Abstract
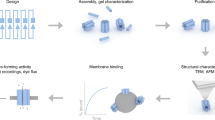
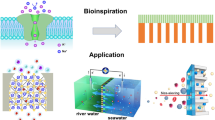
There is much interest in developing synthetic analogues of biological membrane channels1 with high efficiency and exquisite selectivity for transporting ions and molecules. Bottom-up2 and top-down3 methods can produce nanopores of a size comparable to that of endogenous protein channels, but replicating their affinity and transport properties remains challenging. In principle, carbon nanotubes (CNTs) should be an ideal membrane channel platform: they exhibit excellent transport properties4,5,6,7,8 and their narrow hydrophobic inner pores mimic structural motifs typical of biological channels1. Moreover, simulations predict that CNTs with a length comparable to the thickness of a lipid bilayer membrane can self-insert into the membrane9,10. Functionalized CNTs have indeed been found to penetrate lipid membranes and cell walls11,12, and short tubes have been forced into membranes to create sensors13, yet membrane transport applications of short CNTs remain underexplored. Here we show that short CNTs spontaneously insert into lipid bilayers and live cell membranes to form channels that exhibit a unitary conductance of 70–100 picosiemens under physiological conditions. Despite their structural simplicity, these ‘CNT porins’ transport water, protons, small ions and DNA, stochastically switch between metastable conductance substates, and display characteristic macromolecule-induced ionic current blockades. We also show that local channel and membrane charges can control the conductance and ion selectivity of the CNT porins, thereby establishing these nanopores as a promising biomimetic platform for developing cell interfaces, studying transport in biological channels, and creating stochastic sensors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Sui, H., Han, B.-G., Lee, J. K., Walian, P. & Jap, B. K. Structural basis of water-specific transport through the AQP1 water channel. Nature 414, 872–878 (2001)
Langecker, M. et al. Synthetic lipid membrane channels formed by designed DNA nanostructures. Science 338, 932–936 (2012)
Li, J. et al. Ion-beam sculpting at nanometre length scales. Nature 412, 166–169 (2001)
Hinds, B. J. et al. Aligned multiwalled carbon nanotube membranes. Science 303, 62–65 (2004)
Liu, H. et al. Translocation of single-stranded DNA through single-walled carbon nanotubes. Science 327, 64–67 (2010)
Lee, C. Y., Choi, W., Han, J.-H. & Strano, M. S. Coherence resonance in a single-walled carbon nanotube ion channel. Science 329, 1320–1324 (2010)
Zimmerli, U. & Koumoutsakos, P. Simulations of electrophoretic RNA transport through transmembrane carbon nanotubes. Biophys. J. 94, 2546–2557 (2008)
Hummer, G., Rasaiah, J. C. & Noworyta, J. P. Water conduction through the hydrophobic channel of a carbon nanotube. Nature 414, 188–190 (2001)
Lopez, C. F., Nielsen, S. O., Moore, P. B. & Klein, M. L. Understanding nature's design for a nanosyringe. Proc. Natl Acad. Sci. USA 101, 4431–4434 (2004)
Dutt, M., Nayhouse, M. J., Kuksenok, O., Little, S. R. & Balazs, A. C. Interactions of end-functionalized nanotubes with lipid vesicles: spontaneous insertion and nanotube self-organization. Curr. Nanosci. 7, 699–715 (2011)
Lacerda, L. et al. How do functionalized carbon nanotubes land on, bind to and pierce through model and plasma membranes. Nanoscale 5, 10242–10250 (2013)
Kam, N. W. S., O’Connell, M., Wisdom, J. A. & Dai, H. Carbon nanotubes as multifunctional biological transporters and near-infrared agents for selective cancer cell destruction. Proc. Natl Acad. Sci. USA 102, 11600–11605 (2005)
Liu, L., Yang, C., Zhao, K., Li, J. & Wu, H.-C. Ultrashort single-walled carbon nanotubes in a lipid bilayer as a new nanopore sensor. Nature Commun. 4, http://dx.doi.org/10.1038/ncomms3989 (2013)
Sun, X. et al. Optical properties of ultrashort semiconducting single-walled carbon nanotube capsules down to sub-10 nm. J. Am. Chem. Soc. 130, 6551–6555 (2008)
Le Duc, Y. et al. Imidazole-quartet water and proton dipolar channels. Angew. Chem. Int. Ed. 50, 11366–11372 (2011)
Fornasiero, F. et al. Ion exclusion by sub 2-nm carbon nanotube pores. Proc. Natl Acad. Sci. USA 105, 17250–17255 (2008)
Walther, J. H., Ritos, K., Cruz-Chu, E. R., Megaridis, C. M. & Koumoutsakos, P. Barriers to superfast water transport in carbon nanotube membranes. Nano Lett. 13, 1910–1914 (2013)
Gu, L.-Q. & Bayley, H. Interaction of the noncovalent molecular adapter, β-cyclodextrin, with the staphylococcal α-hemolysin pore. Biophys. J. 79, 1967–1975 (2000)
Nestorovich, E. M., Rostovtseva, T. K. & Bezrukov, S. M. Residue ionization and ion transport through OmpF channels. Biophys. J. 85, 3718–3729 (2003)
Powell, M. R. et al. Nanoprecipitation-assisted ion current oscillations. Nature Nanotechnol. 3, 51–57 (2007)
Powell, M. R., Cleary, L., Davenport, M., Shea, K. J. & Siwy, Z. S. Electric-field-induced wetting and dewetting in single hydrophobic nanopores. Nature Nanotechnol. 6, 798–802 (2011)
Lev, A. A. et al. Rapid switching of ion current in narrow pores: implications for biological ion channels. Proc. R. Soc. Lond. B 252, 187–192 (1993)
Shimizu, S. et al. Stochastic pore blocking and gating in PDMS–glass nanopores from vapor–liquid phase transitions. J. Phys. Chem. C 117, 9641–9651 (2013)
Buyukdagli, S., Manghi, M. & Palmeri, J. Ionic capillary evaporation in weakly charged nanopores. Phys. Rev. Lett. 105, 158103 (2010)
Kasianowicz, J. J., Brandin, E., Branton, D. & Deamer, D. W. Characterization of individual polynucleotide molecules using a membrane channel. Proc. Natl Acad. Sci. USA 93, 13770–13773 (1996)
Hall, A. R. et al. Hybrid pore formation by directed insertion of α-haemolysin into solid-state nanopores. Nature Nanotechnol. 5, 874–877 (2010)
Venkatesan, B. M. & Bashir, R. Nanopore sensors for nucleic acid analysis. Nature Nanotechnol. 6, 615–624 (2011)
Majumder, M., Chopra, N. & Hinds, B. J. Effect of tip functionalization on transport through vertically oriented carbon nanotube membranes. J. Am. Chem. Soc. 127, 9062–9070 (2005)
Haque, F., Geng, J., Montemagno, C. & Guo, P. Incorporation of a viral DNA-packaging motor channel in lipid bilayers for real-time, single-molecule sensing of chemicals and double-stranded DNA. Nature Protocols 8, 373–392 (2013)
Neher, E., Sandblom, J. & Eisenman, G. Ionic selectivity, saturation, and block in gramicidin A channels. J. Membr. Biol. 40, 97–116 (1978)
Frolov, V. A., Lizunov, V. A., Dunina-Barkovskaya, A. Y., Samsonov, A. V. & Zimmerberg, J. Shape bistability of a membrane neck: a toggle switch to control vesicle content release. Proc. Natl Acad. Sci. USA 100, 8698–8703 (2003)
Shnyrova, A. V. et al. Geometric catalysis of membrane fission driven by flexible dynamin rings. Science 339, 1433–1436 (2013)
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using µManager. Curr. Protoc. Molec. Biol. 92, 14.20.1–14.20.17 (2010)
Acknowledgements
Parts of this work were supported by the US Department of Energy, Office of Basic Energy Sciences, Division of Materials Sciences and Engineering (characterization and transport studies), and the LDRD programme at LLNL, 12-ERD-073 (synthesis). R.T. acknowledges support from the LSP programme at LLNL. D.M. acknowledges support from an ROTC summer fellowship. V.A.F. acknowledges partial support by the Spanish Ministry of Economy and Competitiveness, grant BFU2012-34885, co-financed with European FEDER funds, and the Basque Government, grant IE12-332. Work at LLNL was performed under the auspices of the US Department of Energy under contract DE-AC52-07NA27344. Work at the Molecular Foundry was supported by the Office of Science, Office of Basic Energy Sciences, of the US Department of Energy under contract DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
J.G. performed and analysed (with D.M.) the planar lipid bilayer transport measurements; K.K. performed bulk transport studies; J.Z., K.K., J.G. and R.T. performed the synthesis and purification, K.R.C. performed AFM analysis, Y.M.W., L.R.C., F.I.A. and K.K. performed TEM analysis; L.R.C. and F.I.A. performed cryo-TEM analysis; A.N. conceived and directed the research, and wrote the manuscript draft; A.E., A.V.S. and V.A.F. designed the cell and GUV reconstitution experiments, and A.E. and A.V.S. performed them. All authors contributed to the data analysis, discussion, and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 TEM images of isolated CNTs.
a–c, Purified CNTs before cutting (a and b, conventional TEM, c imaged on a cryo stage; all air-dried samples). d, Magnified image of one CNT from c. e–g, CNTs mixed with lipid and lightly sonicated to aid nanotube dispersion. h–m, Isolated CNT porins after the cutting procedure.
Extended Data Figure 2 Length distributions of CNT porins measured from TEM images.
a, CNT porin sample after cutting and purification procedures (n = 65). b, Cut and purified sample after passing through a 100 nm pore filter membrane (n = 36). c, As b but after passing through a 50 nm pore filter membrane (n = 156). µ indicates the mean CNT porin length.
Extended Data Figure 3 Characterization of CNT porins.
a, Histogram of the diameters of short CNT fragments measured with AFM. Most CNT porins show diameters between 1 and 2 nm. Inset, high-magnification AFM image of a single CNT on a bare mica surface. b, Histogram of the diameters of CNT porins inserted into the lipid vesicles measured by cryo-TEM. c, Raman spectrum of the short-CNT/lipid complex after 16 h of sonication-assisted cutting. Inset, magnified view of the radial breathing mode region of the CNT spectrum (100–300 cm−1).
Extended Data Figure 4 Cryo-TEM images of CNT–liposome complexes.
Original large field-of-view images (main panels); insets, magnified views of the selected structures, with false-colour lines added to highlight the lipid bilayer (red) and CNT porins (blue).
Extended Data Figure 5 High magnification cryo-TEM imaged of CNT–liposome complexes.
Panels (left to right) show unprocessed, image-processed, and colourized unprocessed examples. See Methods for details of image processing.
Extended Data Figure 6 Single CNT porin conductance at different electrolyte concentrations.
Plots of the average single CNT porin conductance at different values of background electrolyte (KCl) concentrations at pH 7 (red circles) and pH 2 (blue squares). The slopes of the linear fit through the data (dashed lines) correspond to 0.66 nS M−1 at pH 7 and 0.36 nS M−1 at pH 2. Error bars, s.d.; n values as follows (pH = 7: 0.15 M, n = 6; 0.5 M, n = 161; 1 M, n = 39; 2 M, n = 8. pH = 2: 0.5 M, n = 18; 1 M, n = 3; 2 M, n = 36).
Extended Data Figure 7 Single CNT porin conductance at pH 2.
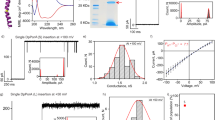
a, Representative conductance traces showing individual CNT porin incorporation at pH 2 and 2 M KCl background electrolyte concentration. Traces i–iv represent incorporation from 4 independent experiments. b, Histogram of conductance values measured for 109 individual CNT channel incorporation events (n = 109). Solid blue line corresponds to the fit of the data to a distribution expected for multiple channel incorporation, where the position and width of the second peak are determined from the position, μ (0.36 ± 0.006 nS), and width, σ (0.10 ± 0.008 nS), of the first peak as 2μ and  . c, Representative conductance traces showing individual CNT porin incorporation at pH 2 and 0.5 M KCl background electrolyte. Traces i–iv represent incorporation from 4 independent experiments. Control trace shows a conductance trace collected at the same conditions without adding CNT porins.
. c, Representative conductance traces showing individual CNT porin incorporation at pH 2 and 0.5 M KCl background electrolyte. Traces i–iv represent incorporation from 4 independent experiments. Control trace shows a conductance trace collected at the same conditions without adding CNT porins.
Extended Data Figure 8 Representative traces and corresponding current level histograms of CNT porin incorporation, gating and DNA translocation in lipid bilayers at pH 7.
a, Control conductance trace recorded in the absence of CNT porin. Although only the first 5 min are shown, the current level remained stable for up to 80 min (n = 3 × 106). b, c, Conductance traces and histograms showing single CNT porin incorporation events and subsequent current fluctuations between gating sub-levels (n > 5 × 104). d, Trace and conductance value histogram showing CNT porin incorporation event and subsequent two DNA translocation events (indicated by black arrows). Here n = 3.6 × 105. The peaks on the histogram correspond to 0.01 nS for pre-incorporation baseline, 0.22 nS for DNA blockade, and 0.67 nS for open pore conductance, which gives the DNA blockade of 0.45 nS and a blockade ratio of 67%. Dashed lines are added to all traces as a guide to the eye.
Extended Data Figure 9 CNT porin conductance in cellular membranes.
a, Schematic of the on-cell patch clamp measurement. b, c, On-cell conductance measurements demonstrate no channel activity (b) and appearance of a small flickering channel (c) in control experiments with HEK293T cells. Holding potentials were −60 mV (b) and +60 mV (c), the time order is from left to right, top to bottom. d, In the presence of CNT porin (0.15% dilution of the stock), large channel activity is detected, holding potential −60 mV. e, Metastable conductance sub-states of 50 pS detected with CNT porin incorporated into the plasma membrane of CHO cell, holding potential −60 mV. f, Mean patch conductance (on cell configuration, 0.15% CNT stock solution in the patch pipette) detected in CHO and HEK293T cells at positive and negative holding potentials. Error bar, s.d.; 4 cells). In ∼70% of HEK293T cells and 95% of CHO cells we did not observe significant dependence of the conductance on the holding potential; only those cells were used for the conductance analysis.
Extended Data Figure 10 CNT porin conductance in patches isolated from planar lipid bilayers and giant unilamellar vesicles.
a, Histogram of conductance values measured for CNT porin incorporation into a planar lipid bilayer mimicking the composition of a charged plasma membrane (red bars, n = 681, 5 membranes, 3 controls without CNT porins showed no events) and a similar lipid bilayer without charged species (black bars, n = 540, 5 membranes, 3 controls without CNT porins showed no events). The background electrolyte concentration was kept at 150 mM. Solid lines correspond to the fit of the data to a distribution expected for multiple channel incorporation. The primary peak positions are 65.1 ± 0.4 pS (charged bilayer) and 92.5 ± 0.6 pS (uncharged bilayer). b, Histogram of conductance values measured for CNT porins reconstituted into giant unilamellar vesicles (GUVs) mimicking the composition of charged plasma membrane (red bars, n = 40, 3 membranes), Solid lines correspond to the fit of the data to a distribution expected for multiple channel incorporation with the primary conductance peak at 60 ± 2 pS. c, Representative control experiment demonstrating lack of spontaneous pore formation in the bilayer patch. d, e, Patch-clamp measurements of the CNT porin incorporation events in planar lipid bilayer (d) and GUV membrane (e).
Rights and permissions
About this article
Cite this article
Geng, J., Kim, K., Zhang, J. et al. Stochastic transport through carbon nanotubes in lipid bilayers and live cell membranes. Nature 514, 612–615 (2014). https://doi.org/10.1038/nature13817
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13817
This article is cited by
-
Outer-surface Covering of Nanochannels with Hydrogel for Highly Sensitive and Specific Cr(VI) Detection Through Analyte-caused Charge Change in Hydrogel
Chemical Research in Chinese Universities (2024)
-
Transient water wires mediate selective proton transport in designed channel proteins
Nature Chemistry (2023)
-
High-resolution discrimination of homologous and isomeric proteinogenic amino acids in nanopore sensors with ultrashort single-walled carbon nanotubes
Nature Communications (2023)
-
Enhanced osmotic transport in individual double-walled carbon nanotube
Nature Communications (2023)
-
Polymer-coated carbon nanotube hybrids with functional peptides for gene delivery into plant mitochondria
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.