of somatic mutations1. If presented on major histocompatibility complex class I molecules (MHCI), peptides containing these mutations could potentially be immunogenic as they should be recognized as ‘non-self’ neo-antigens by the adaptive immune system. Recent work has confirmed that mutant peptides can serve as T-cell epitopes2,3,4,5,6,7,8,9. However, few mutant epitopes have been described because their discovery required the laborious screening of patient tumour-infiltrating lymphocytes for their ability to recognize antigen libraries constructed following tumour exome sequencing. We sought to simplify the discovery of immunogenic mutant peptides by characterizing their general properties. We developed an approach that combines whole-exome and transcriptome sequencing analysis with mass spectrometry to identify neo-epitopes in two widely used murine tumour models. Of the >1,300 amino acid changes identified, ∼13% were predicted to bind MHCI, a small fraction of which were confirmed by mass spectrometry. The peptides were then structurally modelled bound to MHCI. Mutations that were solvent-exposed and therefore accessible to T-cell antigen receptors were predicted to be immunogenic. Vaccination of mice confirmed the approach, with each predicted immunogenic peptide yielding therapeutically active T-cell responses. The predictions also enabled the generation of peptide–MHCI dextramers that could be used to monitor the kinetics and distribution of the anti-tumour T-cell response before and after vaccination. These findings indicate that a suitable prediction algorithm may provide an approach for the pharmacodynamic monitoring of T-cell responses as well as for the development of personalized vaccines in cancer patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013)
Matsushita, H. et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature 482, 400–404 (2012)
DuPage, M., Mazumdar, C., Schmidt, L. M., Cheung, A. F. & Jacks, T. Expression of tumour-specific antigens underlies cancer immunoediting. Nature 482, 405–409 (2012)
Castle, J. C. et al. Exploiting the mutanome for tumor vaccination. Cancer Res. 72, 1081–1091 (2012)
van Rooij, N. et al. Tumor exome analysis reveals neoantigen-specific T-cell reactivity in an ipilimumab-responsive melanoma. J. Clin. Oncol. 31, e439–e442 (2013)
Gros, A. et al. PD-1 identifies the patient-specific CD8+ tumor-reactive repertoire infiltrating human tumors. J. Clin. Invest. 124, 2246–2259 (2014)
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014)
Wick, D. A. et al. Surveillance of the tumor mutanome by T cells during progression from primary to recurrent ovarian cancer. Clin. Cancer Res. 20, 1125–1134 (2014)
Brown, S. D. et al. Neo-antigens predicted by tumor genome meta-analysis correlate with increased patient survival. Genome Res. 24, 743–750 (2014)
Chen, D. S. & Mellman, I. Oncology meets immunology: the cancer-immunity cycle. Immunity 39, 1–10 (2013)
Lundegaard, C. et al. NetMHC-3.0: accurate web accessible predictions of human, mouse and monkey MHC class I affinities for peptides of length 8–11. Nucleic Acids Res. 36, W509–W512 (2008)
Gujar, S. A., Pan, D. A., Marcato, P., Garant, K. A. & Lee, P. W. Oncolytic virus-initiated protective immunity against prostate cancer. Mol. Ther. 19, 797–804 (2011)
Fortier, M. H. et al. The MHC class I peptide repertoire is molded by the transcriptome. J. Exp. Med. 205, 595–610 (2008)
Granados, D. P. et al. MHC I-associated peptides preferentially derive from transcripts bearing miRNA response elements. Blood 119, e181–e191 (2012)
Barbieri, C. E. et al. Exome sequencing identifies recurrent SPOP, FOXA1 and MED12 mutations in prostate cancer. Nature Genet. 44, 685–689 (2012)
Bertsch, E. et al. MED12 and HMGA2 mutations: two independent genetic events in uterine leiomyoma and leiomyosarcoma. Mod. Pathol. 27, 1144–1153 (2014)
Goldberg, A. L. Functions of the proteasome: from protein degradation and immune surveillance to cancer therapy. Biochem. Soc. Trans. 35, 12–17 (2007)
Sette, A. et al. The relationship between class I binding affinity and immunogenicity of potential cytotoxic T cell epitopes. J. Immunol. 153, 5586–5592 (1994)
London, N., Raveh, B., Cohen, E., Fathi, G. & Schueler-Furman, O. Rosetta FlexPepDock web server–high resolution modeling of peptide-protein interactions. Nucleic Acids Res. 39, W249–W253 (2011)
Rudolph, M. G., Stanfield, R. L. & Wilson, I. A. How TCRs bind MHCs, peptides, and coreceptors. Annu. Rev. Immunol. 24, 419–466 (2006)
Jin, H. T. et al. Cooperation of Tim-3 and PD-1 in CD8 T-cell exhaustion during chronic viral infection. Proc. Natl Acad. Sci. USA 107, 14733–14738 (2010)
Jin, H. T., Ahmed, R. & Okazaki, T. Role of PD-1 in regulating T-cell immunity. Curr. Top. Microbiol. Immunol. 350, 17–37 (2011)
Wherry, E. J. T cell exhaustion. Nature Immunol. 12, 492–499 (2011)
Fourcade, J. et al. Upregulation of Tim-3 and PD-1 expression is associated with tumor antigen-specific CD8+ T cell dysfunction in melanoma patients. J. Exp. Med. 207, 2175–2186 (2010)
Fourcade, J. et al. CD8+ T cells specific for tumor antigens can be rendered dysfunctional by the tumor microenvironment through upregulation of the inhibitory receptors BTLA and PD-1. Cancer Res. 72, 887–896 (2012)
Crespo, J., Sun, H., Welling, T. H., Tian, Z. & Zou, W. T cell anergy, exhaustion, senescence, and stemness in the tumor microenvironment. Curr. Opin. Immunol. 25, 214–221 (2013)
Falk, K., Rotzschke, O., Stevanovic, S., Jung, G. & Rammensee, H. G. Allele-specific motifs revealed by sequencing of self-peptides eluted from MHC molecules. Nature 351, 290–296 (1991)
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010)
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26, 2069–2070 (2010)
Butler, N. S. et al. Structural and biological basis of CTL escape in coronavirus-infected mice. J. Immunol. 180, 3926–3937 (2008)
Young, A. C., Zhang, W., Sacchettini, J. C. & Nathenson, S. G. The three-dimensional structure of H-2Db at 2.4 Å resolution: implications for antigen-determinant selection. Cell 76, 39–50 (1994)
Denton, A. E. et al. Affinity thresholds for naive CD8+ CTL activation by peptides and engineered influenza A viruses. J. Immunol. 187, 5733–5744 (2011)
Ostrov, D. A. et al. How H13 histocompatibility peptides differing by a single methyl group and lacking conventional MHC binding anchor motifs determine self-nonself discrimination. J. Immunol. 168, 283–289 (2002)
Wang, B. et al. Peptidic termini play a significant role in TCR recognition. J. Immunol. 169, 3137–3145 (2002)
Zhao, R., Loftus, D. J., Appella, E. & Collins, E. J. Structural evidence of T cell xeno-reactivity in the absence of molecular mimicry. J. Exp. Med. 189, 359–370 (1999)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004)
Acknowledgements
The authors thank A. Bruce and J. Murphy for excellent assistance with artwork.
Author information
Authors and Affiliations
Contributions
M.Y. was involved in planning and performing in vivo experiments, analysing and interpreting data, and writing the manuscript. S.J. analysed and interpreted whole-exome sequencing and RNA sequencing data, generated translated FASTA database, searched for potential neo-epitopes. Q.T.P. and T.K.C. performed mass spectrometric data analysis and peptide validation. P.L. performed the structure prediction of the MHCI–peptide complexes. J.T. performed studies with tumour-bearing mice. S.B. performed and analysed FACS studies on tumour lines. C.F. performed and analysed immunizations experiments. Z.M. oversaw RNA sequencing experiments. I.M. assisted with the study design and the preparation of the manuscript. J.F. and T.W. performed MHCI peptide isolation and mass spectrometric analysis. L.D and J.R.L. oversaw all the work performed, planned experiments, interpreted data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
Mahesh Yadav, Suchit Jhunjhunwala, Qui T. Phung, Patrick Lupardus, Joshua Tanguay, Stephanie Bumbaca, Christian Franci, Tommy K. Cheung, Zora Modrusan, Ira Mellman, Jennie R. Lill, and Lélia Delamarre were employees of Genentech at the time of the work. Jens Fritsche and Toni Weinschenk were employees of Immatics Biotechnologies GmbH at the time of the work. They hence declare competing financial interests.
Extended data figures and tables
Extended Data Figure 1 TRAMP-C1 tumours are less immunogenic than MC-38 tumours.
a, MC-38 or TRAMP-C1 tumours (150–250 mm3 in size) were grown in vivo in C57BL/6 mice. TILs were collected and screened by flow cytometry to quantitate absolute numbers of CD45+ immune cells or CD8 T cells in each tumour sample. n = 9 for MC-38 and n = 5 for TRAMP-C1. **P ≤ 0.01, ***P ≤ 0.001 by two-tailed unpaired Student’s t-test. b, MHCI surface expression on MC-38 and TRAMP-C1 cell lines. Cells grown in vitro were stained with anti-H-2Db (biotinylated) or anti-H-2Kb (PE-conjugated) antibodies or isotype controls and analysed for surface expression by flow cytometry.
Extended Data Figure 2 Tandem mass spectra of endogenous mutant MC-38 MHC antigen peptides and their corresponding synthetic peptides.
a, AALLNSAVL; b, AQLANDVVL; c, ASM(ox)TNM(ox)ELM(ox), ox, oxidized; d, MAPIDHTTM; e, SSPYSLHYL; f, SIIVFNLL; and g, EEKNTGLI. The peak at 460.9511 m/z corresponds to a co-eluting, singly charged contaminant background ion isolated with peptide precursor (EEKNTGLI is an example of a low-confidence spectral identification, based on comparing the endogenous and synthetic spectra). Mutation sites are indicated with the amino acid underlined. Spectral differences in endogenous and synthetic peptide spectra maybe due to co-eluting peptides in the highly complex sample. Although present, not all fragment ions are labelled due to overlapping peak labels.
Extended Data Figure 3 Variant peptide expression is retained in vivo for the chosen peptides.
Total RNA-seq reads covering the variant position for the genes corresponding to the 7 MS/MS-identified variant peptides are shown. Both the reference and the alternate alleles are shown for an in vitro and an in vivo sample of the MC-38 line. Expression of the variant allele is observable both in vitro and in vivo. Alt, alternate allele; Ref, reference allele.
Extended Data Figure 4 Tandem mass spectra of endogenous wild-type MC-38 MHC antigen peptides and their corresponding synthetic peptides.
a–c, Wild-type MC-38 peptides AQLPNDVVL (a), ASM(ox)TNRELM(ox), ox, oxidized (b), and SSISNFQAV (c). Spectral differences in endogenous and synthetic peptide spectra maybe due to co-eluting peptides in the highly complex sample. Although present, not all fragment ions are labelled due to overlapping peak labels.
Extended Data Figure 5 wild-type-counterpart peptides for Adpgk and Reps1 are not immunogenic in vivo.
a–d, C57BL/6 mice were immunized with 100 µg wild-type or mutant Adpgk or Reps1 peptides with adjuvant (100 µg Poly (I:C) plus 50 µg anti-CD40) or adjuvant alone, at day 0 and day 14. At day 21, splenic CD8 T cells from mice immunized with wild-type (GIPVHLELASMTNRELMSSIVHQQVFPT) or mutant (GIPVHLELASMTNMELMSSIVHQQVFPT) Adpgk peptide were stained with mut-Adpgk–H-2Db dextramers (a) or WT-Adpgk–H-2Db dextramers (b). Similarly CD8 T cells from mice immunized with wild-type (GRVLELFRAAQLPNDVVLQIMELCGATR) or mutant Reps1 (GRVLELFRAAQLANDVVLQIMELCGATR) peptide were stained with Mut-Reps1/H-2Db dextramers (c) or WT-Reps1/H-2Db dextramers (d) and analysed by flow cytometry. Frequency of indicated peptide–MHCI dextramer+ of CD8 T cells is shown. Mice immunized with adjuvant alone were used as the reference. Data are from one experiment with n = 5 each group with bars representing means. *P ≤ 0.05 (unpaired two-tailed Student’s t-test). e, CD8 TILs from MC-38 or TRAMP-C1 tumours (300–500 mm3) grown in vivo in C57BL/6 mice were stained with PE-labelled Adpgk–H-2Db dextramers and analysed using flow cytometry. Frequency of Adpgk dextramer+ among CD8 T cells is shown in two different tumour samples.
Extended Data Figure 6 Frequency of peptide-specific CD8 T cells in MC-38 tumours of varying size grown in vivo in C57BL/6 mice.
a, b, CD8 T cells in TILs from tumours of indicated sizes were stained with PE-labelled pooled peptide–MHCI dextramers (Adpgk+Reps1+Dpagt1) and analysed by flow cytometry. The graphs are showing percent of peptide–MHCI dextramer+ among CD8 T cells (a) and the frequency of CD8 T cells among CD45+ cells (b) in TILs. Each point represents an individual mouse and the bars represent means.
Extended Data Figure 7 Frequency of Reps1- and Dpagt1-specific CD8 T cells in blood after immunization with peptides.
Blood CD8 T cells from C57BL/6 mice were analysed a day before inoculation (at day −1) with MC-38 tumours, after immunization with vaccine as outlined in Fig. 4a. The frequencies of peptide-specific CD8 T cells in the adjuvant alone (Adj) or adjuvant plus peptides (Adj+Pep) groups are shown by staining CD8 T cells with indicated peptide–MHCI dextramers and analysed using flow cytometry. The two mice that developed tumours in Adj+Pep group (in Fig. 4b) are highlighted in red filled circles. Pooled data from two experiments are shown with the bars representing means.
Extended Data Figure 8 Frequency of Reps1 and Dpagt1-specific CD8 T cells in spleen after immunization of MC-38 tumour bearing or non-tumour-bearing mice.
Tumour bearing or non-tumour-bearing C57BL/6 mice were immunized with adjuvant (anti-CD40 plus poly(I:C) alone (Adj) or 50 μg of Reps1, Dpagt1 and Adpgk peptides with adjuvant (Adj+Pep)) as outlined in Fig. 4d. a, b, Seven days after immunization splenic CD8 T cells were analysed for peptide-specific CD8 T cells by staining with Reps1–H2-Db-specific (a) or Dpagt1–H2-Kb-specific (b) dextramers and analysed using flow cytometry. Two groups of non-tumour-bearing mice were also immunized and splenic CD8 T cells were analysed at day 7 after immunization. Data are representative of two independent experiments and bars represent means.
Extended Data Figure 9 Increased frequency of Adpgk-specific CD8 T cells and IFN-γ-producing CD4 T cells upon immunization of MC-38 tumour-bearing mice.
a, Frequency of Adpgk–H-2Db dextramer+ CD8 T cells among total live cells in MC-38 tumours. TILs were analysed after immunization of MC-38 tumour bearing C57BL/6 mice with adjuvant alone (Adj) or adjuvant with peptides (Adj+Pep) or no treatment (Control) (as in Fig. 4d). Tumour CD8 T cells were stained with PE-labelled Adpgk–H-2Db dextramers and analysed using flow cytometry. Frequency of Adpgk–H-2Db dextramer+ CD8 T cells of total live cells in tumours is shown. b, Frequency of IFN-γ expressing CD4 T cells in tumours or spleen at day 7-post immunization. As in Fig. 4i, TILs or splenocytes were stimulated with PMA and ionomycin and IFNγ production among CD4 T cells was determined by intracellular cytokine staining. Data are representative of two independent experiments with bars representing means.
Supplementary information
Supplementary Table 1
This table contains variant peptides (MC-38). (XLS 353 kb)
Supplementary Table 2
This table contains variant peptides (TRAMP-C1). (XLS 35 kb)
Supplementary Table 3
This table contains peptides identified by LC-MS. (XLS 1131 kb)
Supplementary Table 4
This table contains RNA-Seq-based expression data for all genes in MC-38 and TRAMP-C1. (XLS 5281 kb)
Supplementary Table 5
This table contains MHC Class I-presented variant peptides. (XLS 30 kb)
Rights and permissions
About this article
Cite this article
Yadav, M., Jhunjhunwala, S., Phung, Q. et al. Predicting immunogenic tumour mutations by combining mass spectrometry and exome sequencing. Nature 515, 572–576 (2014). https://doi.org/10.1038/nature14001
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14001
This article is cited by
-
Assessment of human leukocyte antigen-based neoantigen presentation to determine pan-cancer response to immunotherapy
Nature Communications (2024)
-
Decoding cancer insights: recent progress and strategies in proteomics for biomarker discovery
Journal of Proteins and Proteomics (2024)
-
MHC II immunogenicity shapes the neoepitope landscape in human tumors
Nature Genetics (2023)
-
DNMT and HDAC inhibition induces immunogenic neoantigens from human endogenous retroviral element-derived transcripts
Nature Communications (2023)
-
DNA based neoepitope vaccination induces tumor control in syngeneic mouse models
npj Vaccines (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.