Abstract
Chromosomal rearrangements have a central role in the pathogenesis of human cancers and often result in the expression of therapeutically actionable gene fusions1. A recently discovered example is a fusion between the genes echinoderm microtubule-associated protein like 4 (EML4) and anaplastic lymphoma kinase (ALK), generated by an inversion on the short arm of chromosome 2: inv(2)(p21p23). The EML4–ALK oncogene is detected in a subset of human non-small cell lung cancers (NSCLC)2 and is clinically relevant because it confers sensitivity to ALK inhibitors3. Despite their importance, modelling such genetic events in mice has proven challenging and requires complex manipulation of the germ line. Here we describe an efficient method to induce specific chromosomal rearrangements in vivo using viral-mediated delivery of the CRISPR/Cas9 system to somatic cells of adult animals. We apply it to generate a mouse model of Eml4–Alk-driven lung cancer. The resulting tumours invariably harbour the Eml4–Alk inversion, express the Eml4–Alk fusion gene, display histopathological and molecular features typical of ALK+ human NSCLCs, and respond to treatment with ALK inhibitors. The general strategy described here substantially expands our ability to model human cancers in mice and potentially in other organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Taki, T. & Taniwaki, M. Chromosomal translocations in cancer and their relevance for therapy. Curr. Opin. Oncol. 18, 62–68 (2006)
Soda, M. et al. Identification of the transforming EML4–ALK fusion gene in non-small-cell lung cancer. Nature 448, 561–566 (2007)
Kwak, E. L. et al. Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N. Engl. J. Med. 363, 1693–1703 (2010)
Tuveson, D. A. & Jacks, T. Technologically advanced cancer modeling in mice. Curr. Opin. Genet. Dev. 12, 105–110 (2002)
Sharpless, N. E. & Depinho, R. A. The mighty mouse: genetically engineered mouse models in cancer drug development. Nature Rev. Drug Discov. 5, 741–754 (2006)
Pirazzoli, V. et al. Acquired resistance of EGFR-mutant lung adenocarcinomas to afatinib plus cetuximab is associated with activation of mTORC1. Cell Rep. 7, 999–1008 (2014)
Bergers, G. & Hanahan, D. Modes of resistance to anti-angiogenic therapy. Nature Rev. Cancer 8, 592–603 (2008)
Rottenberg, S. et al. Selective induction of chemotherapy resistance of mammary tumors in a conditional mouse model for hereditary breast cancer. Proc. Natl Acad. Sci. USA 104, 12117–12122 (2007)
Heisterkamp, N. et al. Acute leukaemia in bcr/abl transgenic mice. Nature 344, 251–253 (1990)
Zuber, J. et al. Mouse models of human AML accurately predict chemotherapy response. Genes Dev. 23, 877–889 (2009)
Lange, K. et al. Overexpression of NPM–ALK induces different types of malignant lymphomas in IL-9 transgenic mice. Oncogene 22, 517–527 (2003)
Chiarle, R. et al. NPM–ALK transgenic mice spontaneously develop T-cell lymphomas and plasma cell tumors. Blood 101, 1919–1927 (2003)
Soda, M. et al. A mouse model for EML4–ALK-positive lung cancer. Proc. Natl Acad. Sci. USA 105, 19893–19897 (2008)
Corral, J. et al. An Mll–AF9 fusion gene made by homologous recombination causes acute leukemia in chimeric mice: a method to create fusion oncogenes. Cell 85, 853–861 (1996)
Smith, A. J. et al. A site-directed chromosomal translocation induced in embryonic stem cells by Cre-loxP recombination. Nature Genet. 9, 376–385 (1995)
Collins, E. C., Pannell, R., Simpson, E. M., Forster, A. & Rabbitts, T. H. Inter-chromosomal recombination of Mll and Af9 genes mediated by cre-loxP in mouse development. EMBO Rep. 1, 127–132 (2000)
Piganeau, M. et al. Cancer translocations in human cells induced by zinc finger and TALE nucleases. Genome Res. 23, 1182–1193 (2013)
Torres, R. et al. Engineering human tumour-associated chromosomal translocations with the RNA-guided CRISPR-Cas9 system. Nature Commun. 5, 3964 (2014)
Brunet, E. et al. Chromosomal translocations induced at specified loci in human stem cells. Proc. Natl Acad. Sci. USA 106, 10620–10625 (2009)
Choi, P. S. & Meyerson, M. Targeted genomic rearrangements using CRISPR/Cas technology. Nature Commun. 5, 3728 (2014)
Choi, Y. L. et al. Identification of novel isoforms of the EML4–ALK transforming gene in non-small cell lung cancer. Cancer Res. 68, 4971–4976 (2008)
Horvath, P. & Barrangou, R. CRISPR/Cas, the immune system of bacteria and archaea. Science 327, 167–170 (2010)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Morris, S. W. et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin’s lymphoma. Science 263, 1281–1284 (1994)
DuPage, M., Dooley, A. L. & Jacks, T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nature Protocols 4, 1064–1072 (2009)
Nishino, M. et al. Histologic and cytomorphologic features of ALK-rearranged lung adenocarcinomas. Modern Pathol. 25, 1462–1472 (2012)
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001)
Chiarle, R., Voena, C., Ambrogio, C., Piva, R. & Inghirami, G. The anaplastic lymphoma kinase in the pathogenesis of cancer. Nature Rev. Cancer 8, 11–23 (2008)
Canver, M. C. et al. Characterization of genomic deletion efficiency mediated by clustered regularly interspaced palindromic repeats (CRISPR)/Cas9 nuclease system in mammalian cells. J. Biol. Chem. 289, 21312–21324 (2014)
Acknowledgements
We would like to thank M. Fazio, M. Ladanyi, G. Riely, S. Armstrong, and the members of the Ventura, Lowe and Jacks laboratories for discussion and comments. We also thank J. Hollenstein for editing the manuscript, T. Jacks for providing tumour samples from K-RasG12D mice, and the Cytogenetic Core Facility of MSKCC for tissue processing and histology. This work was supported by grants from the Geoffrey Beene Cancer Research Foundation (A.V.), NCI (Cancer Center Support Grant P30 CA008748, E.d.S.), HHMI (S.W.L.), NCI Project Grant (S.W.L.); and by fellowships from the American Italian Cancer Foundation (D.M.), the Foundation Blanceflor Boncompagni Ludovisi, née Bildt (D.M.), and the Jane Coffin Childs Foundation (E.M.). C.P.C. was supported by an NCI training grant.
Author information
Authors and Affiliations
Contributions
D.M. and A.V. conceived the project, designed and analysed the experiments, and wrote the manuscript. S.W.L. contributed to the interpretation of the results and the writing of the manuscript. D.M. generated and tested the constructs, performed the cell-based experiments, and characterized the Eml4–Alk tumours. E.M., D.M., C.B., Y.-C.H. and P.O. performed the in vivo experiments. E.d.S. supervised the crizotinib treatment experiments and analysed the results. J.A.V., D.M., C.P.C. and A.V. microdissected and analysed lung tumours to detect the Eml4–Alk inversion. C.B., D.M. and A.C. performed the immunostainings. N.R. reviewed the histopathology.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Human and murine Eml4–Alk.
a, Alignment of human EML4 exon 13 and mouse Eml4 exon 14. b, Alignment of the junction between the human EML4–ALK (variant 1) and the predicted murine Eml4–Alk proteins.
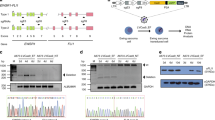
Extended Data Figure 2 Induction of the Npm1–Alk translocation in NIH/3T3 cells.
a, Schematic of the Npm1–Alk translocation. Red arrows indicate the sites recognized by the sgRNAs. b, Sequences recognized by the sgRNAs and location of primers used to detect the Npm1–Alk and Alk–Npm1 rearrangement (top panel). PCR on genomic DNA extracted from NIH/3T3 co-transfected with pX330 constructs expressing the indicated sgRNAs (middle panel). Sequences of four independent subclones obtained from the PCR products and representative chromatogram (bottom panel). c, Detection of the Npm1–Alk fusion transcript by RT–PCR on total RNAs extracted from NIH/3T3 cells co-transfected with the indicated pX330 constructs (left panel). The PCR band was extracted and sequenced to confirm the presence of the correct Npm1–Alk junction (bottom-right panel). Representative results from two independent experiments.
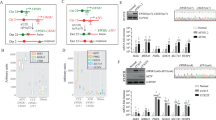
Extended Data Figure 3 Comparison of dual and single sgRNA-expressing plasmids.
a, Schematic of pX330 (A) and its derivatives (B–E) used in these experiments. NIH/3T3 were transfected with these constructs and lysed to extract total RNA and genomic DNA. b, RNAs were analysed by northern blotting with probes against the Alk (left) or Eml4 (right) sgRNAs. c, d, The DNA samples were subjected to surveyor assays (c), or amplified by PCR to detect the Eml4–Alk inversion (d).
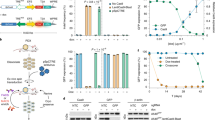
Extended Data Figure 4 Induction of the Eml4–Alk inversion in primary MEFs using an adenoviral vector expressing Flag–Cas and tandem sgRNAs.
a, Schematic of the adenoviral vectors. b, Immunoblot using an anti-Flag antibody on lysates from MEFs infected with the indicated adenoviruses. c, Small-RNA northerns using probes against sgEml4 and sgAlk on total RNAs from cells infected with Ad-Cas9 or Ad-EA. d, PCR-mediated detection of the Eml4–Alk inversion in MEFs infected with Ad-Cas9 or Ad-EA for the indicated number of days. e, Standard curve generated performing quantitative PCR analysis on genomic DNA containing a known fraction of Eml4–Alk alleles. Average of two independent experiments. f, Quantification of the fraction of MEFs harbouring the Eml4–Alk inversion at the indicated time points after infection with Ad-EA or Ad-Cas9. Values are mean of three independent infections ± s.d.
Extended Data Figure 5 Radiologic response of Ad-EA-induced tumours to crizotinib treatment.
µCT images from crizotinib- or vehicle-treated mice at day 0 and after 2 weeks of treatment.
Supplementary information
Vehicle-treatment day 0 (OP1259)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1259 at day 0 of treatment with vehicle (water). (MOV 9878 kb)
Vehicle-treatment week 2 (OP1259)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1259 after 2 weeks of treatment with vehicle (water). (MOV 9574 kb)
Crizotinib-treatment day 0 (OP1300)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1300 at day 0 of treatment with crizotinib. (MOV 11508 kb)
Crizotinib-treatment week 2 (OP1300)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1300 after 2 weeks of treatment with crizotinib. (MOV 10605 kb)
Vehicle-treatment day 0 (OP1280)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1280 at day 0 of treatment with vehicle (water). (MOV 8885 kb)
Vehicle-treatment week 2 (OP1280)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1280 after 2 weeks of treatment with vehicle (water). (MOV 9429 kb)
Crizotinib-treatment day 0 (OP1290)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1290 at day 0 of treatment with crizotinib. (MOV 5581 kb)
Crizotinib-treatment week 2 (OP1290)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1290 after 2 weeks of treatment with crizotinib. (MOV 5166 kb)
Crizotinib-treatment day 0 (OP1293)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1293 at day 0 of treatment with crizotinib. (MOV 5537 kb)
Crizotinib-treatment week 2 (OP1293)
Video generated from axial µCT scans of Ad-EA-infected mouse OP1293 after 2 weeks of treatment with crizotinib. (MOV 5128 kb)
Rights and permissions
About this article
Cite this article
Maddalo, D., Manchado, E., Concepcion, C. et al. In vivo engineering of oncogenic chromosomal rearrangements with the CRISPR/Cas9 system. Nature 516, 423–427 (2014). https://doi.org/10.1038/nature13902
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13902
This article is cited by
-
Downstream STING pathways IRF3 and NF-κB differentially regulate CCL22 in response to cytosolic dsDNA
Cancer Gene Therapy (2024)
-
Genetically-engineered mouse models of small cell lung cancer: the next generation
Oncogene (2024)
-
Efficient and safe therapeutic use of paired Cas9-nickases for primary hyperoxaluria type 1
EMBO Molecular Medicine (2024)
-
Durable responses to alectinib in murine models of EML4-ALK lung cancer requires adaptive immunity
npj Precision Oncology (2023)
-
ALK-positive lung cancer: a moving target
Nature Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.