Abstract
Gain-of-function mutations in the fibroblast growth factor receptor 3 gene (FGFR3) result in skeletal dysplasias, such as thanatophoric dysplasia and achondroplasia (ACH). The lack of disease models using human cells has hampered the identification of a clinically effective treatment for these diseases. Here we show that statin treatment can rescue patient-specific induced pluripotent stem cell (iPSC) models and a mouse model of FGFR3 skeletal dysplasia. We converted fibroblasts from thanatophoric dysplasia type I (TD1) and ACH patients into iPSCs. The chondrogenic differentiation of TD1 iPSCs and ACH iPSCs resulted in the formation of degraded cartilage. We found that statins could correct the degraded cartilage in both chondrogenically differentiated TD1 and ACH iPSCs. Treatment of ACH model mice with statin led to a significant recovery of bone growth. These results suggest that statins could represent a medical treatment for infants and children with TD1 and ACH.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Rousseau, F. et al. Mutations in the gene encoding fibroblast growth factor receptor-3 in achondroplasia. Nature 371, 252–254 (1994)
Shiang, R. et al. Mutations in the transmembrane domain of FGFR3 cause the most common genetic form of dwarfism, achondroplasia. Cell 78, 335–342 (1994)
Warman, M. L. et al. Nosology and classification of genetic skeletal disorders: 2010 revision. Am. J. Med. Genet. A. 155, 943–968 (2011)
Deng, C., Wynshaw-Boris, A., Zhou, F., Kuo, A. & Leder, P. Fibroblast growth factor receptor 3 is a negative regulator of bone growth. Cell 84, 911–921 (1996)
Laederich, M. B. & Horton, W. A. Achondroplasia: pathogenesis and implications for future treatment. Curr. Opin. Pediatr. 22, 516–523 (2010)
Yasoda, A. et al. Systemic administration of C-type natriuretic peptide as a novel therapeutic strategy for skeletal dysplasias. Endocrinology 150, 3138–3144 (2009)
Lorget, F. et al. Evaluation of the therapeutic potential of a CNP analog in a Fgfr3 mouse model recapitulating achondroplasia. Am. J. Hum. Genet. 91, 1108–1114 (2012)
Xie, Y. et al. Intermittent PTH (1–34) injection rescues the retarded skeletal development and postnatal lethality of mice mimicking human achondroplasia and thanatophoric dysplasia. Hum. Mol. Genet. 21, 3941–3955 (2012)
Jin, M. et al. A novel FGFR3-binding peptide inhibits FGFR3 signaling and reverses the lethal phenotype of mice mimicking human thanatophoric dysplasia. Hum. Mol. Genet. 21, 5443–5455 (2012)
Garcia, S. et al. Postnatal soluble FGFR3 therapy rescues achondroplasia symptoms and restores bone growth in mice. Sci. Transl Med. 5, 203ra124 (2013)
Monsonego-Ornan, E., Adar, R., Feferman, T., Segev, O. & Yayon, A. The transmembrane mutation G380R in fibroblast growth factor receptor 3 uncouples ligand-mediated receptor activation from down-regulation. Mol. Cell. Biol. 20, 516–522 (2000)
Naski, M. C., Colvin, J. S., Coffin, J. D. & Ornitz, D. M. Repression of hedgehog signaling and BMP4 expression in growth plate cartilage by fibroblast growth factor receptor 3. Development 125, 4977–4988 (1998)
Okita, K. et al. A more efficient method to generate integration-free human iPS cells. Nature Methods 8, 409–412 (2011)
Oldershaw, R. A. et al. Directed differentiation of human embryonic stem cells toward chondrocytes. Nature Biotechnol. 28, 1187–1194 (2010)
Cho, J. Y. et al. Defective lysosomal targeting of activated fibroblast growth factor receptor 3 in achondroplasia. Proc. Natl Acad. Sci. USA 101, 609–614 (2004)
Harada, D. et al. Sustained phosphorylation of mutated FGFR3 is a crucial feature of genetic dwarfism and induces apoptosis in the ATDC5 chondrogenic cell line via PLCγ-activated STAT1. Bone 41, 273–281 (2007)
Guo, C. et al. Sprouty 2 disturbs FGFR3 degradation in thanatophoric dysplasia type II: a severe form of human achondroplasia. Cell. Signal. 20, 1471–1477 (2008)
Krejci, P. et al. Analysis of STAT1 activation by six FGFR3 mutants associated with skeletal dysplasia undermines dominant role of STAT1 in FGFR3 signaling in cartilage. PLoS ONE 3, e3961 (2008)
Legeai-Mallet, L., Benoist-Lasselin, C., Delezoide, A. L., Munnich, A. & Bonaventure, J. Fibroblast growth factor receptor 3 mutations promote apoptosis but do not alter chondrocyte proliferation in thanatophoric dysplasia. J. Biol. Chem. 273, 13007–13014 (1998)
Yamanaka, Y., Tanaka, H., Koike, M., Nishimura, R. & Seino, Y. PTHrP rescues ATDC5 cells from apoptosis induced by FGF receptor 3 mutation. J. Bone Miner. Res. 18, 1395–1403 (2003)
Yudoh, K. & Karasawa, R. Statin prevents chondrocyte aging and degeneration of articular cartilage in osteoarthritis (OA). Aging 2, 990–998 (2010)
Simopoulou, T., Malizos, K. N., Poultsides, L. & Tsezou, A. Protective effect of atorvastatin in cultured osteoarthritic chondrocytes. J. Orthop. Res. 28, 110–115 (2010)
Baker, J. F., Walsh, P. M., Byrne, D. P. & Mulhall, K. J. Pravastatin suppresses matrix metalloproteinase expression and activity in human articular chondrocytes stimulated by interleukin-1β. J. Orthopaed. Traumatol. 13, 119–123 (2012)
Mundy, G. et al. Stimulation of bone formation in vitro and in rodents by statins. Science 286, 1946–1949 (1999)
Millar, P. J. & Floras, J. S. Statins and the autonomic nervous system. Clinical Sci. 126, 401–415 (2014)
Olivieri, F. et al. Telomere/telomerase system: a new target of statins pleiotropic effect? Curr. Vasc. Pharmacol. 10, 216–224 (2012)
Zhang, J. et al. Statins, autophagy and cancer metastasis. Int. J. Biochem. Cell Biol. 45, 745–752 (2013)
Israel, M. A. et al. Probing sporadic and familial Alzheimer’s disease using induced pluripotent stem cells. Nature 482, 216–220 (2012)
Wolfe, S. M. Dangers of rosuvastatin identified before and after FDA approval. Lancet 363, 2189–2190 (2004)
Kawamoto, T. Use of a new adhesive film for the preparation of multi-purpose fresh-frozen sections from hard tissues, whole-animals, insects and plants. Arch. Histol. Cytol. 66, 123–143 (2003)
Takeo, T. et al. Birth of mice from vitrified/warmed 2-cell embryos transported at a cold temperature. Cryobiology 58, 196–202 (2009)
Faul, F., Erdfelder, E., Lang, A. G. & Buchner, A. G*Power 3: a flexible statistical power analysis program for the social, behavioral, and biomedical sciences. Behav. Res. Methods 39, 175–191 (2007)
Ikegami, D. et al. Identification of small molecular compounds and fabrication of its aqueous solution by laser-ablation, expanding primordial cartilage. Osteoarth. Cartil. 19, 233–241 (2011)
Gosset, M., Berenbaum, F., Thirion, S. & Jacques, C. Primary culture and phenotyping of murine chondrocytes. Nature Protocols 3, 1253–1260 (2008)
Hiramatsu, K. et al. Generation of hyaline cartilaginous tissue from mouse adult dermal fibroblast culture by defined factors. J. Clin. Invest. 121, 640–657 (2011)
Acknowledgements
We thank H. Ohashi, J. Murotsuki, T. Yamada and K. Ozono for preparation of the dermal fibroblasts from a patient and discussion. We thank D. Ornitz and A. Yasoda for the Fgfr3Ach mice. We thank K. Okita and S. Yamanaka for providing the hiPSC line 409B2, and A. Hotta for transducing the piggyBac vectors. We thank A. Motomura, X. Chen, Y. Minegishi, M. Nishino, T. Kobayashi, N. Oda, E. Ikeda and Y. Makita for their assistance. This study was supported in part by the Japan Science Technology Agency (JST), CREST (to N.T.) and Research Center Network for Realization of Regenerative Medicine (to N.T.), and Scientific Research Grant No. 24890101 (to A.Y.) and 26861716 (to A.Y.) and No. 24390354 (to N.T.) from MEXT.
Author information
Authors and Affiliations
Contributions
A.Y. was involved in most of the experiments. M.M. performed the immunohistological and immunoblot analyses, as well as the DNA construction. H.K., T.K. and Y.Y. performed the mouse experiments. M.O. contributed to the generation of iPSCs. H.S. and S.I. were involved in the study design. K.F. performed the immunoblot analyses. N.T. designed the study. A.Y. and N.T. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Characterization of the TD1 and ACH iPSCs.
a, Donor information and characterization of iPSCs. b, Left: a phase contrast image of TD1 iPSCs (TD1-714-3). Right: immunocytochemical staining of TD1 iPSCs (TD1-714-3) for SSEA4 and TRA1-60. The images are representative of two independent experiments. c, The histology of teratomas formed after implantation of TD1 iPSCs (TD1-10749-2) into SCID mice. The images are representative of experiments using three TD1 iPS clones established from three different patients. Scale bars, 50 μm.
Extended Data Figure 2 The results of a histological analysis of particles formed by chondrogenically differentiated wild-type and TD1 iPSCs on day 42.
The data were collected from three independent wild-type and three independent TD1 iPSC lines which were derived from three individuals, respectively. Top: three iPSC lines established from different control individuals all formed cartilaginous particles (top panels), whereas three iPSC lines established from different patients formed particles which lacked cartilaginous elements (bottom panels). Scale bars, 50 μm. Bottom: the area of the Safranin-O-positive region was divided by the total area of the particle. The error bars denote the means ± s.d. The number of particles examined is indicated at the bottom.
Extended Data Figure 3 Results of the expression analysis of chondrogenically differentiated wild-type iPSCs and TD1 iPSCs.
a, The results of a real-time RT–PCR expression analysis of marker genes in chondrogenically differentiated iPSCs lines on day 28. The data were collected from three independent wild-type and three independent TD1 iPSC lines which were derived from three individuals, respectively. Chondrocytes, re-differentiated human fetal chondrocytes; fibroblasts, dermal fibroblasts. *P < 0.05 and **P < 0.01 (n = 3 technical replicates) by the Tukey–Kramer post-hoc test. The error bars denote the means ± s.d. b–d, Immunohistochemical detection of the expression of type I collagen (b), type II collagen (c), SSEA4 (d, upper panels) and TRA1-60 (d, lower panels) in the particles formed by chondrogenically differentiated wild-type iPSCs (409B2) and TD1 iPSCs (TD1-714-3) on day 42. b, c, The images are representative of three independent experiments. d, The images are representative of experiments using three TD1-iPS clones established from three different patients. Scale bars, 50 μm (b, d); 25 μm (c).
Extended Data Figure 4 Expression levels of markers and apoptosis in chondrogenically differentiated wild-type and TD1 iPSCs.
a, Time course of the changes in the expression of markers in iPSCs subjected to chondrogenic differentiation, as determined by real-time RT–PCR (n = 3 technical replicates). b, The results of the TUNEL assay of particles formed by chondrogenically differentiated iPSCs (409B2 and TD1-714-3) on day 21. Scale bars, 50 μm. The images are representative of two independent experiments. c, The ratio of the numbers of TUNEL-positive cells per the total cell number during the chondrogenic differentiation of wild-type iPSCs (409B2) and TD1 iPSCs (TD1-714-3). **P < 0.01 for TD1-iPSC-derived cells compared to wild-type iPSC-derived cells on each day after chondrogenic differentiation (n = 3 particles), t-test. d, Immunohistochemical findings of the expression of cleaved-caspase 3 in the particles formed by chondrogenically differentiated wild-type iPSCs (409B2) and TD1 iPSCs (TD1-714-3) on day 28. Scale bars: 25 μm. The images are representative of two independent experiments. e, The results of a real-time RT–PCR expression analysis of p21 in chondrogenically differentiated TD1 iPSCs on days 0, 14 and 28 (n = 3 technical replicates). The data were collected from three independent wild-type and three independent TD1 iPSC lines which were respectively derived from three individuals. The error bars denote the means ± s.d.
Extended Data Figure 5 Rescuing the chondrogenically differentiated TD1 iPSCs (TD1-714-3) by FGFR3 knockdown and treatment with a FGFR3 neutralizing antibody.
a, Results of a real-time RT–PCR expression analysis of marker genes on day 28. iPSC, undifferentiated TD1 iPSCs; WT, chondrogenically differentiated wild-type iPSCs (409B2) on day 28. *P < 0.05; **P < 0.01, Tukey–Kramer post-hoc test (n = 3 technical replicates). The data were collected from three independent clones respectively bearing three different FGFR3 shRNAs. b, A FGFR3 neutralizing antibody was added to the medium during the chondrogenic differentiation of TD1 iPSCs to induce the formation of cartilaginous particles. IgG was added as a control. The results of a histological analysis of the particles. Left: histological sections of particles were stained with haematoxylin and eosin or Safranin O. Scale bars, 50 μm. Right: the area of the Safranin-O-positive region was divided by the total area of the particle. The number of particles examined is indicated at the bottom. c, The results of a real-time RT–PCR expression analysis of marker genes in chondrogenically differentiated TD1 iPSCs treated with the FGFR3 neutralizing antibody on day 28. *P < 0.05; **P < 0.01 by the t-test (n = 3 technical replicates). Error bars denote the means ± s.d.
Extended Data Figure 6 Screening for molecules that rescue chondrogenically differentiated TD1 iPSCs (TD1-714-3).
a, Candidate molecules or vehicle were added to the medium during the chondrogenic differentiation of TD1 iPSCs. The histology of the particles was examined on day 42. Left: histological sections of particles were stained with haematoxylin and eosin or Safranin O. The addition of a FGFR inhibitor and a G-protein antagonist, NF449, failed to recover the cartilage formation of chondrogenically differentiated TD1 iPSCs under the conditions examined. The FGFR inhibitor used in this experiment inhibits not only FGFR3 but also FGFR1 and FGFR2, which might have adversely affected the cartilage formation. The addition of CNP led to the partial recovery of cartilage formation. Scale bars, 50 μm. Right: the area of the Safranin O-positive region was divided by the total area of the particle. The number of particles examined is indicated at the bottom. The data are representative of two independent experiments. b, The results of a real-time RT–PCR expression analysis of marker genes in chondrogenically differentiated TD1 iPSCs treated with various factors on day 28. The addition of CNP increased the expression of chondrocyte marker genes and decreased the expression of fibroblast marker genes in the chondrogenically differentiated TD1 iPSCs. iPSC, undifferentiated TD1 iPSCs; WT, chondrogenically differentiated wild-type iPSCs (409B2) on day 28. The final concentrations of each molecule were: FGF inhibitor, 1 μM; NF449, 25 μM; and CNP, 100 nM (n = 3 technical replicates). Error bars denote the means ± s.d.
Extended Data Figure 7 Effects of statins on chondrogenically differentiated TD1 iPSCs.
a, The results of a real-time RT–PCR expression analysis of marker genes on day 28. TD1 (TD1-714-3) and wild-type (409B2) iPSCs were chondrogenically differentiated in the presence or absence of lovastatin (1 μM). TD1, chondrogenically differentiated TD1 iPSCs; iPSC, undifferentiated TD1 iPSCs; WT, chondrogenically differentiated wild-type iPSCs (n = 3 technical replicates). The data are representative of two independent experiments. b, Rescue of chondrogenically differentiated TD1 iPSCs by statin treatment. Mevastatin, atorvastatin, pravastatin, rosuvastatin or fluvastatin (each 1 μM) was added to the medium during the chondrogenic differentiation of TD1 iPSCs. On day 42, the particles were subjected to a histological analysis. Top: histological sections of particles were stained with haematoxylin and eosin or Safranin O. Scale bars, 50 μm. Bottom: the area of the Safranin-O-positive region was divided by the total area of the particle. The number of particles examined is indicated at the bottom. The data are representative of three independent experiments. The error bars denote the means ± s.d. *P < 0.05; **P < 0.01 by the Tukey–Kramer post-hoc test.
Extended Data Figure 8 Rescue of chondrogenically differentiated ACH iPSCs by statin treatment.
HDFs were obtained from two ACH patients (ACH-8857 and ACH-8858) bearing a heterozygous G380R mutation in the FGFR3 gene (Extended Data Fig. 1a). HDFs were also obtained from an individual (ACHhomo-8859) whose parents both had ACH, who showed a more severe phenotype of chondrodysplasia than typical ACH, and who was homozygous for the G380R mutation in the FGFR3 gene. We generated more than three iPSC lines for each patient and analysed one iPSC line (ACH-8857-1, ACH-8858-6 and ACHhomo-8859-3) derived from each patient’s HDFs. We confirmed that all iPSC lines expressed SSEA4 and TRA1-60, and formed teratomas containing all three germ layers in mice. Lovastatin and CNP were added to the medium during the chondrogenic differentiation of ACH iPSCs. The histology of particles was examined on day 42. a, Histological sections of particles were stained with haematoxylin and eosin or Safranin O. Scale bars, 50 μm. b, The area of the Safranin-O-positive region was divided by the total area of the particle. The number of particles examined is indicated at the bottom. The error bars denote the means ± s.d.
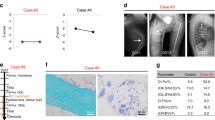
Extended Data Figure 9 X-ray images of Fgfr3Ach and wild-type mice treated with rosuvastatin or vehicle.
The images of the mice with lowest and highest weights in each group are shown. The weight (g) of each mouse is indicated at the bottom of each panel. Scale bars, 10 mm.
Extended Data Figure 10 Organ culture of metatarsal primordial cartilage (a, b), pellet culture of primary chondrocytes (c, d) and culture of primary chondrocytes (e) from Fgfr3Ach mice in the presence or absence of lovastatin.
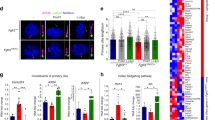
a, Left: images of cartilage on day 7. Right: mean lengths of the cartilage on day 1 and day 7 (n = 8 cartilage samples). b, Cartilage on day 7 was treated with BrdU. Left: histological sections were stained with Safranin O and immunostained with BrdU. Right; number of BrdU-positive cells were divided by the total number of cells (n = 7 cartilage samples). c, Histological sections of pellets cultured for 14 days. The images are representative of three pellets. d, The results of the real-time RT–PCR expression analysis of pellets cultured for 14 days (Sox9, Col2a1 and Acan) and 28 days (Runx2 and Col10a1) (n = 3 technical replicates). The data are representative of two independent experiments. e, Primary chondrocytes from wild-type and Fgfr3Ach mice were cultured in the presence or absence of lovastatin, MG132 or bafilomycin A1, and were subjected to an immunoblot analysis using an anti-FGFR3 antibody. The images are representative of two independent experiments. Error bars denote the means ± s.d. *P < 0.5; **P < 0.01 by the t-test. Scale bars, 50 μm.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary References and Supplementary Tables 1-3. (PDF 241 kb)
Rights and permissions
About this article
Cite this article
Yamashita, A., Morioka, M., Kishi, H. et al. Statin treatment rescues FGFR3 skeletal dysplasia phenotypes. Nature 513, 507–511 (2014). https://doi.org/10.1038/nature13775
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13775
This article is cited by
-
Regeneration of joint surface defects by transplantation of allogeneic cartilage: application of iPS cell-derived cartilage and immunogenicity
Inflammation and Regeneration (2023)
-
Fluvastatin promotes chondrogenic differentiation of adipose-derived mesenchymal stem cells by inducing bone morphogenetic protein 2
BMC Pharmacology and Toxicology (2022)
-
I nuovi farmaci per la terapia dell’acondroplasia
L'Endocrinologo (2022)
-
Identification of a familial cleidocranial dysplasia with a novel RUNX2 mutation and establishment of patient-derived induced pluripotent stem cells
Odontology (2022)
-
Targeted gene correction and functional recovery in achondroplasia patient-derived iPSCs
Stem Cell Research & Therapy (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.