Abstract
The prominent and evolutionarily ancient role of the plant hormone auxin is the regulation of cell expansion1. Cell expansion requires ordered arrangement of the cytoskeleton2 but molecular mechanisms underlying its regulation by signalling molecules including auxin are unknown. Here we show in the model plant Arabidopsis thaliana that in elongating cells exogenous application of auxin or redistribution of endogenous auxin induces very rapid microtubule re-orientation from transverse to longitudinal, coherent with the inhibition of cell expansion. This fast auxin effect requires auxin binding protein 1 (ABP1) and involves a contribution of downstream signalling components such as ROP6 GTPase, ROP-interactive protein RIC1 and the microtubule-severing protein katanin. These components are required for rapid auxin- and ABP1-mediated re-orientation of microtubules to regulate cell elongation in roots and dark-grown hypocotyls as well as asymmetric growth during gravitropic responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Perrot-Rechenmann, C. Cellular responses to auxin: division versus expansion. Cold Spring Harb. Perspect. Biol. 2, a001446 (2010)
Sedbrook, J. C. & Kaloriti, D. Microtubules, MAPs and plant directional cell expansion. Trends Plant Sci. 13, 303–310 (2008)
Chapman, E. J. & Estelle, M. Mechanism of auxin-regulated gene expression in plants. Annu. Rev. Genet. 43, 265–285 (2009)
Lucas, J. & Shaw, S. L. Cortical microtubule arrays in the Arabidopsis seedling. Curr. Opin. Plant Biol. 11, 94–98 (2008)
Blancaflor, E. B. The cytoskeleton and gravitropism in higher plants. J. Plant Growth Regul. 21, 120–136 (2002)
Lindeboom, J. J. et al. A mechanism for reorientation of cortical microtubule arrays driven by microtubule severing. Science 342, 1245533 (2013)
Ubeda-Tomas, S., Beemster, G. T. & Bennett, M. J. Hormonal regulation of root growth: integrating local activities into global behaviour. Trends Plant Sci. 17, 326–331 (2012)
Marc, J. et al. A GFP-MAP4 reporter gene for visualizing cortical microtubule rearrangements in living epidermal cells. Plant Cell 10, 1927–1940 (1998)
Chan, J., Calder, G., Fox, S. & Lloyd, C. Cortical microtubule arrays undergo rotary movements in Arabidopsis hypocotyl epidermal cells. Nature Cell Biol. 9, 171–175 (2007)
Collett, C. E., Harberd, N. P. & Leyser, O. Hormonal interactions in the control of Arabidopsis hypocotyl elongation. Plant Physiol. 124, 553–562 (2000)
Simon, S. et al. Defining the selectivity of processes along the auxin response chain: a study using auxin analogues. New Phytol. 200, 1034–1048 (2013)
Morita, M. T. Directional gravity sensing in gravitropism. Annu. Rev. Plant Biol. 61, 705–720 (2010)
Chan, J., Calder, G. M., Doonan, J. H. & Lloyd, C. W. EB1 reveals mobile microtubule nucleation sites in Arabidopsis. Nature Cell Biol. 5, 967–971 (2003)
Brunoud, G. et al. A novel sensor to map auxin response and distribution at high spatio-temporal resolution. Nature 482, 103–106 (2012)
Tromas, A. et al. Auxin-binding protein 1 is a negative regulator of the SCFTIR1/AFB pathway. Nature Commun. 4, 2496 (2013)
Xu, T. et al. Cell surface- and rho GTPase-based auxin signaling controls cellular interdigitation in Arabidopsis. Cell 143, 99–110 (2010)
Robert, S. et al. ABP1 mediates auxin inhibition of clathrin-dependent endocytosis in Arabidopsis. Cell 143, 111–121 (2010)
Dharmasiri, N. et al. Plant development is regulated by a family of auxin receptor F box proteins. Dev. Cell 9, 109–119 (2005)
Braun, N. et al. Conditional repression of AUXIN BINDING PROTEIN1 reveals that it coordinates cell division and cell expansion during postembryonic shoot development in Arabidopsis and tobacco. Plant Cell 20, 2746–2762 (2008)
Tromas, A. et al. The AUXIN BINDING PROTEIN 1 is required for differential auxin responses mediating root growth. PLoS ONE 4, e6648 (2009)
Hayashi, K. et al. Small-molecule agonists and antagonists of F-box protein-substrate interactions in auxin perception and signaling. Proc. Natl Acad. Sci. USA 105, 5632–5637 (2008)
Shishova, M. & Lindberg, S. A new perspective on auxin perception. J. Plant Physiol. 167, 417–422 (2010)
Uyttewaal, M. et al. Mechanical stress acts via katanin to amplify differences in growth rate between adjacent cells in Arabidopsis. Cell 149, 439–451 (2012)
Lin, D. et al. Rho GTPase signaling activates microtubule severing to promote microtubule ordering in Arabidopsis. Curr. Biol. 23, 290–297 (2013)
Chen, X. et al. ABP1 and ROP6 GTPase signaling regulate clathrin-mediated endocytosis in Arabidopsis roots. Curr. Biol. 22, 1326–1332 (2012)
Paciorek, T. et al. Auxin inhibits endocytosis and promotes its own efflux from cells. Nature 435, 1251–1256 (2005)
Montagnac, G. et al. αTAT1 catalyses microtubule acetylation at clathrin-coated pits. Nature 502, 567–570 (2013)
Boudaoud, A. et al. FibrilTool, an ImageJ plug-in to quantify fibrillar structures in raw microscopy images. Nature Protocols 9, 457–463 (2014)
Salaycik, K. J., Fagerstrom, C. J., Murthy, K., Tulu, U. S. & Wadsworth, P. Quantification of microtubule nucleation, growth and dynamics in wound-edge cells. J. Cell Sci. 118, 4113–4122 (2005)
Leblanc, N. et al. A novel immunological approach establishes that the auxin-binding protein, Nt-abp1, is an element involved in auxin signaling at the plasma membrane. J. Biol. Chem. 274, 28314–28320 (1999)
Acknowledgements
We thank R. Dixit for performing complementary experiments, D. W. Ehrhardt and T. Hashimoto for providing the seeds of TUB6–RFP and EB1b–GFP respectively, E. Zazimalova, J. Petrasek and M. Fendrych for discussing the manuscript and J. Leung for text optimization. This work was supported by the European Research Council (project ERC-2011-StG-20101109-PSDP, to J.F.), ANR blanc AuxiWall project (ANR-11-BSV5-0007, to C.P.-R. and L.G.) and the Agency for Innovation by Science and Technology (IWT) (to H.R.). This work benefited from the facilities and expertise of the Imagif Cell Biology platform (http://www.imagif.cnrs.fr), which is supported by the Conseil Général de l’Essonne.
Author information
Authors and Affiliations
Contributions
X.C., L.G., C.P.-R. and J.F. conceived the study and designed experiments. X.C. performed experiments in roots, and L.G. performed experiments in hypocotyls. H.L., S.P. and A.A. assisted in microscopy and data generation. H.R. generated partial double mutants. R.H. did bioinformatics analysis. E.B. helped with discussion of the data. X.C., L.G., C.P.-R. and J.F. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Auxin induces microtubule rearrangement in root cells.
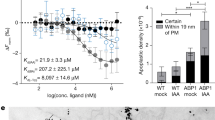
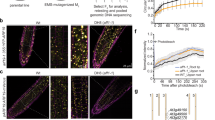
a, Schematic diagram of root and dark-grown hypocotyl growth. The growth direction of the root and hypocotyl is named as the cell growth axis. The observed cells for microtubules array were in the transition zone (highlighted by red line) of roots and in the elongation zone of dark-grown hypocotyls (highlighted by grey frame). The arrays of microtubules in root and hypocotyl were depicted for the expanding cells. b–f, MAP4–GFP or TUA6–RFP visualization of microtubule orientation in roots was performed by time-lapse observation (every 10 min; prime (′), minutes) following 100 nM NAA or IAA treatment, and deviated angles of individual microtubules were quantified as transverse microtubules (90 ± 30°) or longitudinal microtubules (0–60°/120–180°). In c and f, Student’s t-test was calculated for transverse microtubules compared with untreated roots (*P < 0.05; **P < 0.001). g, h, MAP4–GFP visualization and quantification of microtubule orientation in roots after 1 µM 2-NAA treatment for 60 min or after transfer of seedlings on acidified 1/2 Murashige and Skoog medium at pH 4.9 for 30, 90 and 180 minutes. Student’s t-test was calculated for transverse microtubules in treated samples compared with 1/2 Murashige and Skoog medium (pH 5.8) growing roots used as controls (**P < 0.001). i–k, Auxin distribution approximated by DII-VENUS at the lower side (LS) and upper side (US) of 90° re-oriented WT root (in DII-VENUS background). Enlarged pictures (i) are shown as the frames highlighted (k). Signal intensity is represented by the colour code as indicated. The relative signal for the upper side and lower side (j) is expressed compared with the signal in the respective frame before gravistimulation. Student’s t-test was calculated for the signal between the upper and lower sides at each time point (**P < 0.001). In all panels, average values are shown and error bars are s.e.m. Scale bars, 5 μm (b, d, e, g) and 30 μm (k).
Extended Data Figure 2 Functional inactivation of ABP1 results in microtubule defects gradually increasing with time of ABP1 inactivation.
a, b, MAP4–GFP visualization of microtubule orientation in WT and tir1-1 afb1-1 afb2-1 afb3-1 (abbreviated as tir1afb1,2,3) seedlings following 100 nM NAA treatment for 60 min. The proportion of cells with the four categories of microtubule orientation patterns was determined, and Student’s t-test was calculated for the category of transverse microtubule compared with WT treated in the same condition (**P < 0.001). c–f, MAP4–GFP visualization and quantification of microtubule orientation in roots (c, d) or dark-grown hypocotyls (e, f) of WT, SS12S and SS12K seedlings following different times of ethanol induction as indicated. Student’s t-test was calculated for the transverse microtubules compared with WT exposed for the same time to ethanol vapours as the conditional ABP1 lines (*P < 0.05, **P < 0.001). In all panels, average values are shown and error bars are s.e.m. Scale bars, 5 μm (a, c) and 10 μm (e).
Extended Data Figure 3 ABP1 is involved in microtubule rearrangement following gravistimulation.
a, Rearrangement of microtubules at the lower side compared with the upper side of 90° re-oriented roots of WT, SS12S, SS12K and abp1-5 (all expressing MAP4–GFP). Two different types of microtubule orientation (90 ± 30° or 0–60°/120–180°) were quantified. Student’s t-test was calculated for the category of transverse microtubules compared with each 0′ time point and calculated for transverse microtubules in the lower side compared with the upper side at each time point (**P < 0.001). b, c, Auxin distribution simulated by DII-VENUS at the lower side compared with the upper side of 90° re-oriented roots of SS12S and SS12K (all in DII-VENUS background; enlarged pictures were visualized in the frames highlighted). Image stacks were taken every 10 minutes, in total for 60 minutes. The ratio of the lower side signal divided by that of the upper side is shown in the chart (c). Student’s t-test was calculated for the signal ratio at each time point of SS12S/K compared with WT (**P < 0.001). Signal intensity is represented by the colour code as indicated. Data for SS12S and SS12K (b) are compared with WT (Extended Data Fig. 1i–k). d, The deviated angles of 90° gravistimulated-roots of WT, abp1-5, SS12S and SS12K seedlings were calculated every 30 min, in total for 8 h (Student’s t-test, *P < 0.05, **P < 0.001). In all panels, average values are shown and error bars are s.e.m. Scale bars, 5 μm (a) and 30 μm (b).
Extended Data Figure 4 The effect of auxin on fast responsiveness of microtubule dynamics is dependent on ABP1.
a, b, Acquisition and quantification of the rate of EB1b movement in roots of untreated or 100 nM NAA-treated (60 min) WT or SS12K (expressing EB1b–GFP) by measuring EB1b–GFP growth events as highlighted by red lines (Student’s t-test, P > 0.05). Box plots indicate the 25th centile (bottom boundary), median (middle line), 75th centile (top boundary), the nearest observations within 1.5 times, the interquartile range and outliers. c, EB1b movement was simulated as transverse (blue, 90 ± 30°) or longitudinal (red, 0–60°/120–180°) trajectories before (0′) and after (180′′) 100 nM NAA treatment in WT background (colour maps). The blue/red surface ratio is quantified on the chart (n = 5); c corresponds to Fig. 3a. d, Microtubule orientation patterns after 400 µM cordycepin plus NAA co-treatment. Student’s t-test was calculated for the category of transverse microtubule compared with only cordycepin treatment (**P < 0.001). e, EB1b trajectories (simulated by time-stack from 10 min videos) were visualized and quantified after DMSO, IAA (1 µM), PEO-IAA (10 µM) and PEO-IAA (10 µM) plus IAA (1 µM) treatments. The left panel shows successive frames of 90′ acquisitions following IAA application of pre-treated PEO-IAA WT roots. Student’s t-test was calculated for the category of transverse microtubules compared with DMSO treatment at each time point (**P < 0.001). f–i, Projections of EB1b–GFP in SS12K roots (f) and quantification (g) from every 15 s acquisition during 10 min (Supplementary Videos 4 and 6) following DMSO or 100 nM NAA application (n = 10). Blue and red strips represent transverse (90 ± 30°) and oblique/longitudinal (0–60°/120–180°) directions, respectively (f). Colour maps show the simulated transverse or longitudinal trajectories of EB1b before (0′) and after (180′′) 100 nM NAA treatment in SS12K (h) or SS12S (i) roots. The blue/red surface ratio is quantified on the charts (n = 5) (h, i). The data of SS12S (i) correspond to Fig. 3b, and the data of SS12S and SS12K (f–i) are compared with WT (Fig. 3a and Extended Data Fig. 4c). In all panels except b, average values are shown, error bars are s.e.m. and scale bars are 5 μm.
Extended Data Figure 5 Overexpressed ABP1-induced effect of auxin on fast responsiveness of microtubule dynamics.
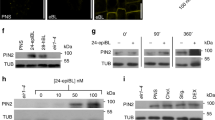
a–c, ABP1 and ABP1–GFP transcripts (a) and ABP1 protein level (b, c) were detected in WT and XVE ≫ ABP1-OE line before and after 2 µM oestradiol induction for 12 h or 48 h before RNA or protein extraction. The transcript levels of ABP1 in WT with DMSO treatment were standardized as ‘1’ (a). The 22 kDa native ABP1 band and 49KDa ABP1–GFP band were detected and quantified in the right chart. The protein level of native ABP1 or ABP1–GFP in WT was standardized as ‘1’ for each ABP1 and ABP1–GFP, respectively (b, c). Student’s t-test, **P < 0.001. d, Time-lapse observation of microtubule orientation in the roots of XVE ≫ ABP1-OE roots expressing TUA6–RFP, WT and abp1-5 (both expressing MAP4–GFP) upon 100 nM NAA treatment. The percentage of re-oriented microtubules (0–60°/120–180°) was quantified. Re-oriented microtubules in the inducible XVE ≫ ABP1-OE TUA6–RFP roots were calculated compared with none-inducible roots, and abp1-5 MAP4–GFP was compared with MAP4–GFP at each time point (Student’s t-test, *P < 0.05, **P < 0.001). In all panels, average values are shown, error bars are s.e.m and scale bars are 5 μm.
Extended Data Figure 6 Calcium starvation disrupts microtubule orientation and high calcium increases microtubule depolymerization.
Orientation and polymerization statuses of microtubules were visualized following transfer of seedlings to different concentrations of CaCl2 for 30, 90 or 180 min. Low calcium levels disrupted microtubule organization, leading to a predominantly random pattern; high calcium caused microtubule depolymerization. Student’s t-test was calculated for the category of transverse microtubules compared with seedlings grown and transferred on standard 1/2 Murashige and Skoog medium (with 1.5 mM CaCl2) (*P < 0.05, **P < 0.001). In all panels, average values are shown, error bars are s.e.m. and scale bars are 5 μm.
Extended Data Figure 7 Auxin–ABP1 controls microtubule arrangement through downstream ROP6–RIC1–KTN1 signalling.
a, MAP4–GFP visualization of microtubule orientation in the root of WT, rop6-1, ric1-1, SS12S ric1-1 and SS12K ric1-1 following DMSO application for 60 min. Pictures in a correspond to quantifications in Fig. 4a. b, c, Microtubule re-orientation patterns were visualized by MAP4–GFP in the roots of WT and rop6-1+/− following DMSO or 100 nM NAA application for 60 min (Student’s t-test, P > 0.05). d, Transcript level of the scFv12 coding the recombinant antibody responsible for ABP1 knockdown in WT, ric1-1, ktn1, SS12S, SS12K, SS12S ric1-1, SS12K ric1-1, SS12S ktn1 and SS12K ktn1 after 48 h ethanol induction. The transcript level of the scFv12 in SS12S was standardized as ‘1’ (Student’s t-test, P > 0.05). e, Microtubule orientation by MAP4–GFP in dark-grown hypocotyls of WT, SS12K, ktn1 and SS12K ktn1 (with 24 h ethanol induction) following DMSO application for 60 min. Pictures in e correspond to Fig. 4b. In all panels, average values are shown and error bars are s.e.m. Scale bars, 5 μm (a, b) and 10 μm (e).
Supplementary information
The trajectories of EB1b in WT background following DMSO treatment
EB1b-GFP seedlings were mounted on DMSO-contained 1/2 MS glass slides and imaged immediately for 10min. EB1b-GFP comets illustrate major transversal MT growth trajectories (90±30°). Corresponding to Fig. 3a. (MOV 3725 kb)
The trajectories of EB1b in WT background following 100nM NAA treatment
EB1b-GFP seedlings were mounted on 1/2 MS glass slides containing 100nM NAA and imaged immediately for 10min. EB1b-GFP moves mainly along 90±30° transversal direction in the beginning (0-60sec), while increasing EB1b tracks switch along oblique/longitudinal direction (0-60°/120-180°) after 75sec. Corresponding to Fig. 3a. (MOV 2184 kb)
The trajectories of EB1b in ABP1 knockdown lines following DMSO treatment
48h ethanol induced SS12S or SS12K seedlings expressing EB1b-GFP were mounted on DMSO-contained 1/2 MS glass slides and imaged immediately for 10min. High proportion of EB1b-GFP moves in oblique/longitudinal directions. Corresponding to Fig. 3b, Extended Data Fig. 4d. (MOV 1813 kb)
The trajectories of EB1b in ABP1 knockdown lines following DMSO treatment
48h ethanol induced SS12S or SS12K seedlings expressing EB1b-GFP were mounted on DMSO-contained 1/2 MS glass slides and imaged immediately for 10min. High proportion of EB1b-GFP moves in oblique/longitudinal directions. Corresponding to Fig. 3b, Extended Data Fig. 4d. (MOV 1383 kb)
The trajectories of EB1b in SS12S or SS12K background following 100nM NAA treatment
48h ethanol induced SS12S or SS12K seedlings expressing EB1b-GFP were mounted on 1/2 MS glass slides containing 100nM NAA and imaged immediately for 10min. Compared with WT, no consistent switch of EB1b trajectories to longitudinal directions but only few stochastic changes were observed. Corresponding to Fig. 3b, Extended Data Fig. 4d. (MOV 1717 kb)
The trajectories of EB1b in SS12S or SS12K background following 100nM NAA treatment
48h ethanol induced SS12S or SS12K seedlings expressing EB1b-GFP were mounted on 1/2 MS glass slides containing 100nM NAA and imaged immediately for 10min. Compared with WT, no consistent switch of EB1b trajectories to longitudinal directions but only few stochastic changes were observed. Corresponding to Fig. 3b, Extended Data Fig. 4d. (MOV 2100 kb)
Rights and permissions
About this article
Cite this article
Chen, X., Grandont, L., Li, H. et al. Inhibition of cell expansion by rapid ABP1-mediated auxin effect on microtubules. Nature 516, 90–93 (2014). https://doi.org/10.1038/nature13889
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13889
This article is cited by
-
New PEO-IAA-Inspired Anti-Auxins: Synthesis, Biological Activity, and Possible Application in Hemp (Cannabis Sativa L.) Micropropagation
Journal of Plant Growth Regulation (2023)
-
In silico analysis of key regulatory networks related to microfibril angle in Populus trichocarpa Hook.
Biologia (2022)
-
The plant AlcR-pAlcA ethanol-inducible system displays gross growth artefacts independently of downstream pAlcA-regulated inducible constructs
Scientific Reports (2021)
-
Cell surface and intracellular auxin signalling for H+ fluxes in root growth
Nature (2021)
-
Mathematical Modelling of Auxin Transport in Plant Tissues: Flux Meets Signalling and Growth
Bulletin of Mathematical Biology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.