Abstract
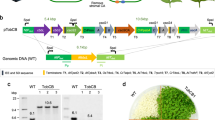
In photosynthetic organisms, d-ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) is the major enzyme assimilating atmospheric CO2 into the biosphere1. Owing to the wasteful oxygenase activity and slow turnover of Rubisco, the enzyme is among the most important targets for improving the photosynthetic efficiency of vascular plants2,3. It has been anticipated that introducing the CO2-concentrating mechanism (CCM) from cyanobacteria into plants could enhance crop yield4,5,6. However, the complex nature of Rubisco’s assembly has made manipulation of the enzyme extremely challenging, and attempts to replace it in plants with the enzymes from cyanobacteria and red algae have not been successful7,8. Here we report two transplastomic tobacco lines with functional Rubisco from the cyanobacterium Synechococcus elongatus PCC7942 (Se7942). We knocked out the native tobacco gene encoding the large subunit of Rubisco by inserting the large and small subunit genes of the Se7942 enzyme, in combination with either the corresponding Se7942 assembly chaperone, RbcX, or an internal carboxysomal protein, CcmM35, which incorporates three small subunit-like domains9,10. Se7942 Rubisco and CcmM35 formed macromolecular complexes within the chloroplast stroma, mirroring an early step in the biogenesis of cyanobacterial β-carboxysomes11,12. Both transformed lines were photosynthetically competent, supporting autotrophic growth, and their respective forms of Rubisco had higher rates of CO2 fixation per unit of enzyme than the tobacco control. These transplastomic tobacco lines represent an important step towards improved photosynthesis in plants and will be valuable hosts for future addition of the remaining components of the cyanobacterial CCM, such as inorganic carbon transporters and the β-carboxysome shell proteins4,5,6.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Andersson, I. & Backlund, A. Structure and function of Rubisco. Plant Physiol. Biochem. 46, 275–291 (2008)
Whitney, S. M., Houtz, R. L. & Alonso, H. Advancing our understanding and capacity to engineer nature’s CO2-sequestering enzyme, Rubisco. Plant Physiol. 155, 27–35 (2011)
Parry, M. A. J. et al. Rubisco activity and regulation as targets for crop improvement. J. Exp. Bot. 64, 717–730 (2013)
Zarzycki, J., Axen, S. D., Kinney, J. N. & Kerfeld, C. A. Cyanobacterial-based approaches to improving photosynthesis in plants. J. Exp. Bot. 64, 787–798 (2013)
McGrath, J. M. & Long, S. P. Can the cyanobacterial carbon-concentrating mechanism increase photosynthesis in crop species? A theoretical analysis. Plant Physiol. 164, 2247–2261 (2014)
Price, G. D. et al. The cyanobacterial CCM as a source of genes for improving photosynthetic CO2 fixation in crop species. J. Exp. Bot. 64, 753–768 (2013)
Whitney, S. M., Baldet, P., Hudson, G. S., Andrews, T. J. & Form, I. Rubiscos from non-green algae are expressed abundantly but not assembled in tobacco chloroplasts. Plant J. 26, 535–547 (2001)
Kanevski, I., Maliga, P., Rhoades, D. F. & Gutteridge, S. Plastome engineering of ribulose-1,5-bisphosphate carboxylase/oxygenase in tobacco to form a sunflower large subunit and tobacco small subunit hybrid. Plant Physiol. 119, 133–142 (1999)
Saschenbrecker, S. et al. Structure and function of RbcX, an assembly chaperone for hexadecameric rubisco. Cell 129, 1189–1200 (2007)
Long, B. M., Badger, M. R., Whitney, S. M. & Price, G. D. Analysis of carboxysomes from Synechococcus PCC7942 reveals multiple Rubisco complexes with carboxysomal proteins CcmM and CcaA. J. Biol. Chem. 282, 29323–29335 (2007)
Cameron, J. C., Wilson, S. C., Bernstein, S. L. & Kerfeld, C. A. Biogenesis of a bacterial organelle: the carboxysome assembly pathway. Cell 155, 1131–1140 (2013)
Chen, A. H., Robinson-Mosher, A., Savage, D. F., Silver, P. A. & Polka, J. K. The bacterial carbon-fixing organelle is formed by shell envelopment of preassembled cargo. PLoS ONE 8, e76127 (2013)
Parry, M. A. J., Andralojc, P. J., Mitchell, R. A. C., Madgwick, P. J. & Keys, A. J. Manipulation of Rubisco: the amount, activity, function and regulation. J. Exp. Bot. 54, 1321–1333 (2003)
Zhu, X. G., Long, S. P. & Ort, D. R. Improving photosynthetic efficiency for greater yield. Annu. Rev. Plant Biol. 61, 235–261 (2010)
Dhingra, A., Portis, A. R. & Daniell, H. Enhanced translation of a chloroplast-expressed RbcS gene restores small subunit levels and photosynthesis in nuclear RbcS antisense plants. Proc. Natl Acad. Sci. USA 101, 6315–6320 (2004)
von Caemmerer, S., Quick, W. P. & Furbank, R. T. The development of C4 rice: current progress and future challenges. Science 336, 1671–1672 (2012)
Whitney, S. M. & Andrews, T. J. Plastome-encoded bacterial ribulose-1,5-bisphosphate carboxylase/oxygenase (RubisCO) supports photosynthesis and growth in tobacco. Proc. Natl Acad. Sci. USA 98, 14738–14743 (2001)
Maliga, P. & Tungsuchat-Huang, T. Plastid transformation in Nicotiana tabacum and Nicotiana sylvestris by biolistic DNA delivery to leaves. Methods Mol. Biol. 1132, 147–163 (2014)
Zhou, F., Karcher, D. & Bock, R. Identification of a plastid intercistronic expression element (IEE) facilitating the expression of stable translatable monocistronic mRNAs from operons. Plant J. 52, 961–972 (2007)
Drechsel, O. & Bock, R. Selection of Shine–Dalgarno sequences in plastids. Nucleic Acids Res. 39, 1427–1438 (2011)
Long, B. M., Rae, B. D., Badger, M. R. & Price, G. D. Over-expression of the beta-carboxysomal CcmM protein in Synechococcus PCC7942 reveals a tight co-regulation of carboxysomal carbonic anhydrase (CcaA) and M58 content. Photosynth. Res. 109, 33–45 (2011)
Long, B. M., Tucker, L., Badger, M. R. & Price, G. D. Functional cyanobacterial β-carboxysomes have an absolute requirement for both long and short forms of the CcmM protein. Plant Physiol. 153, 285–293 (2010)
Yokota, A. & Canvin, D. T. Ribulose bisphosphate carboxylase/oxygenase content determined with [14C]carboxypentitol bisphosphate in plants and algae. Plant Physiol. 77, 735–739 (1985)
Mueller-Cajar, O. & Whitney, S. M. Evolving improved Synechococcus Rubisco functional expression in Escherichia coli . Biochem. J. 414, 205–214 (2008)
Parry, M. A. J., Keys, A. J., Madgwick, P. J., Carmo-Silva, A. E. & Andralojc, P. J. Rubisco regulation: a role for inhibitors. J. Exp. Bot. 59, 1569–1580 (2008)
Emlyn-Jones, D., Woodger, F. J., Price, G. D. & Whitney, S. M. RbcX can function as a Rubisco chaperonin, but is non-essential in Synechococcus PCC7942. Plant Cell Physiol. 47, 1630–1640 (2006)
Lin, M. T. et al. β-carboxysomal proteins assemble into highly organized structures in Nicotiana chloroplasts. Plant J. 79, 1–12 (2014)
Gray, B. N., Yang, H., Ahner, B. A. & Hanson, M. R. An efficient downstream box fusion allows high-level accumulation of active bacterial beta-glucosidase in tobacco chloroplasts. Plant Mol. Biol. 76, 345–355 (2011)
Bainbridge, G. et al. Engineering Rubisco to change its catalytic properties. J. Exp. Bot. 46, 1269–1276 (1995)
Wintermans, J. F. & de Mots, A. Spectrophotometric characteristics of chlorophylls a and b and their pheophytins in ethanol. Biochim. Biophys. Acta 109, 448–453 (1965)
Acknowledgements
We thank C. Kerfeld (Michigan State University) for helpful discussion and providing us with the Se7942 genomic DNA and purified His-tagged CcmM protein, W. Li (Cornell University) for technical assistance in generating, selecting and analysing the tobacco chloroplast transformants and M. Waqar Hameed (Cornell University) for the codon-optimized cyanobacterial Rubisco genes. This material is based upon work supported by the National Science Foundation under grant number EF-1105584 to M.R.H., Biotechnology and Biological Sciences Research Council under grant number BB/I024488/1 to M.A.J.P. and the National Institute of General Medical Sciences of the National Institutes of Health under award number F32GM103019 to M.T.L. P.J.A. and M.A.J.P. also acknowledge support from the 20:20 Wheat Institute Strategic Program (BBSRC BB/J/00426X/1).
Author information
Authors and Affiliations
Contributions
M.T.L. designed and generated the DNA constructs and the transgenic tobacco lines. A.O. carried out the TEM imaging, protein analyses and Rubisco activity assays. M.R.H., P.J.A. and M.A.J.P. supervised the project. All authors interpreted results and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Rubisco and CcmM35 content of SeLSM35 tobacco leaves.
The stated concentrations of purified Se Rubisco (a) and CcmM35 (b) proteins were used as standards. a, Immunoblot using an antibody against cyanobacterial LSU (top) and the standard curve used to estimate the amount of cyanobacterial Rubisco in samples S1–S3 extracted from SeLSM35 tobacco leaves (bottom). b, Immunoblot using an antibody against CcmM (top) and the standard curve used to estimate the amount of CcmM35 in samples S4–S6 extracted from SeLSM35 tobacco leaves (bottom). The band intensities in the two standard curves were obtained with ImageJ software and the standard curves with Microsoft Excel. c, The absolute and relative amounts (mean ± standard deviation) of CcmM35 and cyanobacterial Rubisco in SeLSM35 tobacco line from two separate measurements. Each Rubisco holoenzyme is assumed to be composed of 8 LSU and an unknown quantity of SSU.
Extended Data Figure 2 Electron micrographs of ultrathin sections of leaf mesophyll cells from the chloroplast transformant SeLSM35.
Large compartments containing cyanobacterial Rubisco and CcmM35 in the chloroplast stroma are indicated by black arrows. Leaf tissues were prepared by high pressure freeze fixation (HPF) in combination with immunogold labelling using an antibody against CcmM. A secondary antibody conjugated with 10-nm gold particles was used for the labelling. Scale bars, 500 nm.
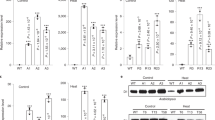
Extended Data Figure 3 Rubisco-specific 14CO2 fixation by crude leaf homogenates from tobacco lines expressing cyanobacterial Rubisco (SeLSX and SeLSM35) and wild-type tobacco (WT).
a, Carboxylase activity assayed with (+) and without (−) RuBP. b, Carboxylase activity assayed with (+) and without (−) the inhibitor CABP. The rates of carboxylase activity (mols fixed per mol act sites per s) are the means ± standard deviation derived from the 2, 4 and 10 min data obtained in assays at 125 µM CO2 (corresponding to 10 mM NaH14CO3, at pH 8.0).
Source data
Rights and permissions
About this article
Cite this article
Lin, M., Occhialini, A., Andralojc, P. et al. A faster Rubisco with potential to increase photosynthesis in crops. Nature 513, 547–550 (2014). https://doi.org/10.1038/nature13776
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13776
This article is cited by
-
RuBisCO activity assays: a simplified biochemical redox approach for in vitro quantification and an RNA sensor approach for in vivo monitoring
Microbial Cell Factories (2024)
-
Engineering α-carboxysomes into plant chloroplasts to support autotrophic photosynthesis
Nature Communications (2023)
-
Nanomaterials in agriculture for plant health and food safety: a comprehensive review on the current state of agro-nanoscience
3 Biotech (2023)
-
Plastid engineering using episomal DNA
Plant Cell Reports (2023)
-
Singular adaptations in the carbon assimilation mechanism of the polyextremophile cyanobacterium Chroococcidiopsis thermalis
Photosynthesis Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.