Abstract
The TMEM16 family of proteins, also known as anoctamins, features a remarkable functional diversity. This family contains the long sought-after Ca2+-activated chloride channels as well as lipid scramblases and cation channels. Here we present the crystal structure of a TMEM16 family member from the fungus Nectria haematococca that operates as a Ca2+-activated lipid scramblase. Each subunit of the homodimeric protein contains ten transmembrane helices and a hydrophilic membrane-traversing cavity that is exposed to the lipid bilayer as a potential site of catalysis. This cavity harbours a conserved Ca2+-binding site located within the hydrophobic core of the membrane. Mutations of residues involved in Ca2+ coordination affect both lipid scrambling in N. haematococca TMEM16 and ion conduction in the Cl− channel TMEM16A. The structure reveals the general architecture of the family and its mode of Ca2+ activation. It also provides insight into potential scrambling mechanisms and serves as a framework to unravel the conduction of ions in certain TMEM16 proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Milenkovic, V. M., Brockmann, M., Stohr, H., Weber, B. H. & Strauss, O. Evolution and functional divergence of the anoctamin family of membrane proteins. BMC Evol. Biol. 10, 319 (2010)
Pedemonte, N. & Galietta, L. J. Structure and function of TMEM16 proteins (anoctamins). Physiol. Rev. 94, 419–459 (2014)
Yang, Y. D. et al. TMEM16A confers receptor-activated calcium-dependent chloride conductance. Nature 455, 1210–1215 (2008)
Caputo, A. et al. TMEM16A, a membrane protein associated with calcium-dependent chloride channel activity. Science 322, 590–594 (2008)
Schroeder, B. C., Cheng, T., Jan, Y. N. & Jan, L. Y. Expression cloning of TMEM16A as a calcium-activated chloride channel subunit. Cell 134, 1019–1029 (2008)
Pifferi, S., Cenedese, V. & Menini, A. Anoctamin 2/TMEM16B: a calcium-activated chloride channel in olfactory transduction. Exp. Physiol. 97, 193–199 (2012)
Ferrera, L., Zegarra-Moran, O. & Galietta, L. J. Ca2+-activated Cl– channels. Compr. Physiol. 1, 2155–2174 (2011)
Huang, F., Wong, X. & Jan, L. Y. International Union of Basic and Clinical Pharmacology. LXXXV: calcium-activated chloride channels. Pharmacol. Rev. 64, 1–15 (2012)
Billig, G. M., Pal, B., Fidzinski, P. & Jentsch, T. J. Ca2+-activated Cl– currents are dispensable for olfaction. Nature Neurosci. 14, 763–769 (2011)
Yang, H. et al. TMEM16F forms a Ca2+-activated cation channel required for lipid scrambling in platelets during blood coagulation. Cell 151, 111–122 (2012)
Martins, J. R. et al. Anoctamin 6 is an essential component of the outwardly rectifying chloride channel. Proc. Natl Acad. Sci. USA 108, 18168–18172 (2011)
Suzuki, J., Umeda, M., Sims, P. J. & Nagata, S. Calcium-dependent phospholipid scrambling by TMEM16F. Nature 468, 834–838 (2010)
Suzuki, J. et al. Calcium-dependent phospholipid scramblase activity of TMEM16 protein family members. J. Biol. Chem. 288, 13305–13316 (2013)
Malvezzi, M. et al. Ca2+-dependent phospholipid scrambling by a reconstituted TMEM16 ion channel. Nature Commun. 4, 2367 (2013)
Kunzelmann, K. et al. Molecular functions of anoctamin 6 (TMEM16F): a chloride channel, cation channel, or phospholipid scramblase? Pflugers Arch. 466, 407–414 (2014)
Kuruma, A. & Hartzell, H. C. Bimodal control of a Ca2+-activated Cl− channel by different Ca2+ signals. J. Gen. Physiol. 115, 59–80 (2000)
Ni, Y. L., Kuan, A. S. & Chen, T. Y. Activation and inhibition of TMEM16A calcium-activated chloride channels. PLoS ONE 9, e86734 (2014)
Yu, K., Duran, C., Qu, Z., Cui, Y. Y. & Hartzell, H. C. Explaining calcium-dependent gating of anoctamin-1 chloride channels requires a revised topology. Circ. Res. 110, 990–999 (2012)
Terashima, H., Picollo, A. & Accardi, A. Purified TMEM16A is sufficient to form Ca2+-activated Cl– channels. Proc. Natl Acad. Sci. USA 110, 19354–19359 (2013)
Menon, I. et al. Opsin is a phospholipid flippase. Curr. Biol. 21, 149–153 (2011)
Tien, J. et al. A comprehensive search for calcium binding sites critical for TMEM16A calcium-activated chloride channel activity. eLife 3, e02772 (2014)
Fallah, G. et al. TMEM16A(a)/anoctamin-1 shares a homodimeric architecture with CLC chloride channels. Mol. Cell. Proteomics 10, M110.004697 (2011)
Sheridan, J. T. et al. Characterization of the oligomeric structure of the Ca2+-activated Cl– channel Ano1/TMEM16A. J. Biol. Chem. 286, 1381–1388 (2011)
Tien, J., Lee, H. Y., Minor, D. L., Jr, Jan, Y. N. & Jan, L. Y. Identification of a dimerization domain in the TMEM16A calcium-activated chloride channel (CaCC). Proc. Natl Acad. Sci. USA 110, 6352–6357 (2013)
Hartzell, H. C., Yu, K., Xiao, Q., Chien, L. T. & Qu, Z. Anoctamin/TMEM16 family members are Ca2+-activated Cl− channels. J. Physiol. (Lond.) 587, 2127–2139 (2009)
Sanyal, S. & Menon, A. K. Flipping lipids: why an’ what’s the reason for? ACS Chem. Biol. 4, 895–909 (2009)
Suzuki, J., Denning, D. P., Imanishi, E., Horvitz, H. R. & Nagata, S. Xk-related protein 8 and CED-8 promote phosphatidylserine exposure in apoptotic cells. Science 341, 403–406 (2013)
Sanyal, S. & Menon, A. K. Stereoselective transbilayer translocation of mannosyl phosphoryl dolichol by an endoplasmic reticulum flippase. Proc. Natl Acad. Sci. USA 107, 11289–11294 (2010)
Mohammadi, T. et al. Identification of FtsW as a transporter of lipid-linked cell wall precursors across the membrane. EMBO J. 30, 1425–1432 (2011)
Zwaal, R. F., Comfurius, P. & Bevers, E. M. Scott syndrome, a bleeding disorder caused by defective scrambling of membrane phospholipids. Biochim. Biophys. Acta 1636, 119–128 (2004)
Dekkers, D. W., Comfurius, P., Bevers, E. M. & Zwaal, R. F. Comparison between Ca2+-induced scrambling of various fluorescently labelled lipid analogues in red blood cells. Biochem. J. 362, 741–747 (2002)
Sham, L. T. et al. Bacterial cell wall. MurJ is the flippase of lipid-linked precursors for peptidoglycan biogenesis. Science 345, 220–222 (2014)
Suzuki, T., Suzuki, J. & Nagata, S. Functional swapping between transmembrane proteins TMEM16A and TMEM16F. J. Biol. Chem. 289, 7438–7447 (2014)
Miller, C. Open-state substructure of single chloride channels from Torpedo electroplax. Phil. Trans. R. Soc. Lond. B 299, 401–411 (1982)
Kmit, A. et al. Calcium-activated and apoptotic phospholipid scrambling induced by Ano6 can occur independently of Ano6 ion currents. Cell Death Dis. 4, e611 (2013)
Tian, Y., Schreiber, R. & Kunzelmann, K. Anoctamins are a family of Ca2+-activated Cl− channels. J. Cell Sci. 125, 4991–4998 (2012)
Baldridge, R. D. & Graham, T. R. Identification of residues defining phospholipid flippase substrate specificity of type IV P-type ATPases. Proc. Natl Acad. Sci. USA 109, E290–E298 (2012)
Vestergaard, A. L. et al. Critical roles of isoleucine-364 and adjacent residues in a hydrophobic gate control of phospholipid transport by the mammalian P4-ATPase ATP8A2. Proc. Natl Acad. Sci. USA 111, E1334–E1343 (2014)
Pifferi, S., Dibattista, M. & Menini, A. TMEM16B induces chloride currents activated by calcium in mammalian cells. Pflugers Arch. 458, 1023–1038 (2009)
Woodhull, A. M. Ionic blockage of sodium channels in nerve. J. Gen. Physiol. 61, 687–708 (1973)
Arreola, J., Melvin, J. E. & Begenisich, T. Activation of calcium-dependent chloride channels in rat parotid acinar cells. J. Gen. Physiol. 108, 35–47 (1996)
Xiao, Q. et al. Voltage- and calcium-dependent gating of TMEM16A/Ano1 chloride channels are physically coupled by the first intracellular loop. Proc. Natl Acad. Sci. USA 108, 8891–8896 (2011)
Vocke, K. et al. Calmodulin-dependent activation and inactivation of anoctamin calcium-gated chloride channels. J. Gen. Physiol. 142, 381–404 (2013)
Yu, K., Zhu, J., Qu, Z., Cui, Y. Y. & Hartzell, H. C. Activation of the Ano1 (TMEM16A) chloride channel by calcium is not mediated by calmodulin. J. Gen. Physiol. 143, 253–267 (2014)
Tian, Y. et al. Calmodulin-dependent activation of the epithelial calcium-dependent chloride channel TMEM16A. FASEB J. 25, 3. 1058–1068 (2011)
Jung, J. et al. Dynamic modulation of ANO1/TMEM16A HCO3− permeability by Ca2+/calmodulin. Proc. Natl Acad. Sci. USA 110, 360–365 (2013)
Geertsma, E. R. & Dutzler, R. A versatile and efficient high-throughput cloning tool for structural biology. Biochemistry 50, 3272–3278 (2011)
Keefe, A. D., Wilson, D. S., Seelig, B. & Szostak, J. W. One-step purification of recombinant proteins using a nanomolar-affinity streptavidin-binding peptide, the SBP-tag. Protein Expr. Purif. 23, 440–446 (2001)
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nature Biotechnol. 20, 87–90 (2002)
Schenck, S., Wojcik, S. M., Brose, N. & Takamori, S. A chloride conductance in VGLUT1 underlies maximal glutamate loading into synaptic vesicles. Nature Neurosci. 12, 156–162 (2009)
Drew, D. et al. GFP-based optimization scheme for the overexpression and purification of eukaryotic membrane proteins in Saccharomyces cerevisiae. Nature Protocols 3, 784–798 (2008)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993)
The Collaborative Computational Project, Number 4. The CCP4 suite: programs for X-ray crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D 58, 1772–1779 (2002)
Pape, T. & Schneider, T. R. HKL2MAP: a graphical user interface for phasing with SHELX programs. J. Appl. Crystallogr. 37, 843–844 (2004)
de La Fortelle, E. & Bricogne, G. in Methods in Enzymology Vol. 276 (eds Carter, C. W. & Sweet, R. M. ) 492–494 (Academic, 1997)
Cowtan, K. ‘dm’: An automated procedure for phase improvement by density modification. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography 31, 34–38 (1994)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Geertsma, E. R., Nik Mahmood, N. A., Schuurman-Wolters, G. K. & Poolman, B. Membrane reconstitution of ABC transporters and assays of translocator function. Nature Protocols 3, 256–266 (2008)
Garcia-Celma, J., Szydelko, A. & Dutzler, R. Functional characterization of a ClC transporter by solid-supported membrane electrophysiology. J. Gen. Physiol. 141, 479–491 (2013)
Accardi, A., Kolmakova-Partensky, L., Williams, C. & Miller, C. Ionic currents mediated by a prokaryotic homologue of CLC Cl− channels. J. Gen. Physiol. 123, 109–119 (2004)
Zimmermann, I. & Dutzler, R. Ligand activation of the prokaryotic pentameric ligand-gated ion channel ELIC. PLoS Biol. 9, e1001101 (2011)
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011)
Acknowledgements
This research was supported by a grant from the European Research Council (no. 339116, AnoBest) and by the Swiss National Science Foundation through the National Centre of Competence in Research TransCure. We thank the staff of the X06SA beamline for support during data collection, B. Blattman and C. Stutz-Ducommun of the Protein Crystallization Center at UZH, for their support with crystallization, B. Dreier for help with MALS experiments, A. Szydelko for providing ecClC as negative control in the scramblase assay and D. Drew for the FGY217 yeast strain. All members of the Dutzler laboratory are acknowledged for help in all stages of the project.
Author information
Authors and Affiliations
Contributions
J.D.B. screened homologues, purified and crystallized nhTMEM16 and performed scrambling experiments. N.K.L. screened and crystallized homologues, performed electrophysiological recordings and did the MALS measurement. S.S. started the project, made expression vectors and aided in cell culture. A.D. screened homologues. R.D. assisted J.D.B. and N.K.L. during structure determination. J.D.B., N.K.L., S.S. and R.D. jointly planned experiments, analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Structure-based sequence alignment.
Sequences were aligned with Clustal Omega67 and edited manually. Identical residues are highlighted in green, homologous residues in yellow and residues of the Ca2+-binding site in red. Secondary structure elements are shown below. a, Comparison of nhTMEM16 and afTMEM16. The numbering corresponds to nhTMEM16. b, Comparison of the membrane domains of selected TMEM16 proteins, m refers to murine, hs to human proteins. Long insertions in loop regions of mammalian family members (indicated by -xxx-) are not shown in the alignment. The positions of residues in α-helix 10 involved in an inter-subunit salt bridge at the dimer interface are highlighted in cyan. c, Comparison of the observed and predicted topology of TMEM16 proteins. Sequence alignment of the membrane spanning regions of mTMEM16A and nhTMEM16 with the observed (green) and predicted topology25 (red) indicated. Identical residues are highlighted in green, homologous residues in yellow, residues of the Ca2+-binding site in red and the inter-subunit salt bridge at the dimer interface in cyan. The difference between the predicted and observed transmembrane segments is due to the failure of sequence-based approaches to identify the correct boundaries of several helices and to detect α-helix 6 at all and helices 7 and 8 as separate entities.
Extended Data Figure 2 Multi-angle light scattering and lipid scrambling.
a, Gel filtration and light scattering results for nhTMEM16 in the detergent DDM. The continuous black trace corresponds to the absorption at 280 nm. Molecular weights of the protein and the protein-detergent complex are shown in red and green, respectively. b, Inaccessibility of NBD groups trapped within liposomes. Dithionite is incapable of reducing the soluble NBD-dextran trapped in the interior of proteoliposomes containing nhTMEM16. Traces of proteoliposomes containing nhTMEM16 and empty liposomes are shown in red and black respectively. Traces from proteoliposomes of nhTMEM16 containing NBD-PE at equivalent dithionite levels are shown for comparison (blue). Asterisk marks addition of 2.5 mM dithionite. c, SDS–PAGE gel of the Ca2+-binding site triple-mutant E452Q/E535Q/D539N (M) and nhTMEM16 (WT) used for reconstitution illustrating the purity of the sample. The molecular weight marker (MW) is shown on the left with selected bands labelled. d, Analysis of phospholipid scrambling. Time dependent fluorescence decrease of NBD-PE upon reduction by 30 mM dithionite (t = 0). The traces are as in Figs 1b–d and 5a. A fit to a single exponential decay is shown as dotted lines for protein-free/Ca2+ and nhTMEM16/Ca2+ with time constants of 15 s and 22 s, respectively. A fit to a sum of two exponential functions is shown for nhTMEM16/no Ca2+ and E452Q/E535Q/D539N/Ca2+ with time constants of 25 and 21 s for the fast component and 175 and 803 s for the slow component, respectively.
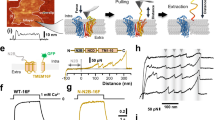
Extended Data Figure 3 Search for ion channel activity in nhTMEM16.
a, Freeze-fracture electron microscopy image of a proteoliposome containing nhTMEM16 formed from a 3:1 mixture of E. coli polar lipids/egg PC. Reconstituted proteins are labelled with red asterisks. b, Planar lipid bilayer experiments. Currents recorded after fusion of proteoliposomes containing nhTMEM16 expressed and purified from S. cerevisiae (Sc) in the absence of Ca2+ (top, left), with 300 μM Ca2+ added on both sides of the bilayer (top right) and of proteoliposomes containing nhTMEM16 expressed and purified from HEK tsA201 cells in the presence of 300 μM Ca2+ added on both sides of the bilayer (bottom left). Currents recorded after fusion of liposomes of the same lipid composition not containing any protein are shown for comparison (bottom right). Displayed traces were recorded at a holding potential of 100 mV in symmetric solutions containing 150 mM NaCl and 10 mM HEPES pH 7.4. Selected current levels (in pA) are indicated on the left. c, Fluorescence confocal microscopy images of HEK tsA201 cells expressing a mTMEM16A–YFP fusion construct (left) or a nhTMEM16–YFP fusion construct (right). d, Recordings from excised inside-out patches. Representative current response in a membrane patch excised from cells expressing a mTMEM16A–YFP fusion construct upon rapid exchange into solutions containing the indicated amount of Ca2+ (left) and equivalent recordings from patches that were excised from cells expressing a nhTMEM16–YFP fusion protein (right). The voltage was clamped at 80 mV. The fluorescence of transfected cells expressing mTMEM16A–YFP used for recording is shown below. No activity of nhTMEM16–YFP was observed in any of more than 30 patches. e, Patch-clamp recording in the whole-cell configuration. Representative currents from a HEK293T cell expressing a mTMEM16A–YFP construct recorded from a solution containing either 0.1 µM (left) or 20 µM (right) free Ca2+ in the patch pipette. f, Representative currents from a cell expressing a nhTMEM16–YFP fusion protein recorded from a solution containing 20 µM free Ca2+ in the patch pipette (left). Current response from mock-transfected cells recorded under the same conditions is shown for comparison (right). Insets show part of the traces with magnified current scale.
Extended Data Figure 4 Crystallography.
a, Table describing data collection and refinement statistics of five data sets presented in this study. nhTMEM16/CF1 and nhTMEM16/CF2 are data sets used for the building and refinement of the crystal structures of CF1 and CF2 respectively that have been deposited in the PDB. nhTMEM16/SeMet, a data set of a selenomethionine derivative collected at the Se anomalous absorption edge, was used for obtaining initial phases of CF2. nhTMEM16/anom is a data set used for the identification of the Ca2+-binding site by anomalous scattering and nhTMEM16/no Ca2+ is from a protein purified in the presence of EDTA and crystallized without addition of Ca2+. b, Stereo view of the Ca2+-binding region in CF1. The model of the protein displayed as sticks is shown with experimental electron density superimposed. The map was calculated at 3.3 Å with Se-Met SAD phases that were improved by solvent flattening, cyclic twofold NCS and cross-crystal averaging (blue mesh, contoured at 1σ). Ca2+ ions are shown as blue spheres.
Extended Data Figure 5 Electron density.
a, Stereo view of the Ca2+-binding region in CF1. The model of the protein displayed as sticks is shown with 2Fo – Fc electron density superimposed (cyan mesh, contoured at 1σ after sharpening with b = 50). The density at 3.3 Å was calculated with phases from the refined model. Ca2+ ions are shown as blue spheres. b, 2Fo – Fc electron density of the Ca2+-binding region in CF2 (calculated at 3.4 Å and contoured at 1σ after sharpening with b = 50, orange) superimposed on the refined model. c, Stereo view of the Ca2+-binding region of a structure obtained from protein purified in the presence of EDTA and crystallized in CF2 without addition of Ca2+. 2Fo – Fc electron density (cyan mesh, calculated at 4.2 Å and contoured at 1σ after sharpening with a b = 50) and Fo – Fc density (contoured at 3σ, green) is superimposed on the refined model. No ions were included in the refinement. d, Close-up of the Ca2+-binding site. Anomalous difference density (left, calculated at 6 Å and contoured at 4σ, magenta) and Fo – Fc density (right, contoured at 3σ, green) indicates the presence of bound Ca2+ ions.
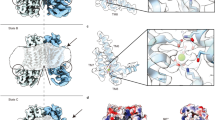
Extended Data Figure 6 nhTMEM16 dimer.
Stereo views of a ribbon representation of the dimeric protein. Bound Ca2+ ions are shown as blue spheres. a, View from within the membrane; b, view from the extracellular side; c, view from the cytoplasm.
Extended Data Figure 7 Model of lipid interactions and dimer cavity.
a, Model of nhTMEM16 embedded in a lipid membrane (left). The protein was positioned within the model of a PC bilayer (obtained from http://www.lobos.nih.gov/mbs/coords.shtml). A ribbon representation of the protein and the molecular surface are shown. Lipids are displayed as CPK models. Same view of the protein with regions on the surface presumably in contact with the membrane coloured in orange (right). b, Putative location of α-helices 0a and 0b relative to the lipid bilayer. c, Inter-subunit interactions between residues of α-helix 10. The protein is shown as sticks with 2Fo – Fc density (CF1, calculated at 3.3 Å and contoured at 1σ after sharpening with b = 50, cyan mesh) superimposed (left). A sequence alignment of the corresponding region underlines the conservation of interacting residues. Amino acids of the salt bridge in nhTMEM16 are highlighted in cyan, the numbering corresponds to nhTMEM16. d, View on the dimer cavity from the dimer interface. The molecular surface is coloured according to the properties of contacting residues (red, acidic; blue, basic; green, polar). A modelled lipid indicates the boundary of the inner leaflet of the bilayer. e, Stereo view of the cleft between α-helices 3 and 10. The protein is shown as stick model. The molecular surface is coloured according to the properties of contacting residues (yellow, hydrophobic; orange, aromatic). Lipids indicate the membrane boundary. f, Residual density in the dimer cavity. The molecular surface is coloured in white. 2Fo – Fc density (CF2, contoured at 1σ after sharpening with b = 50, orange) and Fo – Fc density (contoured at 3σ, green) are shown. The view is as in d.
Extended Data Figure 8 Subunit cavity and Ca2+-binding site.
a, Stereo view of the subunit cavity viewed from within the membrane. Protein residues and the molecular surface are shown. b, Residual density in the subunit cavity. The molecular surface of the protein is shown. 2Fo – Fc density (CF2, contoured at 1σ after sharpening with b = 50, orange) and Fo – Fc density (contoured at 3σ, green) are displayed. c, Model of the subunit cavity in different TMEM16 proteins. The molecular surface is coloured according to the properties of contacting residues (red, acidic; blue, basic; green, polar). Putative surface-exposed residues were obtained from a sequence alignment with nhTMEM16. d, Location of the Ca2+-binding site in relation to the lipid bilayer. Modelled lipids of the inner leaflet of the bilayer are shown as sticks.
Extended Data Figure 9 Electrophysiology.
Current response in HEK293T cells overexpressing mTMEM16A–YFP and point mutants of the Ca2+-binding site. All recordings were measured from single excised patches in the inside-out configuration after changing to intracellular solutions containing the indicated Ca2+ concentrations. a, WT, with voltage protocol shown as inset. b, Mutant N650A, with the voltage protocol shown as inset. c–g, Recordings of mutants E654A, c, E702Q, d, E705Q, e, E634Q, f, and mutant D738N, g.
Extended Data Figure 10 Ca2+ activation of mTMEM16A.
Representative current traces of mTMEM16A and mutants of the Ca2+-binding site. Currents were measured from excised inside-out patches of HEK293T cells expressing the respective protein at 80 mV. The Ca2+ concentration is indicated, selected traces are shown in colour. a, WT, b, N650A, c, E734Q, d, E702Q, e, D738N f, E705Q and g, E654A. h, Fluorescence microscopy image of mTMEM16A mutants expressed in HEK293T cells. Expression of fluorescently labelled protein is shown for mutants E654Q (left) and E654A (right). i, Analysis of the EC50 of Ca2+ activation for different binding site mutants. The data show averages of fits to 3–4 independent recordings. Errors are s.d.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion and additional references. (PDF 249 kb)
Rights and permissions
About this article
Cite this article
Brunner, J., Lim, N., Schenck, S. et al. X-ray structure of a calcium-activated TMEM16 lipid scramblase. Nature 516, 207–212 (2014). https://doi.org/10.1038/nature13984
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13984
This article is cited by
-
Structural heterogeneity of the ion and lipid channel TMEM16F
Nature Communications (2024)
-
Two broadly conserved families of polyprenyl-phosphate transporters
Nature (2023)
-
Extracellular calcium functions as a molecular glue for transmembrane helices to activate the scramblase Xkr4
Nature Communications (2023)
-
Substrate binding-induced conformational transitions in the omega-3 fatty acid transporter MFSD2A
Nature Communications (2023)
-
Molecular logic of salt taste reception in special reference to transmembrane channel-like 4 (TMC4)
The Journal of Physiological Sciences (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.