Abstract
The ribosome is a molecular machine responsible for protein synthesis and a major target for small-molecule inhibitors. Compared to the wealth of structural information available on ribosome-targeting antibiotics in bacteria, our understanding of the binding mode of ribosome inhibitors in eukaryotes is currently limited. Here we used X-ray crystallography to determine 16 high-resolution structures of 80S ribosomes from Saccharomyces cerevisiae in complexes with 12 eukaryote-specific and 4 broad-spectrum inhibitors. All inhibitors were found associated with messenger RNA and transfer RNA binding sites. In combination with kinetic experiments, the structures suggest a model for the action of cycloheximide and lactimidomycin, which explains why lactimidomycin, the larger compound, specifically targets the first elongation cycle. The study defines common principles of targeting and resistance, provides insights into translation inhibitor mode of action and reveals the structural determinants responsible for species selectivity which could guide future drug development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Atomic coordinates and structure factors for the reported crystal structures have been deposited in the Protein Data Bank (http://www.pdb.org/pdb/home/home.do) under accession codes 4U3M (anisomycin), 4U56 (blasticidin S), 4U3N (CCA), 4U55 (cryptopleurine), 4U3U (cycloheximide), 4U53 (deoxynivalenol), 4U4N (edeine), 4U4O (geneticin G418), 4U4Q (homoharringtonine), 4U4R (lactimidomycin), 4U4U (lycorine), 4U52 (nagilactone C), 4U51 (narcilasine), 4U4Y (pactamycin), 4U4Z (phyllanthoside), 4U6F (T-2 toxin) and 4U50 (verrucarin).
References
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334, 1524–1529 (2011)
Melnikov, S. et al. One core, two shells: bacterial and eukaryotic ribosomes. Nature Struct. Mol. Biol. 19, 560–567 (2012)
Jenner, L. et al. Crystal structure of the 80S yeast ribosome. Curr. Opin. Struct. Biol. 22, 759–767 (2012)
Sonenberg, N. & Hinnebusch, A. G. Regulation of translation initiation in eukaryotes: mechanisms and biological targets. Cell 136, 731–745 (2009)
Wilson, D. N. Ribosome-targeting antibiotics and mechanisms of bacterial resistance. Nature Rev. Microbiol. 12, 35–48 (2014)
Blaha, G. M., Polikanov, Y. S. & Steitz, T. A. Elements of ribosomal drug resistance and specificity. Curr. Opin. Struct. Biol. 22, 750–758 (2012)
Zhou, J. et al. Design at the atomic level: generation of novel hybrid biaryloxazolidinones as promising new antibiotics. Bioorg. Med. Chem. Lett. 18, 6179–6183 (2008)
Hobbie, S. N. et al. Genetic reconstruction of protozoan rRNA decoding sites provides a rationale for paromomycin activity against Leishmania and Trypanosoma . PLoS Negl. Trop. Dis. 5, e1161 (2011)
Lu, W., Roongsawang, N. & Mahmud, T. Biosynthetic studies and genetic engineering of pactamycin analogs with improved selectivity toward malarial parasites. Chem. Biol. 18, 425–431 (2011)
Santagata, S. et al. Tight coordination of protein translation and HSF1 activation supports the anabolic malignant state. Science 341, 1238303 (2013)
Bidou, L., Allamand, V., Rousset, J. P. & Namy, O. Sense from nonsense: therapies for premature stop codon diseases. Trends Mol. Med. 18, 679–688 (2012)
Darnell, J. C. & Klann, E. The translation of translational control by FMRP: therapeutic targets for FXS. Nature Neurosci. 16, 1530–1536 (2013)
Gürel, G., Blaha, G., Steitz, T. A. & Moore, P. B. Structures of triacetyloleandomycin and mycalamide A bind to the large ribosomal subunit of Haloarcula marismortui . Antimicrob. Agents Chemother. 53, 5010–5014 (2009)
Gürel, G., Blaha, G., Moore, P. B. & Steitz, T. A. U2504 determines the species specificity of the A-site cleft antibiotics: the structures of tiamulin, homoharringtonine, and bruceantin bound to the ribosome. J. Mol. Biol. 389, 146–156 (2009)
Klinge, S., Voigts-Hoffmann, F., Leibundgut, M., Arpagaus, S. & Ban, N. Crystal structure of the eukaryotic 60S ribosomal subunit in complex with initiation factor 6. Science 334, 941–948 (2011)
Decatur, W. A. & Fournier, M. J. rRNA modifications and ribosome function. Trends Biochem. Sci. 27, 344–351 (2002)
Chan, J., Khan, S. N., Harvey, I., Merrick, W. & Pelletier, J. Eukaryotic protein synthesis inhibitors identified by comparison of cytotoxicity profiles. RNA 10, 528–543 (2004)
Schneider-Poetsch, T. et al. Inhibition of eukaryotic translation elongation by cycloheximide and lactimidomycin. Nature Chem. Biol. 6, 209–217 (2010)
Lee, S., Liu, B., Huang, S. X., Shen, B. & Qian, S. B. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc. Natl Acad. Sci. USA 109, E2424–E2432 (2012)
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009)
Wintermeyer, W. & Zachau, H. G. Fluorescent derivatives of yeast tRNAPhe . Eur. J. Biochem. 98, 465–475 (1979)
Lill, R., Robertson, J. M. & Wintermeyer, W. Affinities of tRNA binding sites of ribosomes from Escherichia coli . Biochemistry 25, 3245–3255 (1986)
Svidritskiy, E., Ling, C., Ermolenko, D. N. & Korostelev, A. A. Blasticidin S inhibits translation by trapping deformed tRNA on the ribosome. Proc. Natl Acad. Sci. USA 110, 12283–12288 (2013)
Cannone, J. J. et al. The comparative RNA web (CRW) site: an online database of comparative sequence and structure information for ribosomal, intron, and other RNAs. BMC Bioinformatics 3, 2 (2002)
Demeshkina, N., Jenner, L., Westhof, E., Yusupov, M. & Yusupova, G. A new understanding of the decoding principle on the ribosome. Nature 484, 256–259 (2012)
Ogle, J. M., Murphy, F. V., Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002)
Shulman, E. et al. Designer aminoglycosides that selectively inhibit cytoplasmic rather than mitochondrial ribosomes show decreased ototoxicity: a strategy for the treatment of genetic diseases. J. Biol. Chem. 289, 2318–2330 (2014)
Fan-Minogue, H. & Bedwell, D. M. Eukaryotic ribosomal RNA determinants of aminoglycoside resistance and their role in translational fidelity. RNA 14, 148–157 (2008)
Recht, M. I., Douthwaite, S. & Puglisi, J. D. Basis for prokaryotic specificity of action of aminoglycoside antibiotics. EMBO J. 18, 3133–3138 (1999)
Becker, B. & Cooper, M. A. Aminoglycoside antibiotics in the 21st century. ACS Chem. Biol. 8, 105–115 (2013)
Perez-Fernandez, D. et al. 4′-O-substitutions determine selectivity of aminoglycoside antibiotics. Nature Commun. 5, 3112 (2014)
Dinos, G. et al. Dissecting the ribosomal inhibition mechanisms of edeine and pactamycin: the universally conserved residues G693 and C795 regulate P-site RNA binding. Mol. Cell 13, 113–124 (2004)
Pioletti, M. et al. Crystal structures of complexes of the small ribosomal subunit with tetracycline, edeine and IF3. EMBO J. 20, 1829–1839 (2001)
Kozak, M. & Shatkin, A. J. Migration of 40 S ribosomal subunits on messenger RNA in the presence of edeine. J. Biol. Chem. 253, 6568–6577 (1978)
Dölz, H., Vazquez, D. & Jimenez, A. Quantitation of the specific interaction of [14a-3H]cryptopleurine with 80S and 40S ribosomal species from the yeast Saccharomyces cerevisiae . Biochemistry 21, 3181–3187 (1982)
Ingolia, N. T., Lareau, L. F. & Weissman, J. S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011)
Chen, Y. & Li, S. Omacetaxine mepesuccinate in the treatment of intractable chronic myeloid leukemia. Onco Targets Ther 7, 177–186 (2014)
Evidente, A. et al. Biological evaluation of structurally diverse amaryllidaceae alkaloids and their synthetic derivatives: discovery of novel leads for anticancer drug design. Planta Med. 75, 501–507 (2009)
McCormick, S. P., Stanley, A. M., Stover, N. A. & Alexander, N. J. Trichothecenes: from simple to complex mycotoxins. Toxins 3, 802–814 (2011)
Shirai, A., Sadaie, M., Shinmyozu, K. & Nakayama, J. Methylation of ribosomal protein L42 regulates ribosomal function and stress-adapted cell growth. J. Biol. Chem. 285, 22448–22460 (2010)
Kawai, S. et al. Drastic alteration of cycloheximide sensitivity by substitution of one amino acid in the L41 ribosomal protein of yeasts. J. Bacteriol. 174, 254–262 (1992)
Kaàufer, N. F., Fried, H. M., Schwindinger, W. F., Jasin, M. & Warner, J. R. Cycloheximide resistance in yeast: the gene and its protein. Nucleic Acids Res. 11, 3123–3135 (1983)
Sánchez, L., Vásquez, D. & Jiménez, A. Genetics and biochemistry of cryptopleurine resistance in the yeast Saccharomyces cerevisiae . Mol. Gen. Genet. 156, 319–326 (1977)
Mueller, M., Wang, M. & Schulze-Briese, C. Optimal fine phi-slicing for single-photon-counting pixel detectors. Acta Crystallogr. D 68, 42–56 (2012)
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Bolton, E., Wang, Y., Thiessen, P. A. & Bryant, S. H. PubChem: integrated platform of small molecules and biological activities. Annu. Rep. Comput. Chem. 4, 217–241 (2008)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Bruno, I. J. et al. Retrieval of crystallographically-derived molecular geometry information. J. Chem. Inf. Comput. Sci. 44, 2133–2144 (2004)
Ban, N. et al. A new system for naming ribosomal proteins. Curr. Opin. Struct. Biol. 24, 165–169 (2014)
Schneider-Poetsch, T. et al. Inhibition of eukaryotic translation elongation by cycloheximide and lactimidomycin. Nature Chem. Biol. 6, 209–217 (2010)
Cundliffe, E., Cannon, M. & Davies, J. Mechanism of inhibition of eukaryotic protein synthesis by trichothecene fungal toxins. Proc. Natl Acad. Sci. USA 71, 30–34 (1974)
Middlebrook, J. L. & Leatherman, D. L. Binding of T-2 toxin to eukaryotic cell ribosomes. Biochem. Pharmacol. 38, 3103–3110 (1989)
Ehrlich, K. C. & Daigle, K. W. Protein synthesis inhibition by 8-oxo-12,13-epoxytrichothecenes. Biochim. Biophys. Acta 923, 206–213 (1987)
Fresno, M., Jiménez, A. & Vázquez, D. Inhibition of translation in eukaryotic systems by harringtonine. Eur. J. Biochem. 72, 323–330 (1977)
Gürel, G., Blaha, G., Moore, P. B. & Steitz, T. A. U2504 determines the species specificity of the A-site cleft antibiotics: the structures of tiamulin, homoharringtonine, and bruceantin bound to the ribosome. J. Mol. Biol. 389, 146–156 (2009)
Tujebajeva, R. M., Graifer, D. M., Karpova, G. G. & Ajtkhozhina, N. A. Alkaloid homoharringtonine inhibits polypeptide chain elongation on human ribosomes on the step of peptide bond formation. FEBS Lett. 257, 254–256 (1989)
Carrasco, L., Fresno, M. & Vazquez, D. Narciclasine: an antitumour alkaloid which blocks peptide bond formation by eukaryotic ribosomes. FEBS Lett. 52, 236–239 (1975)
Baez, A. & Vazquez, D. Binding of [3H]narciclasine to eukaryotic ribosomes. A study on a structure-activity relationship. Biochim. Biophys. Acta 518, 95–103 (1978)
Jimenez, A., Santos, A., Alonso, G. & Vazquez, D. Inhibitors of protein synthesis in eukarytic cells. Comparative effects of some amaryllidaceae alkaloids. Biochim. Biophys. Acta 425, 342–348 (1976)
Kukhanova, M., Victorova, L. & Krayevsky, A. Peptidyltransferase center of ribosomes. On the mechanism of action of alkaloid lycorine. FEBS Lett. 160, 129–133 (1983)
Grollman, A. P. Inhibitors of protein biosynthesis. II. Mode of action of anisomycin. J. Biol. Chem. 242, 3226–3233 (1967)
Hansen, J. L., Moore, P. B. & Steitz, T. A. Structures of five antibiotics bound at the peptidyl transferase center of the large ribosomal subunit. J. Mol. Biol. 330, 1061–1075 (2003)
Yamaguchi, H., Yamamoto, C. & Tanaka, N. Inhibition of protein synthesis by blasticidin S. I. Studies with cell-free systems from bacterial and mammalian cells. J. Biochem. 57, 667–677 (1965)
Svidritskiy, E., Ling, C., Ermolenko, D. N. & Korostelev, A. A. Blasticidin S inhibits translation by trapping deformed tRNA on the ribosome. Proc. Natl Acad. Sci. USA 110, 12283–12288 (2013)
Eustice, D. C. & Wilhelm, J. M. Mechanisms of action of aminoglycoside antibiotics in eucaryotic protein synthesis. Antimicrob. Agents Chemother. 26, 53–60 (1984)
Manuvakhova, M., Keeling, K. & Bedwell, D. M. Aminoglycoside antibiotics mediate context-dependent suppression of termination codons in a mammalian translation system. RNA 6, 1044–1055 (2000)
Contreras, A., Vazquez, D. & Carrasco, L. Inhibition, by selected antibiotics, of protein synthesis in cells growing in tissue cultures. J. Antibiot. 31, 598–602 (1978)
Brodersen, D. E. et al. The structural basis for the action of the antibiotics tetracycline, pactamycin, and hygromycin B on the 30S ribosomal subunit. Cell 103, 1143–1154 (2000)
Dinos, G. et al. Dissecting the ribosomal inhibition mechanisms of edeine and pactamycin: the universally conserved residues G693 and C795 regulate P-site RNA binding. Mol. Cell 13, 113–124 (2004)
Bucher, K. & Skogerson, L. Cryptopleurine–an inhibitor of translocation. Biochemistry 15, 4755–4759 (1976)
Kozak, M. & Shatkin, A. J. Migration of 40 S ribosomal subunits on messenger RNA in the presence of edeine. J. Biol. Chem. 253, 6568–6577 (1978)
Pioletti, M. et al. Crystal structures of complexes of the small ribosomal subunit with tetracycline, edeine and IF3. EMBO J. 20, 1829–1839 (2001)
Dehoux, P., Davies, J. & Cannon, M. Natural cycloheximide resistance in yeast. The role of ribosomal protein L41. Eur. J. Biochem. 213, 841–848 (1993)
Kawai, S. et al. Drastic alteration of cycloheximide sensitivity by substitution of one amino acid in the L41 ribosomal protein of yeasts. J. Bacteriol. 174, 254–262 (1992)
Schultz, L. D. & Friesen, J. D. Nucleotide sequence of the tcml gene (ribosomal protein L3) of Saccharomyces cerevisiae . J. Bacteriol. 155, 8–14 (1983)
Koshinsky, H. A., Schappert, K. T. & Khachatourians, G. G. Isolation and characterization of Saccharomyces cerevisiae mutants resistant to T-2 toxin. Curr. Genet. 13, 363–368 (1988)
Adam, G. et al. Molecular mechanisms of deoxynivalenol resistance in the yeast Saccharomyces cerevisiae . Mycotoxin Res. 17 (Suppl. 1). 19–23 (2001)
Jimenez, A., Sanchez, L. & Vazquez, D. Simultaneous ribosomal resistance to trichodermin and anisomycin in Saccharomyces cerevisiae mutants. Biochim. Biophys. Acta 383, 427–434 (1975)
Hobden, A. N. & Cundliffe, E. Ribosomal resistance to the 12,13-epoxytrichothecene antibiotics in the producing organism Myrothecium verrucaria . Biochem. J. 190, 765–770 (1980)
Jimenez, A., Sanchez, L. & Vazquez, D. Location of resistance to the alkaloid narciclasine in the 60S ribosomal subunit. FEBS Lett. 55, 53–56 (1975)
Nelson, J. A. E., Savereide, P. B. & Lefebvre, P. A. The CRY1 gene in Chlamydomonas reinhardtii: structure and use as a dominant selectable marker for nuclear transformation. Mol. Cell. Biol. 14, 4011–4019 (1994)
Paulovich, A. G., Thompson, J. R., Larkin, J. C., Li, Z. & Woolford, J. L., Jr Molecular genetics of cryptopleurine resistance in Saccharomyces cerevisiae: expression of a ribosomal protein gene family. Genetics 135, 719–730 (1993)
Acknowledgements
We thank J. Liu (Johns Hopkins Medical Institute), D. Wilson (Gene Center Munich), P. Hazendonk (Agriculture and Agri-Food Canada) and the NIH/NCI Developmental Therapeutics Program for providing materials. We acknowledge SOLEIL synchrotron (France), all staff members of PROXIMA1 beamline, especially A. Thompson and P. Legrand for their assistance during data collection. We thank A. Perez Lara, MPI Göttingen, for the help with the ITC experiments and S. Melnikov, IGBMC, for reading the manuscript. I.P. acknowledges support from AFM-Telethon post-doctoral fellowship. This work was supported by the SATT Conectus Technology Maturation grant I12-042 (to N.G.D.L.), the ERC Advanced grant 294312, the Human Frontier Science Program grant RGP0062/2012 and the Russian Government Program of Competitive Growth of Kazan Federal University (to M.Y.), the French National Research Agency grant ANR-11-BSV8-006 01 (to G.Y.) and the Deutsche Forschungsgemeinschaft grant (to M.V.R.).
Author information
Authors and Affiliations
Contributions
M.Y. supervised the study. N.G.D.L. designed the experiments. N.G.D.L. and I.P. conducted purification, crystallization and post-crystallization treatment experiments, collected X-ray diffraction data and carried out the structure determination. N.G.D.L., I.P., G.Y. and M.Y. analysed the crystal structures. M.V.R. and W.H. designed, performed and interpreted rapid kinetic experiments. N.G.D.L. wrote the initial manuscript to which M.V.R., G.Y. and M.Y. contributed specialist insights. All authors helped with refining the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
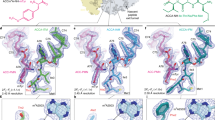
Extended Data Figure 1 Unbiased positive electron density of small-molecule inhibitors and CCA-trinucleotide.
Fo–Fc positive electron density maps of the 16 small-molecule inhibitors and the CCA tri-nucleotide. The maps were contoured at 3.0–3.5σ.
Extended Data Figure 2 Electron density of small-molecule inhibitors and CCA-trinucleotide.
2Fo–Fc electron density maps of the 16 small-molecule inhibitors and the CCA tri-nucleotide. The maps were contoured at 1.0–1.5σ.
Extended Data Figure 3 Structures of homoharringtonine, anisomycin, blasticidin S and pactamycin in eukaryotes, archaea and bacteria.
Complexes with bacterial and archaeal structures were aligned with the 25S rRNA or the 18S rRNA of the yeast ribosome. Differences in the binding pocket were found only in the case of anisomycin as described in Extended Data Fig. 8. Coordinates were taken from the PDB databank; PDB entries are indicated in parentheses.
Extended Data Figure 4 Structural differences in protein eL42 may preclude the binding of lactimidomycin and phyllanthoside to the archaeal ribosome.
In archaea, the protein eL42 (in red, PDB 1JJ2) is shorter than its eukaryotic counterpart (yellow) and adopts a markedly different conformation that clashes with lactimidomycin (pink) and phyllanthoside (cyan). Residues of protein eL42 from archaea involved in the steric clash with both inhibitors are depicted in red with sticks and van der Waals spheres. Although the 60S tRNA E-site is targeted by small-molecule inhibitors in archaea and eukaryotes, remarkably no antibiotics targeting this site in bacteria have been described.
Extended Data Figure 5 Close-up view of CCA tri-nucleotide binding site.
CCA tri-nucleotide (white) bound to the 60S tRNA E-site. The binding pocket is formed by 25S rRNA nucleotides (blue) and part of protein eL42 (yellow). In eukaryotes, the protein eL42 remodels the 60S E-site and participates actively in positioning the CCA-end. Although C75 is stabilized by stacking and hydrogen bonds interactions with eL42, the terminal residue A76 of deacylated tRNA enters the pocket and forms a non-canonical base pair with a conserved residue of the 25S rRNA.
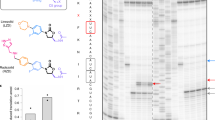
Extended Data Figure 6 Kinetic study of lactimidomycin and cycloheximide.
a, Deacylated tRNA binding to the bacterial 70S and eukaryotic 80S ribosomes. Time courses of tRNAPhe (Prf) binding to the S. cerevisiae 80S (blue) and E. coli 70S (red) ribosomes measured by the stopped-flow technique. b, Competition binding assays. Dose response curves for lactimidomycin (closed circles) and cycloheximide (open circles). Inset, the binding of the tRNA to the 70S ribosome was not affected in the presence of lactimidomycin (blue) and cycloheximide (magenta). Control without inhibitors is shown in black. c, Measurement of cycloheximide affinity to the 80S ribosome by isothermal titration calorimetry. The curves present the thermodynamic parameters of cycloheximide binding to 80S ribosomes (black circles) and control buffer (red circles). N, number of binding sites. The affinity was determined in 4 independent experiments.
Extended Data Figure 7 Close-up view of blasticidin S binding site.
Blasticidin S (pink) bound to the 60S tRNA P-site. The binding pocket is formed exclusively by nucleotides of the 25S rRNA (yellow). Dashed lines indicate hydrogen contacts with G2619 that precludes the formation of the base pair with C75 of the tRNA in the P-site.
Extended Data Figure 8 Conformational changes in the peptidyl transferase centre and differences with the archaeal ribosome.
a, A-site inhibitors induce conformational changes upon binding to the peptidyl transferase centre of the yeast ribosome. Superimposition of the vacant 80S ribosome (PDB 3U5A–3U5D, blue) and the 80S ribosome in complexes with A-site inhibitors (25S rRNA in yellow). The structure of anisomycin (orange) was chosen as a reference to represent the peptidyl transferase centre A-site inhibitors. Residue U2875 (U2506) undergoes the most drastic change resulting in the breakdown of a canonical base pair formed by G2952 (U2583) and its subsequent flipping out. The reorientation of U2875 (U2506) participates in preventing the binding of aminoacyl-tRNA. b, U2875 adopts a different conformation upon binding of anisomycin (orange) to the peptidyl transferase centre A-site (yellow) in eukaryotes in contrast to its homologue (U2541) in archaea (magenta). Superimposition of the 50S large subunit from H. marismortui in complex with anisomycin (PDB 1K73) and the 80S ribosome in complex with anisomycin.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1. (PDF 144 kb)
Rights and permissions
About this article
Cite this article
Garreau de Loubresse, N., Prokhorova, I., Holtkamp, W. et al. Structural basis for the inhibition of the eukaryotic ribosome. Nature 513, 517–522 (2014). https://doi.org/10.1038/nature13737
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13737
This article is cited by
-
A Legionella toxin exhibits tRNA mimicry and glycosyl transferase activity to target the translation machinery and trigger a ribotoxic stress response
Nature Cell Biology (2023)
-
USP36 stabilizes nucleolar Snail1 to promote ribosome biogenesis and cancer cell survival upon ribotoxic stress
Nature Communications (2023)
-
Cis-regulatory elements and transcription factors related to auxin signaling in the streptophyte algae Klebsormidium nitens
Scientific Reports (2023)
-
Influence of deoxynivalenol-contaminated feed on the immune response of pigs after PRRSV vaccination and infection
Archives of Toxicology (2023)
-
Prognostic value of amino acid metabolism-related gene expression in invasive breast carcinoma
Journal of Cancer Research and Clinical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.