Abstract
Clustered regularly interspaced short palindromic repeats (CRISPR) together with CRISPR-associated (Cas) proteins form the CRISPR/Cas system to defend against foreign nucleic acids of bacterial and archaeal origin1,2,3,4,5,6,7,8,9. In the I–E subtype CRISPR/Cas system, eleven subunits from five Cas proteins (CasA1B2C6D1E1) assemble along a CRISPR RNA (crRNA) to form the Cascade complex10,11,12,13. Here we report on the 3.05 Å crystal structure of the 405-kilodalton Escherichia coli Cascade complex that provides molecular details beyond those available from earlier lower-resolution cryo-electron microscopy structures. The bound 61-nucleotide crRNA spans the entire 11-protein subunit-containing complex, where it interacts with all six CasC subunits (named CasC1–6), with its 5′ and 3′ terminal repeats anchored by CasD and CasE, respectively. The crRNA spacer region is positioned along a continuous groove on the concave surface generated by the aligned CasC1–6 subunits. The five long β-hairpins that project from individual CasC2–6 subunits extend across the crRNA, with each β-hairpin inserting into the gap between the last stacked base and its adjacent splayed counterpart, and positioned within the groove of the preceding CasC subunit. Therefore, instead of continuously stacking, the crRNA spacer region is divided into five equal fragments, with each fragment containing five stacked bases flanked by one flipped-out base. Each of those crRNA spacer fragments interacts with CasC in a similar fashion. Furthermore, our structure explains why the seed sequence, with its outward-directed bases, has a critical role in target DNA recognition. In conclusion, our structure of the Cascade complex provides novel molecular details of protein–protein and protein–RNA alignments and interactions required for generation of a complex mediating RNA-guided immune surveillance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wiedenheft, B., Sternberg, S. H. & Doudna, J. A. RNA-guided genetic silencing systems in bacteria and archaea. Nature 482, 331–338 (2012)
Barrangou, R. et al. CRISPR provides acquired resistance against viruses in prokaryotes. Science 315, 1709–1712 (2007)
Garneau, J. E. et al. The CRISPR/Cas bacterial immune system cleaves bacteriophage and plasmid DNA. Nature 468, 67–71 (2010)
Andersson, A. F. & Banfield, J. F. Virus population dynamics and acquired virus resistance in natural microbial communities. Science 320, 1047–1050 (2008)
Jansen, R., Embden, J. D., Gaastra, W. & Schouls, L. M. Identification of genes that are associated with DNA repeats in prokaryotes. Mol. Microbiol. 43, 1565–1575 (2002)
van der Oost, J., Jore, M. M., Westra, E. R., Lundgren, M. & Brouns, S. J. CRISPR-based adaptive and heritable immunity in prokaryotes. Trends Biochem. Sci. 34, 401–407 (2009)
Westra, E. R. et al. The CRISPRs, they are a-changin': how prokaryotes generate adaptive immunity. Annu. Rev. Genet. 46, 311–339 (2012)
van der Oost, J., Westra, E. R., Jackson, R. N. & Wiedenheft, B. Unravelling the structural and mechanistic basis of CRISPR-Cas systems. Nature Rev. Microbiol. 12, 479–492 (2014)
Makarova, K. S., Wolf, Y. I. & Koonin, E. V. The basic building blocks and evolution of CRISPR-cas systems. Biochem. Soc. Trans. 41, 1392–1400 (2013)
Wiedenheft, B. et al. Structures of the RNA-guided surveillance complex from a bacterial immune system. Nature 477, 486–489 (2011)
Jore, M. M. et al. Structural basis for CRISPR RNA-guided DNA recognition by Cascade. Nature Struct. Mol. Biol. 18, 529–536 (2011)
Reeks, J., Naismith, J. H. & White, M. F. CRISPR interference: a structural perspective. Biochem. J. 453, 155–166 (2013)
Brouns, S. J. et al. Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960–964 (2008)
Sashital, D. G., Jinek, M. & Doudna, J. A. An RNA-induced conformational change required for CRISPR RNA cleavage by the endoribonuclease Cse3. Nature Struct. Mol. Biol. 18, 680–687 (2011)
Gesner, E. M., Schellenberg, M. J., Garside, E. L., George, M. M. & Macmillan, A. M. Recognition and maturation of effector RNAs in a CRISPR interference pathway. Nature Struct. Mol. Biol. 18, 688–692 (2011)
Haurwitz, R. E., Jinek, M., Wiedenheft, B., Zhou, K. & Doudna, J. A. Sequence- and structure-specific RNA processing by a CRISPR endonuclease. Science 329, 1355–1358 (2010)
Nam, K. H. et al. Cas5d protein processes pre-crRNA and assembles into a cascade-like interference complex in subtype I–C/Dvulg CRISPR-Cas system. Structure 20, 1574–1584 (2012)
Lintner, N. G. et al. Structural and functional characterization of an archaeal clustered regularly interspaced short palindromic repeat (CRISPR)-associated complex for antiviral defense (CASCADE). J. Biol. Chem. 286, 21643–21656 (2011)
Hrle, A. et al. Structure and RNA-binding properties of the type III-A CRISPR-associated protein Csm3. RNA Biol. 10, 1670–1678 (2013)
Wiedenheft, B. et al. RNA-guided complex from a bacterial immune system enhances target recognition through seed sequence interactions. Proc. Natl Acad. Sci. USA 108, 10092–10097 (2011)
Semenova, E. et al. Interference by clustered regularly interspaced short palindromic repeat (CRISPR) RNA is governed by a seed sequence. Proc. Natl Acad. Sci. USA 108, 10098–10103 (2011)
Künne, T., Swarts, D. C. & Brouns, S. J. Planting the seed: target recognition of short guide RNAs. Trends Microbiol. 22, 74–83 (2014)
Sashital, D. G., Wiedenheft, B. & Doudna, J. A. Mechanism of foreign DNA selection in a bacterial adaptive immune system. Mol. Cell 46, 606–615 (2012)
Hochstrasser, M. L. et al. CasA mediates Cas3-catalyzed target degradation during CRISPR RNA-guided interference. Proc. Natl Acad. Sci. USA 111, 6618–6623 (2014)
Wang, Y., Sheng, G., Juranek, S., Tuschl, T. & Patel, D. J. Structure of the guide-strand-containing argonaute silencing complex. Nature 456, 209–213 (2008)
Westra, E. R. et al. CRISPR immunity relies on the consecutive binding and degradation of negatively supercoiled invader DNA by Cascade and Cas3. Mol. Cell 46, 595–605 (2012)
Mulepati, S. & Bailey, S. In vitro reconstitution of an Escherichia coli RNA-guided immune system reveals unidirectional, ATP-dependent degradation of DNA target. J. Biol. Chem. 288, 22184–22192 (2013)
Sinkunas, T. et al. In vitro reconstitution of Cascade-mediated CRISPR immunity in Streptococcus thermophilus. EMBO J. 32, 385–394 (2013)
Jackson, R. N., Lavin, M., Carter, J. & Wiedenheft, B. Fitting CRISPR-associated Cas3 into the helicase family tree. Curr. Opin. Struct. Biol. 24, 106–114 (2014)
Sinkunas, T. et al. Cas3 is a single-stranded DNA nuclease and ATP-dependent helicase in the CRISPR/Cas immune system. EMBO J. 30, 1335–1342 (2011)
Battye, T. G., Kontogiannis, L., Johnson, O., Powell, H. R. & Leslie, A. G. iMOSFLM: a new graphical interface for diffraction-image processing with MOSFLM. Acta Crystallogr. D 67, 271–281 (2011)
Otwinowski, Z. & Minor, W. Processing of X-Ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Xiong, Y. From electron microscopy to X-ray crystallography: molecular-replacement case studies. Acta Crystallogr. D 64, 76–82 (2008)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993)
Acknowledgements
We thank the staff at beamline BL-17U at Shanghai Synchrotron Radiation Facility (SSRF), and beamlines BL-1A, BL-5A and BL-17A at Photon Factory. The research was funded by Chinese Ministry of Science and Technology (2014CB910102 and 2011CBA01105), the Natural Science Foundation of China (31222014 and 31170705), and the Strategic Priority Research program of the Chinese Academy of Sciences (XDB08010203) to Y.W., and was supported by the Research Startup Fund from South University of Science and Technology of China and Shenzhen Government to Z.W. We thank D. Patel for assistance with manuscript editing, H. Wang, H. Li and F. Sun for experimental assistance, and T. Juelich for critical reading and editing of our manuscript.
Author information
Authors and Affiliations
Contributions
H.Z., G.S. and M.W. expressed and purified and grew crystals of the Cascade complex. H.Z. and Y.W. collected x-ray diffraction data, J.W. made all constructs and did biochemical assays and Z.W., Y.W., G.B., H.Z. and W.G. solved the Cascade complex structure, Y.W. and Z.W. wrote the paper. All studies were undertaken under the supervision of Y.W.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Overall surface view of the Cascade complex with the same orientations as those used for Fig. 1b.
Extended Data Figure 2 Schematic summary of Cas-protein–crRNA interactions.
Hydrogen bonds and salt bridges are indicated by dashed lines. Cation-π interactions are indicated by wavy lines.
Extended Data Figure 3 Structure of CasE subunit in the Cascade complex.
a, CasE is shown as a surface representation, and is labelled according to electrostatic potential (red, negative charge; blue, positive charge), and RNA is shown in ribbon representation (orange). b, c, Magnified view of the sequence-specific interaction between CasE and the major groove of 3′-end repeat (b), and the interaction between CasE and 3′-end repeat backbone (c). d, Expanded view of the interaction between CasE and CasC1.
Extended Data Figure 4 The β-hairpin arm of CasD is critical for Cascade function.
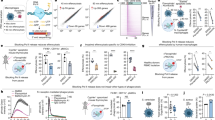
a, pACYC-target invasion assay. Competent E. coli BL21 (DE3) cells expressing either wild type Cascade (WT) or a mutant form in which the hairpin-arm was replaced with a (GGS)4 linker CasD mutant (DΔ), or R108A, R108A/Y142A CasD mutants, were transformed with either pACYC plasmid (Control) or pACYC-target plasmids (Target). Colony-forming units per microgram pACYC (CFU per μg) are depicted for each of the strains. The statistics was based on 10 replicates. Error bars represent the standard error of the mean (s.e.m.). Upon transformation with pACYC plasmid control, E. coli cells expressing WT Cascade exhibited much higher transformation efficiencies than cells transformed with pACYC-target plasmid. However, the transformation efficiencies were high in E. coli cells expressing mutant CasD and transformed with either pACYC control or pACYC-target. These results show that the hairpin-arm of CasD is essential for proper function of the Cascade complex. b, Plasmid loss assay. E. coli BL21 (DE3) cells expressing wild-type or casDΔ(75–104)-replaced Cascade complex and pACYC-target, were serially diluted and grown on either non-selective or selective plates upon induction with 0.5 mM IPTG. Growth of BL21 cells expressing WT Cascade was seriously inhibited on selection medium compared with growth on non-selection medium. However, BL21 cells with mutant CasD displayed similar growth on selection and non-selection medium, indicating that the target sequence was retained upon introduction of the CasD mutant. c, Interface between CasD (green) and CasA (salmon). d, Interface between CasD (green) and CasC6 (blue). e, Pull-down and gel-filtration analysis of the Cascade assembly. WT Cascade complex was formed effectively in the Ni-NTA column and was eluted out with elution buffer containing 100 mM imidazole. In contrast, very limited amounts of the co-expressed Cascade complex with the DΔ mutant or CΔ mutant were obtained in the 100 mM imidazole elution, with the majority of the Cas proteins were in the flow-through. The fractions eluted with 100 mM imidazole were collected and applied for the analytical gel filtration assay. Compared with the WT Cascade, both DΔ mutant and CΔ mutant Cascade did not form a stable complex in gel filtration, further confirming the critical role of the β-hairpin arm of CasD and CasC for the assembly of the Cascade complex. The CasC mutant (CΔ) contains F200A mutation as well as replacement mutation, where residues D204 to L206 were replaced by Ala. f, Structural comparison among E. coli CasD (green) and Cas5d in the RNA-free state of Streptococcus pyogenes (wheat, PDB id: 3VZH) and Bacillus halodurans (grey, PDB id: 4F3M). The β-hairpin arm of CasD is highlighted by a red dashed circle.
Extended Data Figure 5 The unique multi-kinked conformation within the crRNA spacer region.
a, The six CasC subunits form a right-handed helix, with a groove for crRNA binding. Except for CasC1, the β-hairpin (highlighted by a black arrow) of each CasC molecule extends up into the groove of its preceding CasC, and thus results in five kinks observed for the crRNA spacer. b, Superposition of CasC5 (cyan) and CasC6 (blue). The top of long β-hairpin in CasD is shown as green spheres, depicting the clash between the distal domain of CasC6 and the long β-hairpin top given it adopts the same conformation as CasC5. c, CasC (cyan) shares a similar overall fold with Sulfolobus solfataricus Cas7 (grey, PDB id 3PS0). d, The pACYC-target invasion assay showing that mutating three interacting residues (D204CasC, D205CasC, L206CasC) at the β-hairpin arm largely impairs Cascade activity. e, Four designed DNA targets with complementary sequence (WT) to the spacer segment of crRNA or with non-complementary mutations at kinked sites (M1), at immediately upstream nucleotides (M2), and at immediately downstream nucleotides (M3). The five kinked sites are highlighted as yellow background, with the mutated sequences in purple. f, ITC-based analysis of the Cascade complex and four DNA targets interactions. ITC analysis showed that the mutations of the flipping of these residues had no obvious effect on target recognition. However, mutations in either the first or the last nucleotides in the 5-nucleotide stacked region largely affected target binding.
Extended Data Figure 6 Structural analysis of the crRNA spacer segments.
a, The first 5-nucleotide segment in the spacer region is positioned in the positively charged groove of CasC. b, Overlap of the five 5-nucleotide segments within the crRNA space region and their interacting Cas proteins. The 5-nucleotide segments are interrupted by single flipped-out bases, which are indicated by a black arrow, thereby resulting in kinks at these positions. The conformations of the five segments together with the kink sites are essentially identical. Like those of CasC2–6, the β-hairpin arm of CasD is involved in maintaining the kink at G(−1). Interestingly, although the overall foldings are distinct between CasD and CasC, their β-hairpin arms share similar conformations, which are highlighted by a black box. c–g, Structural comparison of the five 5-nucleotide stacked segments with Cas proteins shown in the same orientation. The Cascade complex is colour-coded by chains as shown in Fig. 1.
Extended Data Figure 7 Inter-subunit interactions required for Cascade assembly.
a, Ribbon representation of the organization of CasA and CasB dimers and their associations with the CasC helix. The inter-subunit interfaces are highlighted with dashed-line boxes. b–f, The structural details of the CasA–CasC (b), CasA–CasB2 (c), CasB1–CasB2 (d), CasB1–CasC (e) and CasB2–CasC (f) interactions. The five triads, which connect CasA and the CasB dimer to the CasC helix, are highlighted with red circles. Hydrogen bonds and salt bridges are indicated by dashed lines.
Extended Data Figure 8 Structural comparison between target-bound and unbound Cascade.
a, The building of the structural model of the target-bound Cascade. The subunits were individually docked into the 9Å-electron microscopy map. The fitted model is shown in two views with the electron microscopy density overlapping (contoured at 2σ). b, Ribbon representation of the target-bound Cascade. c, Subunit motions. The movement was represented by red-green lines, which were drawn by connecting each Cα atom (green) in the target-bound Cascade to corresponding Cα atom (red) in the unbound Cascade after the two Cascade structures were superimposed. Thus, the lengths of the lines correlate with the motion scale. The motions of the outer and inner layers are shown by superimposing the target-bound with unbound Cascade structure in d and e, respectively. The target-free structures are rendered semi-transparent. The translational and rotational motions are indicated by arrows showing the motional direction with translation distances and rotation angles labelled, respectively. f, g, Comparison of electrostatic surface potentials for the CasB dimer in the target-free (f) and target-bound Cascade (g) with cartoon representations of the crRNA spacer and its paring target (green). A highly positively charged groove of the CasB dimer fits well with the negatively charged target backbone in g, but not in f. Also, the paring target partially clashes with the CasB dimer in f, with the clashing site highlighted by a yellow circle. The structural analysis suggests that the motion of the CasB dimer identified in the target-bound Cascade complex is required for proper target binding.
Extended Data Figure 9 Electron density maps showing the model quality.
a, The 3.5 Å-electron density map contoured at 1.5σ generated from the phase after density modification treatment with the final structural model superimposed. Cα backbones are shown in with the same colour-coding as used in Fig. 1. b, The 2Fo − Fc map contoured at 1σ of the whole crRNA is shown with the refined RNA model superimposed. Zoom-in views of the 5′-end, 3′-end and the 5-nucleotide stacked segment are shown in the lower left, lower right and upper insets, respectively.
Rights and permissions
About this article
Cite this article
Zhao, H., Sheng, G., Wang, J. et al. Crystal structure of the RNA-guided immune surveillance Cascade complex in Escherichia coli. Nature 515, 147–150 (2014). https://doi.org/10.1038/nature13733
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13733
This article is cited by
-
Dynamic mechanisms of CRISPR interference by Escherichia coli CRISPR-Cas3
Nature Communications (2022)
-
DNA interference is controlled by R-loop length in a type I-F1 CRISPR-Cas system
BMC Biology (2020)
-
Targeted transcriptional modulation with type I CRISPR–Cas systems in human cells
Nature Biotechnology (2019)
-
The repurposing of type I-E CRISPR-Cascade for gene activation in plants
Communications Biology (2019)
-
CRISPR analysis suggests that small circular single-stranded DNA smacoviruses infect Archaea instead of humans
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.