Abstract
Elongation of the head-to-tail body axis by convergent extension is a conserved developmental process throughout metazoans. In Drosophila, patterns of transcription factor expression provide spatial cues that induce systematically oriented cell movements and promote tissue elongation. However, the mechanisms by which patterned transcriptional inputs control cell polarity and behaviour have long been elusive. We demonstrate that three Toll family receptors, Toll-2, Toll-6 and Toll-8, are expressed in overlapping transverse stripes along the anterior–posterior axis and act in combination to direct planar polarity and polarized cell rearrangements during convergent extension. Simultaneous disruption of all three receptors strongly reduces actomyosin-driven junctional remodelling and axis elongation, and an ectopic stripe of Toll receptor expression is sufficient to induce planar polarized actomyosin contractility. These results demonstrate that tissue-level patterns of Toll receptor expression provide spatial signals that link positional information from the anterior–posterior patterning system to the essential cell behaviours that drive convergent extension.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Keller, R. et al. Mechanisms of convergence and extension by cell intercalation. Phil. Trans. R. Soc. Lond. B 355, 897–922 (2000)
Zallen, J. A. Planar polarity and tissue morphogenesis. Cell 129, 1051–1063 (2007)
Wallingford, J. B. Planar cell polarity and the developmental control of cell behavior in vertebrate embryos. Annu. Rev. Cell Dev. Biol. 28, 627–653 (2012)
Solnica-Krezel, L. & Sepich, D. S. Gastrulation: making and shaping germ layers. Annu. Rev. Cell Dev. Biol. 28, 687–717 (2012)
Walck-Shannon, E. & Hardin, J. Cell intercalation from top to bottom. Nature Rev. Mol. Cell Biol. 15, 34–48 (2014)
Zallen, J. A. & Wieschaus, E. Patterned gene expression directs bipolar planar polarity in Drosophila. Dev. Cell 6, 343–355 (2004)
Bertet, C., Sulak, L. & Lecuit, T. Myosin-dependent junction remodelling controls planar cell intercalation and axis elongation. Nature 429, 667–671 (2004)
Blankenship, J. T., Backovic, S. T., Sanny, J. S. P., Weitz, O. & Zallen, J. A. Multicellular rosette formation links planar cell polarity to tissue morphogenesis. Dev. Cell 11, 459–470 (2006)
Rauzi, M., Verant, P., Lecuit, T. & Lenne, P.-F. Nature and anisotropy of cortical forces orienting Drosophila tissue morphogenesis. Nature Cell Biol. 10, 1401–1410 (2008)
Fernández-González, R., Simões de M, S., Röper, J.-C., Eaton, S. & Zallen, J. A. Myosin II dynamics are regulated by tension in intercalating cells. Dev. Cell 17, 736–743 (2009)
Nishimura, T. & Takeichi, M. Shroom3-mediated recruitment of Rho kinases to the apical cell junctions regulates epithelial and neuroepithelial planar remodeling. Development 135, 1493–1502 (2008)
Nishimura, T., Honda, H. & Takeichi, M. Planar cell polarity links axes of spatial dynamics in neural-tube closure. Cell 149, 1084–1097 (2012)
Lienkamp, S. S. et al. Vertebrate kidney tubules elongate using a planar cell polarity-dependent, rosette-based mechanism of convergent extension. Nature Genet. 44, 1382–1387 (2012)
Mahaffey, J. P., Grego-Bessa, J., Liem, K. F. & Anderson, K. V. Cofilin and Vangl2 cooperate in the initiation of planar cell polarity in the mouse embryo. Development 140, 1262–1271 (2013)
Shindo, A. & Wallingford, J. B. PCP and septins compartmentalize cortical actomyosin to direct collective cell movement. Science 343, 649–652 (2014)
Williams, M., Yen, W., Lu, X. & Sutherland, A. Distinct apical and basolateral mechanisms drive planar cell polarity-dependent convergent extension of the mouse neural plate. Dev. Cell 29, 34–46 (2014)
Irvine, K. D. & Wieschaus, E. Cell intercalation during Drosophila germband extension and its regulation by pair-rule segmentation genes. Development 120, 827–841 (1994)
Ninomiya, H., Elinson, R. P. & Winklbauer, R. Antero-posterior tissue polarity links mesoderm convergent extension to axial patterning. Nature 430, 364–367 (2004)
St Johnston, D. & Nüsslein-Volhard, C. The origin of pattern and polarity in the Drosophila embryo. Cell 68, 201–219 (1992)
Butler, L. C. et al. Cell shape changes indicate a role for extrinsic tensile forces in Drosophila germ-band extension. Nature Cell Biol. 11, 859–864 (2009)
Simões de M, S. et al. Rho-kinase directs Bazooka/Par-3 planar polarity during Drosophila axis elongation. Dev. Cell 19, 377–388 (2010)
Wieschaus, E., Sweeton, D. & Costa, M. in Gastrulation 213–223 (Springer, 1992)
Brennan, C. A. & Anderson, K. V. Drosophila: the genetics of innate immune recognition and response. Annu. Rev. Immunol. 22, 457–483 (2004)
Janeway, C. A. & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002)
Leulier, F. & Lemaitre, B. Toll-like receptors—taking an evolutionary approach. Nature Rev. Genet. 9, 165–178 (2008)
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nature Immunol. 11, 373–384 (2010)
Tauszig, S., Jouanguy, E., Hoffmann, J. A. & Imler, J. L. Toll-related receptors and the control of antimicrobial peptide expression in Drosophila. Proc. Natl Acad. Sci. USA 97, 10520–10525 (2000)
Morisato, D. & Anderson, K. V. Signaling pathways that establish the dorsal-ventral pattern of the Drosophila embryo. Annu. Rev. Genet. 29, 371–399 (1995)
Chiang, C. & Beachy, P. A. Expression of a novel Toll-like gene spans the parasegment boundary and contributes to hedgehog function in the adult eye of Drosophila. Mech. Dev. 47, 225–239 (1994)
Kambris, Z., Hoffmann, J. A., Imler, J.-L. & Capovilla, M. Tissue and stage-specific expression of the Tolls in Drosophila embryos. Gene Expr. Patterns 2, 311–317 (2002)
Eldon, E. et al. The Drosophila 18 wheeler is required for morphogenesis and has striking similarities to Toll. Development 120, 885–899 (1994)
Keith, F. J. & Gay, N. J. The Drosophila membrane receptor Toll can function to promote cellular adhesion. EMBO J. 9, 4299–4306 (1990)
Kim, S., Chung, S., Yoon, J., Choi, K.-W. & Yim, J. Ectopic expression of Tollo/Toll-8 antagonizes Dpp signaling and induces cell sorting in the Drosophila wing. Genesis 44, 541–549 (2006)
Kleve, C. D., Siler, D. A., Syed, S. K. & Eldon, E. D. Expression of 18-wheeler in the follicle cell epithelium affects cell migration and egg morphology in Drosophila. Dev. Dyn. 235, 1953–1961 (2006)
Kolesnikov, T. & Beckendorf, S. K. 18 wheeler regulates apical constriction of salivary gland cells via the Rho-GTPase-signaling pathway. Dev. Biol. 307, 53–61 (2007)
Paré, A. et al. Visualization of individual Scr mRNAs during Drosophila embryogenesis yields evidence for transcriptional bursting. Curr. Biol. 19, 2037–2042 (2009)
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011)
Tamada, M., Farrell, D. L. & Zallen, J. A. Abl regulates planar polarized junctional dynamics through β-catenin tyrosine phosphorylation. Dev. Cell 22, 309–319 (2012)
Kasza, K. E., Farrell, D. L. & Zallen, J. A. Spatiotemporal control of epithelial remodeling by regulated myosin phosphorylation. Proc. Natl Acad. Sci. USA 111, 11732–11737 (2014)
McIlroy, G. et al. Toll-6 and Toll-7 function as neurotrophin receptors in the Drosophila melanogaster CNS. Nature Neurosci. 16, 1248–1256 (2013)
Ballard, S. L., Miller, D. L. & Ganetzky, B. Retrograde neurotrophin signaling through Tollo regulates synaptic growth in Drosophila. J. Cell Biol. 204, 1157–1172 (2014)
Özkan, E. et al. An extracellular interactome of immunoglobulin and LRR proteins reveals receptor–ligand networks. Cell 154, 228–239 (2013)
de Wit, J., Hong, W., Luo, L. & Ghosh, A. Role of leucine-rich repeat proteins in the development and function of neural circuits. Annu. Rev. Cell Dev. Biol. 27, 697–729 (2011)
Rakoff-Nahoum, S. & Medzhitov, R. Toll-like receptors and cancer. Nature Rev. Cancer 9, 57–63 (2009)
Grote, K., Schütt, H. & Schieffer, B. Toll-like receptors in angiogenesis. Scientific World J. 11, 981–991 (2011)
Huebener, P. & Schwabe, R. F. Regulation of wound healing and organ fibrosis by toll-like receptors. Biochim. Biophy. Acta 1832, 1005–1017 (2013)
West, M. A. et al. Enhanced dendritic cell antigen capture via toll-like receptor-induced actin remodeling. Science 305, 1153–1157 (2004)
Ma, Y. et al. Toll-like receptor 8 functions as a negative regulator of neurite outgrowth and inducer of neuronal apoptosis. J. Cell Biol. 175, 209–215 (2006)
Cameron, J. S. et al. Toll-like receptor 3 is a potent negative regulator of axonal growth in mammals. J. Neurosci. 27, 13033–13041 (2007)
Rose, D. et al. Toll, a muscle cell surface molecule, locally inhibits synaptic initiation of the RP3 motoneuron growth cone in Drosophila. Development 124, 1561–1571 (1997)
Gerttula, S., Jin, Y. & Anderson, K. V. Zygotic expression and activity of the Drosophila Toll gene, a gene required maternally for embryonic dorsal-ventral pattern formation. Genetics 119, 123–133 (1988)
Yagi, Y., Nishida, Y. & Ip, Y. T. Functional analysis of Toll-related genes in Drosophila. Dev. Growth Differ. 52, 771–783 (2010)
Nüsslein-Volhard, C. & Wieschaus, E. Mutations affecting segment number and polarity in Drosophila. Nature 287, 795–801 (1980)
Duffy, J. B. & Gergen, J. P. The Drosophila segmentation gene runt acts as a position-specific numerator element necessary for the uniform expression of the sex-determining gene Sex-lethal. Genes Dev. 5, 2176–2187 (1991)
Ludwig Manu, M. Z., R., White, K. P. & Kreitman, M. Consequences of eukaryotic enhancer architecture for gene expression dynamics, development, and fitness. PLoS Genet. 7, e1002364 (2011)
Martin, A. C., Gelbart, M., Fernández-González, R., Kaschube, M. & Wieschaus, E. F. Integration of contractile forces during tissue invagination. J. Cell Biol. 188, 735–749 (2010)
Royou, A., Field, C., Sisson, J. C., Sullivan, W. & Karess, R. Reassessing the role and dynamics of nonmuscle myosin II during furrow formation in early Drosophila embryos. Mol. Biol. Cell 15, 838–850 (2004)
Schmittgen, T. D. & Livak, K. J. Analyzing real-time PCR data by the comparative CT method. Nature Protocols 3, 1101–1108 (2008)
Nagaso, H., Murata, T., Day, N. & Yokoyama, K. K. Simultaneous detection of RNA and protein by in situ hybridization and immunological staining. J. Histochem. Cytochem. 49, 1177–1182 (2001)
Kosman, D. et al. Multiplex detection of RNA expression in Drosophila embryos. Science 305, 846 (2004)
Kosman, D., Small, S. & Reinitz, J. Rapid preparation of a panel of polyclonal antibodies to Drosophila segmentation proteins. Dev. Genes Evol. 208, 290–294 (1998)
Hutson, M. S. et al. Forces for morphogenesis investigated with laser microsurgery and quantitative modeling. Science 300, 145–149 (2003)
Acknowledgements
We thank K. Anderson, K. Kasza, W. Razzell, G. Sabio, M. Shirasu-Hiza, A. Spencer, M. Tamada and R. Zallen for comments on the manuscript, B. Glick for the fast-folding YFP, M. Buszczak for pUASp-w-attB, and the BAC-Recombineering Core Facility at the University of Chicago for Toll-8–YFP. This work was funded by NIH/NIGMS grants GM079340 and GM102803 to J.A.Z. J.A.Z. is an Early Career Scientist of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
A.C.P., A.V. and J.A.Z. designed the study. A.C.P., A.V., C.T.F. and Z.M. performed the experiments, D.L.F. and A.M. performed the computational analysis, and A.C.P. and J.A.Z. wrote the manuscript. All authors participated in analysis of the data and in producing the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Targeting of Eve, Runt and Toll receptors by dsRNA injection.
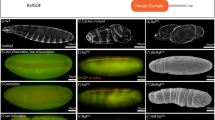
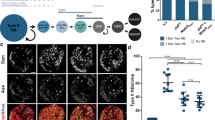
a–c, Control and dsRNA-injected embryos stained for Runt (red, middle) and Wingless (Wg) (green, bottom) proteins. a, In uninjected wild-type embryos, Runt is expressed in seven broad stripes and Wg is expressed in 14 narrow stripes. b, In embryos injected with eve dsRNA alone, Runt is more uniformly expressed, and the Wg expression pattern collapses into fewer, broader stripes, similar to eve mutants (data not shown). c, In embryos co-injected with eve and runt dsRNAs, Runt protein is undetectable, indicating that the runt dsRNA effectively inhibits Runt expression, and the Wg expression pattern collapses into fewer, broader stripes, similar to eve and runt mutants (data not shown). Anterior left, ventral down. Scale bars, 100 μm. d–f, Quantitative RT–PCR analysis of Toll-2 (2), Toll-6 (6), Toll-7 (7) and Toll-8 (8) expression in late stage 6 embryos before axis elongation. CT values were normalized to the internal control gene RpL32. d, Relative transcript levels in WT embryos were calculated using the 2−ΔCT method. Toll-2, Toll-6 and Toll-8 were expressed at comparable levels, whereas Toll-7 was expressed at much lower levels. e, Toll-8 expression in Toll-859/145 embryos was reduced 76-fold compared with WT embryos. f, Gene expression was specifically reduced in embryos injected with single dsRNAs targeting Toll-2, Toll-6 or Toll-8 compared with embryos injected with a control Toll-3 dsRNA, as determined using the 2−ΔΔCT method.
Extended Data Figure 2 Expression patterns of Toll-2, Toll-6 and Toll-8 relative to Runt.
a–f, Toll-2, Toll-6 and Toll-8 transcripts (green top, white bottom) and Runt protein (magenta) in wild-type (WT) embryos during early (stage 7, a–c) and mid-elongation (stage 8, d–f). The embryos are the same as in Fig. 1a–f. Coloured bars indicate the position of the Toll-2, Toll-6 and Toll-8 stripes (green) relative to Runt (magenta). Anterior left, ventral down. Scale bars, 100 μm.
Extended Data Figure 3 Time-lapse imaging of embryos defective for combinations of Toll-2, Toll-6 and Toll-8.
a–e, Axis elongation (tissue AP length relative to t = 0) (first row), total cell rearrangements (average number of neighbours lost per cell) (second row), T1 processes resulting from the contraction of single edges7 (third row), and rosette rearrangements resulting from the contraction of multiple connected edges8 (fourth row) over time in wild-type embryos (a) and embryos defective for one (b), two (c), or three (d, e) Toll receptors. Images were acquired every 15 s. f–i, Axis elongation (f), average number of cell rearrangements (g), T1 processes (h), and rosettes (i) per cell at t = 30 min. j, Edge contraction rate for AP edges (oriented 75–90° relative to the AP axis) at mid-stage 7 (averaged from t = 5–8 min after the onset of elongation). k, The orientation of shrinking edges relative to the AP axis (0°) was similar for all conditions. Single average values were obtained for each embryo, and plots show the mean ± s.e.m. across embryos. *P = 0.01–0.03, **P < 0.005 (unpaired t-test). n = 3–8 embryos per genotype, 164–365 cells per embryo (see Supplementary Table 2 for full list of n values). l, Cross sections of the ventrolateral epithelium in wild-type and Toll-2Δ76, Toll-61B, Toll-714F, Toll-859 mutant (Toll-2,6,7,8) embryos, showing that apical–basal polarity is unaffected in quadruple mutants. Myosin II (green) and Par-3 (red, white) are enriched at apical adherens junctions. Apical up, basal down. Scale bars, 10 μm. WT (Spider–GFP in a, e–k; Resille–GFP in a, f–k; and Resille–GFP + Toll-3 dsRNA in a–d and f–k); Toll-2 (Resille–GFP + Toll-2 dsRNA); Toll-6 (Resille–GFP + Toll-6 dsRNA); Toll-8 (Resille–GFP; Toll-859/145); Toll-2,6 (Resille–GFP + Toll-2/Toll-6 dsRNAs); Toll-2,8 (Resille–GFP; Toll-859/145+ Toll-2 dsRNA); Toll-6,8 (Toll-2Δ76/CyO; Toll-859, Toll-65A, Spider–GFP); Toll-2,6,8 (Toll-2Δ76; Toll-859, Toll-65A, Spider–GFP), Toll-2,6,8 i1 (Resille–GFP; Toll-859/145+ Toll-2/Toll-6 dsRNAs set 1); Toll-2,6,8 i2 (Resille–GFP; Toll-859/145+ Toll-2/Toll-6 dsRNAs set 2); runt (runtLB5; Spider–GFP/+); and eve (eveR13; Spider–GFP/+).
Extended Data Figure 4 Generation of double, triple and quadruple Toll receptor mutants.
a, The crossing strategy used to generate Toll-2,6,8 triple mutants and Toll-2,6,7,8 quadruple mutants. Toll-7 and Toll-2 are 285 kb apart on the right arm of chromosome II and Toll-8 and Toll-6 are 94 kb apart on the left arm of chromosome III. b, Three unique Toll-6 null alleles (Toll-61B, Toll-64B and Toll-65A) were generated on the Toll-859 chromosome using TALEN-mediated mutagenesis to create Toll-8, Toll-6 double-mutant chromosomes. c, Six unique Toll-7 null alleles (Toll-71C, Toll-74D, Toll-75A, Toll-75F, Toll-714F and Toll-716A) were generated on the Toll-2Δ76 chromosome using TALEN-mediated mutagenesis to create Toll-7, Toll-2 double-mutant chromosomes. Each Toll-6 and Toll-7 allele is a frame-shift mutation leading to premature translational termination. TALENs were designed to induce double-stranded breaks immediately downstream of the ATG translational start sites. Orange letters indicate the TALEN binding sites, and the spacer regions are shown in bold. The AvaII and AvaI restriction sites used for screening are indicated with dotted boxes. Shown below are the predicted amino acid sequences of the mutant proteins compared with the wild-type sequence. Residues that are identical in the mutant and wild-type proteins are shown in green.
Extended Data Figure 5 Distributions of cell polarity measurements in Toll receptor mutants.
a–l, Planar polarity distributions for myosin II (left panels) and Par-3 (right panels) in Toll-2 single mutants (a, b), Toll-6,8 double mutants (c, d), Toll-2,6,8 triple mutants (e, f), Toll-2,6,7,8 quadruple mutants (g, h), runt mutants (i, j) and eve mutants (k, l). Vertical lines indicate the means of the distributions. Error bars indicate s.e.m. between embryos. Mean planar polarity was shifted towards 1 (absolute ratio; 0 on the log2 scale) in Toll-2 single mutants (P = 0.005 for myosin and P < 0.00005 for Par-3), Toll-2,6,8 triple mutants (P < 0.00002 for myosin and Par-3), Toll-2,6,7,8 quadruple mutants (P < 0.00001 for myosin and Par-3), runt mutants (P = 0.002 for myosin and P < 0.00001 for Par-3) and eve mutants (P < 0.00001 for myosin and Par-3), indicating reduced planar polarity (unpaired t-test with the means of the distributions used as the test statistic). Planar polarity in Toll-2,6,7,8 quadruple mutants was not significantly enhanced relative to triple mutants. Single values were obtained for each embryo, and plots show the mean ± s.e.m. across embryos. n = 2,166–4,909 cells in 7–20 embryos per genotype (Supplementary Table 2).
Extended Data Figure 6 Toll receptor expression affects planar polarity in a regional manner.
a, Single-cell analysis of Par-3 planar polarity in wild-type (WT) (left) and Toll-2 mutant (right) embryos. Toll-8-expressing cells were identified by fluorescence in situ hybridization. Cyan lines, boundaries between stripes; Toll-8+, Toll-8-expressing cells. AP enriched (red), DV enriched (blue). Cells without at least one AP and one DV edge were not scored (grey). b, c, Myosin II (cyan, white) and Toll-2 mRNA (red) in stage 7 WT (b) and eve mutant (c) embryos. Arrows, residual myosin cables in eve embryos. Anterior left, ventral down. Scale bars, 20 μm.
Supplementary information
Supplementary Table 1
This file contains toll receptor expression levels and significantly misregulated genes in eve/runt dsRNA-injected embryos. The transcriptomes of embryos injected with both eve and runt dsRNAs were compared with water-injected controls using RNA sequencing (Gene Expression Omnibus accession number GSE61689). Embryos were hand-selected at the late blastoderm stage, 15-30 min prior to the start of axis elongation (late stage 5/early stage 6). (Top) Toll-8 (Tollo) and Toll-2 (18w) were strongly expressed and Toll-6 and Toll-7 were weakly expressed at this stage. Toll (Tl) is maternally and zygotically expressed at this stage51. Expression of the other Drosophila Toll family genes was not detected. (Middle) Transcripts that were significantly upregulated in eve/runt double RNAi embryos (p<0.01). (Bottom) Transcripts that were significantly downregulated in eve/runt double RNAi embryos (p<0.01). (XLSX 16 kb)
Supplementary Table 2
This file contains a summary of genotypes used and the number of cells and embryos in each experiment. (XLSX 15 kb)
Cell intercalation in wild-type and Toll-2,6,8 dsRNA-injected embryos
Time-lapse videos of a wild-type control-injected embryo (WT) (top) and a Toll-8 mutant embryo injected with Toll-2/Toll-6 dsRNAs (Toll-2,6,8) (bottom). Cell membranes were labeled with Resille:GFP. Cells are labeled according to number of neighbors lost (purple=0, blue=1, green=2, yellow=3, red=4). Anterior left, ventral down; t = 0 is the onset of elongation. Images were acquired at 40× magnification with 15 s intervals between time points. (MOV 9109 kb)
Cell intercalation in wild-type and Toll-2,6,8 triple-mutant embryos
Time-lapse videos of a wild-type embryo (WT) (top) and a Toll-2,6,8 triple mutant embryo (bottom). Cell membranes were labeled with Spider:GFP. Cells are labeled according to number of neighbors lost (purple=0, blue=1, green=2, yellow=3, red=4). Anterior left, ventral down; t = 0 is the onset of elongation. Images were acquired at 40× magnification with 15 s intervals between time points. (MOV 14150 kb)
Myosin II localization in a late stage wild-type embryo
Time-lapse video of myosin:GFP in a wild-type stage 15 embryo. Yellow arrows indicate the anterior boundaries of two engrailed stripes, anterior to the denticle field (white dots). Anterior left, ventral down; t = 0 is the beginning of stage 15 (time in h:min:sec). Images were acquired at 40× magnification with 30 s intervals between time points. (MOV 7160 kb)
Myosin II is recruited to the anterior boundary of ectopic Toll-2 expression
Time-lapse video of myosin:GFP in a stage 15 embryo expressing Toll-2:HA in ectopic stripes under the control of engrailed-Gal4. Yellow arrows indicate the anterior boundaries of two engrailed stripes, which form an ectopic myosin cable. Anterior left, ventral down; t = 0 is the beginning of stage 15 (time in h:min:sec). Images were acquired at 40× magnification with 30 s intervals between time points. (MOV 6513 kb)
Myosin II is recruited to the anterior boundary of ectopic Toll-8 expression
Time-lapse video of myosin:GFP in a stage 15 embryo expressing Toll-8:HA in ectopic stripes under the control of engrailed-Gal4. Yellow arrows indicate the anterior boundaries of two engrailed stripes, which form an ectopic myosin cable. Anterior left, ventral down; t = 0 is the beginning of stage 15 (time in h:min:sec). Images were acquired at 40× magnification with 30 s intervals between time points. (MOV 7256 kb)
Rights and permissions
About this article
Cite this article
Paré, A., Vichas, A., Fincher, C. et al. A positional Toll receptor code directs convergent extension in Drosophila. Nature 515, 523–527 (2014). https://doi.org/10.1038/nature13953
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13953
This article is cited by
-
Homeotic compartment curvature and tension control spatiotemporal folding dynamics
Nature Communications (2023)
-
Serotonin signaling regulates actomyosin contractility during morphogenesis in evolutionarily divergent lineages
Nature Communications (2023)
-
Patterned mechanical feedback establishes a global myosin gradient
Nature Communications (2022)
-
Programmed and self-organized flow of information during morphogenesis
Nature Reviews Molecular Cell Biology (2021)
-
Differential cell adhesion implemented by Drosophila Toll corrects local distortions of the anterior-posterior compartment boundary
Nature Communications (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.