Abstract
Non-caloric artificial sweeteners (NAS) are among the most widely used food additives worldwide, regularly consumed by lean and obese individuals alike. NAS consumption is considered safe and beneficial owing to their low caloric content, yet supporting scientific data remain sparse and controversial. Here we demonstrate that consumption of commonly used NAS formulations drives the development of glucose intolerance through induction of compositional and functional alterations to the intestinal microbiota. These NAS-mediated deleterious metabolic effects are abrogated by antibiotic treatment, and are fully transferrable to germ-free mice upon faecal transplantation of microbiota configurations from NAS-consuming mice, or of microbiota anaerobically incubated in the presence of NAS. We identify NAS-altered microbial metabolic pathways that are linked to host susceptibility to metabolic disease, and demonstrate similar NAS-induced dysbiosis and glucose intolerance in healthy human subjects. Collectively, our results link NAS consumption, dysbiosis and metabolic abnormalities, thereby calling for a reassessment of massive NAS usage.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gardner, C. et al. Nonnutritive sweeteners: current use and health perspectives. Diabetes Care 35, 1798–1808 (2012)
Fitch, C. & Keim, K. S. Position of the Academy of Nutrition and Dietetics: use of nutritive and nonnutritive sweeteners. Journal of the Academy of Nutrition and Dietetics 112, 739–758 (2012)
Tordoff, M. G. & Alleva, A. M. Effect of drinking soda sweetened with aspartame or high-fructose corn syrup on food intake and body weight. Am. J. Clin. Nutr. 51, 963–969 (1990)
Horwitz, D. L., McLane, M. & Kobe, P. Response to single dose of aspartame or saccharin by NIDDM patients. Diabetes Care 11, 230–234 (1988)
Nettleton, J. A. et al. Diet soda intake and risk of incident metabolic syndrome and type 2 diabetes in the Multi-Ethnic Study of Atherosclerosis (MESA). Diabetes Care 32, 688–694 (2009)
Roberts, A., Renwick, A. G., Sims, J. & Snodin, D. J. Sucralose metabolism and pharmacokinetics in man. Food Chem. Toxicol. 38 (Suppl. 2). 31–41 (2000)
Byard, J. L. & Goldberg, L. The metabolism of saccharin in laboratory animals. Food Cosmet. Toxicol. 11, 391–402 (1973)
Clemente, J. C., Ursell, L. K., Parfrey, L. W. & Knight, R. The impact of the gut microbiota on human health: an integrative view. Cell 148, 1258–1270 (2012)
Claesson, M. J. et al. Gut microbiota composition correlates with diet and health in the elderly. Nature 488, 178–184 (2012)
Muegge, B. D. et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332, 970–974 (2011)
Turnbaugh, P. J. et al. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444, 1027–1031 (2006)
Ley, R. E., Turnbaugh, P. J., Klein, S. & Gordon, J. I. Microbial ecology: human gut microbes associated with obesity. Nature 444, 1022–1023 (2006)
Qin, J. et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 490, 55–60 (2012)
Henao-Mejia, J. et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482, 179–185 (2012)
David, L. A. et al. Diet rapidly and reproducibly alters the human gut microbiome. Nature 505, 559–563 (2014)
Peterson, J. et al. The NIH human microbiome project. Genome Res. 19, 2317–2323 (2009)
Koropatkin, N. M., Cameron, E. A. & Martens, E. C. How glycan metabolism shapes the human gut microbiota. Nature Rev. Microbiol. 10, 323–335 (2012)
Schwiertz, A. et al. Microbiota and SCFA in lean and overweight healthy subjects. Obesity 18, 190–195 (2010)
Bergman, E. N. Energy contributions of volatile fatty acids from the gastrointestinal tract in various species. Physiol. Rev. 70, 567–590 (1990)
Karlsson, F. H. et al. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 498, 99–103 (2013)
Connor, S. C., Hansen, M. K., Corner, A., Smith, R. F. & Ryan, T. E. Integration of metabolomics and transcriptomics data to aid biomarker discovery in type 2 diabetes. Mol. Biosyst. 6, 909–921 (2010)
Cani, P. D. et al. Changes in gut microbiota control metabolic endotoxemia-induced inflammation in high-fat diet-induced obesity and diabetes in mice. Diabetes 57, 1470–1481 (2008)
Markle, J. G. et al. Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339, 1084–1088 (2013)
Sonnenburg, J. L. et al. Glycan foraging in vivo by an intestine-adapted bacterial symbiont. Science 307, 1955–1959 (2005)
Cani, P. D. et al. Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56, 1761–1772 (2007)
Smith, M. I. et al. Gut microbiomes of Malawian twin pairs discordant for kwashiorkor. Science 339, 548–554 (2013)
Taylor, J. D., Richards, R. K. & Wiegand, R. G. Toxicological studies with sodium cyclamate and saccharin. Food Cosmet. Toxicol. 6, 313–327 (1968)
Goldsmith, L. A. Acute and subchronic toxicity of sucralose. Food Chem. Toxicol. 38 (Suppl. 2). 53–69 (2000)
Magnuson, B. A. et al. Aspartame: a safety evaluation based on current use levels, regulations, and toxicological and epidemiological studies. Crit. Rev. Toxicol. 37, 629–727 (2007)
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature Methods 7, 335–336 (2010)
Elinav, E. et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145, 745–757 (2011)
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006)
Blecher-Gonen, R. et al. High-throughput chromatin immunoprecipitation for genome-wide mapping of in vivo protein–DNA interactions and epigenomic states. Nature Protocols 8, 539–554 (2013)
Qin, J. et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464, 59–65 (2010)
Marco-Sola, S., Sammeth, M., Guigó, R. & Ribeca, P. The GEM mapper: fast, accurate and versatile alignment by filtration. Nature Methods 9, 1185–1188 (2012)
Francis, O. E. et al. Pathoscope: species identification and strain attribution with unassembled sequencing data. Genome Res. 23, 1721–1729 (2013)
Kolodkin-Gal, I. et al. d-amino acids trigger biofilm disassembly. Science 328, 627–629 (2010)
Shahar, D., Fraser, D., Shai, I. & Vardi, H. Development of a food frequency questionnaire (FFQ) for an elderly population based on a population survey. J. Nutr. 133, 3625–3629 (2003)
Shahar, D., Shai, I., Vardi, H., Brener-Azrad, A. & Fraser, D. Development of a semi-quantitative Food Frequency Questionnaire (FFQ) to assess dietary intake of multiethnic populations. Eur. J. Epidemiol. 18, 855–861 (2003)
Shai, I. et al. Dietary evaluation and attenuation of relative risk: multiple comparisons between blood and urinary biomarkers, food frequency, and 24-hour recall questionnaires: the DEARR study. J. Nutr. 135, 573–579 (2005)
Acknowledgements
We thank the members of the Elinav and Segal laboratories for discussions. We acknowledge C. Bar-Nathan for germ-free mouse caretaking. We thank the Weizmann Institute management and the Nancy and Stephen Grand Israel National Center for Personalized Medicine (INCPM) for providing financial and infrastructure support. We thank G. Malka, N. Kosower and R. Bikovsky for coordinating the human clinical trials, and M. Pevsner-Fischer, T. Avnit-Sagi and M. Lotan-Pompan for assistance with microbiome sample processing. C.A.T. is the recipient of a Boehringer Ingelheim Fonds PhD Fellowship. G.Z.-S. is supported by the Morris Kahn Fellowships for Systems Biology. This work was supported by grants from the National Institute of Health (NIH) and the European Research Council (ERC) to E.S., and support and grants to E.E. provided by Y. and R. Ungar, the Abisch Frenkel Foundation for the Promotion of Life Sciences, the Gurwin Family Fund for Scientific Research, Leona M. and Harry B. Helmsley Charitable Trust, Crown Endowment Fund for Immunological Research, estate of J. Gitlitz, estate of L. Hershkovich, Rising Tide foundation, Minerva Stiftung foundation, and the European Research Council. E.E. is the incumbent of the Rina Gudinski Career Development Chair.
Author information
Authors and Affiliations
Contributions
J.S. conceived the project, designed and performed experiments, interpreted the results, and wrote the manuscript. T.K., D.Z. and G.Z.-S. performed the computational and metagenomic microbiota analysis and the analysis of the retrospective and prospective human study, and are listed alphabetically. C.A.T., O.M., A.W. and H.S. helped with experiments. Y.K. helped with the metabolic cage experiments. S.G. designed the metagenomic library protocols and generated the libraries. I.K.-G. performed the SCFA quantification experiments. D.I., N.Z., and Z.H. performed and supervised human experimentation. A.H. supervised the germ-free mouse experiments. E.S. and E.E. conceived and directed the project, designed experiments, interpreted the results, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
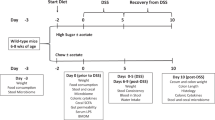
Extended Data Figure 1 Experimental scheme.
10-week-old C57Bl/6 male mice were treated with the following dietary regimes. a, Drinking commercially available non-caloric artificial sweeteners (NAS; saccharin, sucralose and aspartame) or glucose, sucrose or water as controls and fed a normal-chow (NC) diet. b, Drinking commercially available saccharin or glucose as control and fed a high-fat diet (HFD). c, Drinking pure saccharin or water and fed HFD. d, As in c, but with outbred Swiss-Webster mice. Glucose tolerance tests, microbiome analysis and supplementation of drinking water with antibiotics were performed on the indicated time points. e, Schematic of faecal transplant experiments.
Extended Data Figure 2 Artificial sweeteners induce glucose intolerance.
a, AUC of mice fed HFD and commercial saccharin (N = 10) or glucose (N = 9). b, AUC of HFD-fed mice drinking 0.1 mg ml−1 saccharin or water for 5 weeks (N = 20), followed by ‘antibiotics A’ (N = 10). c, d, OGTT and AUC of HFD-fed outbred Swiss-Webster mice (N = 5) drinking pure saccharin or water. e, f, Faecal samples were transferred from donor mice (N = 10) drinking commercially available, pure saccharin, glucose or water controls into 8-week-old male Swiss-Webster germ-free recipient mice. AUC of germ-free mice 6 days following transplant of microbiota from commercial saccharin- (N = 12) and glucose-fed mice (N = 11) (e); or pure saccharin- (N = 16) and water-fed (N = 16) donors (f). Symbols (GTT) or horizontal lines (AUC), means; error bars, s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, ANOVA and Tukey post hoc analysis (GTT) or unpaired two-sided Student t-test (AUC). Each experiment was repeated twice.
Extended Data Figure 3 Metabolic characterization of mice consuming commercial NAS formulations.
10-week-old C57Bl/6 mice (N = 4) were given commercially available artificial sweeteners (saccharin, sucralose and aspartame) or controls (water, sucrose or glucose, N = 4 in each group) and fed normal-chow diet. After 11 weeks, metabolic parameters were characterized using the PhenoMaster metabolic cages system for 80 h. Light and dark phases are denoted by white and black rectangles on the x-axis, respectively, and grey bars for the dark phase. a, Liquids intake. b, AUC of a. c, Chow consumption. d, AUC of c. e, Total caloric intake from chow and liquid during 72 h (see methods for calculation). f, Respiratory exchange rate (RER). g, AUC of f. h, Physical activity as distance. i, AUC of h. j, Energy expenditure. k, Mass change compared to original mouse weight during 15 weeks (N = 10). l, AUC of k. The metabolic cages characterization and weight-gain monitoring were repeated twice.
Extended Data Figure 4 Metabolic characterization of mice consuming HFD and pure saccharin or water.
10-week-old C57Bl/6 mice (N = 8) were fed HFD, with or without supplementing drinking water with 0.1 mg ml−1 pure saccharin. After 5 weeks, metabolic parameters were characterized using the PhenoMaster metabolic cages system for 70 h. Light and dark phases are denoted by white and black rectangles on the x-axis, respectively, and grey bars for the dark phase. a, Liquids intake. b, AUC of a. c, Chow consumption. d, AUC of c. e, Respiratory exchange rate (RER). f, AUC of e. g, Physical activity as distance. h, AUC of g. i, Energy expenditure. The metabolic cages characterization was repeated twice.
Extended Data Figure 5 Glucose intolerant NAS-drinking mice display normal insulin levels and tolerance.
a, Fasting plasma insulin measured after 11 weeks of commercial NAS or controls (N = 10). b, Same as a, but measured after 5 weeks of HFD and pure saccharin or water (N = 20). c, Insulin tolerance test performed after 12 weeks of commercial NAS or controls (N = 10). Horizontal lines (a, b) or symbols (c) represent means; error bars, s.e.m. All measurements were performed on two independent cohorts.
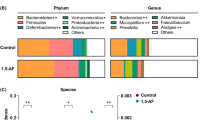
Extended Data Figure 6 Dysbiosis in saccharin-consuming mice and germ-free recipients.
Heat map representing W11 logarithmic-scale fold taxonomic differences between commercial saccharin and water or caloric sweetener consumers (N = 5 in each group). Right column, taxonomical differences in germ-free mice following faecal transplantation from commercial saccharin- (recipients N = 15) or glucose-consuming mice (N = 13). OTU number (GreenGenes) and the lowest taxonomic level identified are denoted.
Extended Data Figure 7 Functional analysis of saccharin-modulated microbiota.
a, b, Changes in bacterial relative abundance occur throughout the bacterial genome. Shown are changes in sequencing coverage along 10,000 bp genomic regions of Bacteroides vulgatus (a) and Akkermansia muciniphila (b), with bins ordered by abundance in week 0 of saccharin-treated mice. c, Fold change in relative abundance of modules belonging to phosphotransferase systems (PTS) between week 11 and week 0 in mice drinking commercial saccharin, glucose or water. Module diagram source: KEGG database. d, Enriched KEGG pathways (fold change > 1.38 as cutoff) in mice consuming HFD and pure saccharin versus water compared to the fold change in relative abundance of the same pathways in mice consuming commercial saccharin (week 11/week 0).
Extended Data Figure 8 Saccharin directly modulates the microbiota.
a, Experimental schematic. b, Relative taxonomic abundance of anaerobically cultured microbiota. c, AUC of germ-free mice 6 days following transplantation with saccharin-enriched or control faecal cultures (N = 10 and N = 9, respectively). Horizontal lines, means; error bars, s.e.m. **P < 0.01, unpaired two-sided Student t-test. The experiment was repeated twice.
Extended Data Figure 9 Impaired glycaemic control associated with acute saccharin consumption in humans is transferable to germ-free mice.
a, Experimental schematic (N = 7). b, c, Daily incremental AUC of days 1–4 versus days 5–7 in four responders (b) or three non-responders (c). d, Principal coordinates analysis (PCoA) of weighted UniFrac distances of 16S rRNA sequences demonstrating separation on principal coordinates 2 (PC2), 3 (PC3) and 4 (PC4) of microbiota from responders (N samples = 12) versus non-responders (N = 8) during days 5–7. e, Order-level relative abundance of taxa samples from days 1–7 of responders and non-responders. f, AUC in germ-free mice (N = 6) 6 days following faecal transplantation from samples of responder 1 (R1) collected before and after 7 days of saccharin consumption. g, h, OGTT and AUC in germ-free mice (N = 5) 6 days after receiving faecal samples collected from responder 4 (R4) before and after 7 days of saccharin consumption. i, AUC in germ-free mice (N = 5) 6 days following faecal transplantation from samples of non-responder 3 (NR3) collected before and after 7 days of saccharin consumption. j, k, OGTT and AUC in germ-free mice (N = 5) 6 days after receiving faecal samples collected from non-responder 2 (NR2) before and after 7 days of saccharin consumption. l, Fold taxonomical abundance changes of selected OTUs, altered in germ-free recipients of D7 versus D1 microbiomes from R1. Dot colour same as in e, bacterial orders. Symbols (GTT) or horizontal lines (AUC), means; error bars, s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001, two-way ANOVA and Bonferroni post-hoc analysis (GTT), unpaired two-sided Student t-test (AUC).
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-7. (XLSX 845 kb)
Rights and permissions
About this article
Cite this article
Suez, J., Korem, T., Zeevi, D. et al. Artificial sweeteners induce glucose intolerance by altering the gut microbiota. Nature 514, 181–186 (2014). https://doi.org/10.1038/nature13793
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13793
This article is cited by
-
Rebaudioside D decreases adiposity and hepatic lipid accumulation in a mouse model of obesity
Scientific Reports (2024)
-
Ultra-processed foods and food additives in gut health and disease
Nature Reviews Gastroenterology & Hepatology (2024)
-
Decoding the mystery of non-nutritive sweeteners
International Journal of Diabetes in Developing Countries (2024)
-
Triangulating nutrigenomics, metabolomics and microbiomics toward personalized nutrition and healthy living
Human Genomics (2023)
-
Association between diet soft drink consumption and metabolic dysfunction-associated steatotic liver disease: findings from the NHANES
BMC Public Health (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.