Abstract
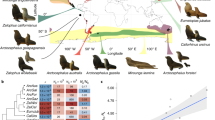
Genetic diversity is the amount of variation observed between DNA sequences from distinct individuals of a given species. This pivotal concept of population genetics has implications for species health, domestication, management and conservation. Levels of genetic diversity seem to vary greatly in natural populations and species, but the determinants of this variation, and particularly the relative influences of species biology and ecology versus population history, are still largely mysterious1,2. Here we show that the diversity of a species is predictable, and is determined in the first place by its ecological strategy. We investigated the genome-wide diversity of 76 non-model animal species by sequencing the transcriptome of two to ten individuals in each species. The distribution of genetic diversity between species revealed no detectable influence of geographic range or invasive status but was accurately predicted by key species traits related to parental investment: long-lived or low-fecundity species with brooding ability were genetically less diverse than short-lived or highly fecund ones. Our analysis demonstrates the influence of long-term life-history strategies on species response to short-term environmental perturbations, a result with immediate implications for conservation policies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Change history
12 November 2014
A minor change was made to the author affiliations.
References
Lewontin, R. The Genetic Basis of Evolutionary Change (Columbia Univ. Press, 1974)
Leffler, E. M. et al. Revisiting an old riddle: what determines genetic diversity levels within species? PLoS Biol. 10, e1001388 (2012)
Keller, L. F. & Waller, D. M. Inbreeding effects in wild populations. Trends Ecol. Evol. 17, 19–23 (2002)
Reusch, T. B. H., Ehlers, A., Hämmerli, A. & Worm, B. Ecosystem recovery after climatic extremes enhanced by genotypic diversity. Proc. Natl Acad. Sci. USA 102, 2826–2831 (2005)
Hughes, A. R., Inouye, B. D., Johnson, M. T. J., Underwood, N. & Vellend, M. Ecological consequences of genetic diversity. Ecol. Lett. 11, 609–623 (2008)
Johnson, W. E. et al. Genetic restoration of the Florida panther. Science 329, 1641–1645 (2010)
Nair, P. Conservation genomics. Proc. Natl Acad. Sci. USA 111, 569 (2014)
Bazin, E., Glémin, S. & Galtier, N. Population size does not influence mitochondrial genetic diversity in animals. Science 312, 570–572 (2006)
Nabholz, B., Mauffrey, J.-F., Bazin, E., Galtier, N. & Glemin, S. Determination of mitochondrial genetic diversity in mammals. Genetics 178, 351–361 (2008)
Perry, G. et al. Comparative RNA sequencing reveals substantial genetic variation in endangered primates. Genome Res. 22, 602–610 (2012)
Banks, S. C. et al. How does ecological disturbance influence genetic diversity? Trends Ecol. Evol. 28, 670–679 (2013)
Hewitt, G. The genetic legacy of the Quaternary ice ages. Nature 405, 907–913 (2000)
Ellegren, H. Genome sequencing and population genomics in non-model organisms. Trends Ecol. Evol. 29, 51–63 (2014)
Cahais, V. et al. Reference-free transcriptome assembly in non-model animals from next-generation sequencing data. Mol. Ecol. Resources 12, 834–845 (2012)
Tsagkogeorga, G., Cahais, V. & Galtier, N. The population genomics of a fast evolver: high levels of diversity, functional constraint, and molecular adaptation in the tunicate Ciona intestinalis. Genome Biol. Evol. 4, 740–749 (2012)
Gayral, P. et al. Reference-free population genomics from next-generation transcriptome data and the vertebrate-invertebrate gap. PLoS Genet. 9, e1003457 (2013)
MacArthur, R. H. & Wilson, E. O. The Theory of Island Biogeography (Princeton Univ. Press, 1967)
Lynch, M. Evolution of the mutation rate. Trends Genet. 26, 345–352 (2010)
Ohta, T. Slightly deleterious mutant substitutions in evolution. Nature 246, 96–98 (1973)
Pinsky, M. L. et al. Unexpected patterns of fisheries collapse in the world’s oceans. Proc. Natl Acad. Sci. USA 108, 8317–8322 (2011)
Gayral, P. et al. Next-generation sequencing of transcriptomes: a guide to RNA isolation in non-model animals. Mol. Ecol. Resources 11, 650–661 (2011)
Guéguen, L. et al. Bio++: efficient, extensible libraries and tools for molecular evolution. Mol. Biol. Evol. 30, 1745–1750 (2013)
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984)
Philippe, H. et al. Phylogenomics revives traditional views on deep animal relationships. Curr. Biol. 19, 706–712 (2009)
Wilfert, L., Gadau, J. & Schmid-Hempel, P. Variation in genomic recombination rates among animal taxa and the case of social insects. Heredity 98, 189–197 (2007)
Acknowledgements
We thank the following for providing samples: F. Delsuc, E. Douzery, M. Tilak, G. Dugas, S. Harispe, C. Benoist, D. Bouchon (woodlice), J. Bierne, M. Bierne, B. Houseaux, M. Strand, C. Lemaire, D. Lallias, Service Modèle Biologique Station Marine Roscoff (nemertines), X. Turon, S. Lopez-Legentil (Cystodytes), P. Jarne, P. David, R. Dillon, J. Auld, R. Relyea, C. Lively, J. Jokela, V. Poullain, T. Stewart (snails), S. Lapègue, V. Boulo, F. Batista, D. Lallias, L. Fast Jensen, M. Cantou (oysters), J. Do Nascimento, C. Daguin-Thiébaut, M. Cantou (crabs), L. Bonnaud (cuttlefish), D. Aurelle (gorgonians), F. Viard, Y. Pechenik, A. Cahill, R. Colins (slipper limpets), L. Dupont (earthworms), D. Jollivet (trumpet worms), M. A. Felix, I. Nuez (nematodes), N. Rodes, T. Lenormand, E. Flaven (brine shrimps), Rotterdam Zoo, Zurich Zoo, C. Libert, Montpellier Zoo, S. Martin, la Ferme aux Crocodiles, O. Verneau, C. Ayres, M. Carretero, M. Vanberger, K. Pobolsaj, M. Zuffi, C. Palacios, L. du Preez, B. Halpern, Budapest Zoo (turtles), P. Peret, C. Doutrelant, B. Halpern, B. Rosivall (tits), M. de Dinechin, B. Rey (penguins), Z. Melo-Fereira, P. Alves (hares), N. Brand, M. Chapuisat (bees), R. Blatrix, A. Lenoir, I. Nodet, A. Lugagne, S. Blanquart, L. Serres-Giardi, V. Roustang, N. François, G. Ballantyne, A. Carbonnel, Y. Samuel, G. James, G. Kalytta, F. Guerrini, S. Stenzel, J. Beekman, X. Cerda, S. Ikoen (ants), I. Hanski, S. Ikonen, J. Kullberg, Z. Kolev (fritillary butterflies), F. Viard, X. Turon, Di Jiang, D. Chourrout, B. Vercaemer, E. Newman-Smith, Ascidian Stock Center, Service Modèle Biologique Station Marine Roscoff (ciona), L. Excoffier, G. Heckel (voles), F. Dedeine (termites), C. Atyame, O. Duron, M. Weill (mosquitoes), M. Cantou, H. Violette, F. Batista, J. Hondeville (seahorses), C. Fraïsse, G. Pogson, N. Saarman, J. Normand (mussels), E. Poulin, C. Gonzalez-Weivar, and J. P. Feral (sea urchins). This work was supported by European Research Council advanced grant 232971 (PopPhyl).
Author information
Authors and Affiliations
Contributions
N.G. conceived the project. P.G., M.B., N.F., Y.C., L.A.W., G.T., A.C., A.W., J.R., N.G. and N.B. performed sampling and laboratory work. A.B., V.C., E.L., J.R., J.M.L., C.R., P.G., G.T., B.N., R.D., K.B., S.G. and N.G. developed the data analysis pipeline. J.R. collected life-history/geographic variables and produced figures. J.R., A.B., V.C., L.D., E.L. and N.G. analysed the data. S.G., N.B., B.N., J.R. and N.G. provided interpretations and models. J.R., N.B., S.G. and N.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Data sets are freely available from the Sequence Read Archive (SRA) database (http://www.ncbi.nlm.nih.gov/sra) under project ID SRP042651 and from the Datasets section of the PopPhyl website (http://kimura.univ-montp2.fr/PopPhyl), in which predicted single nucleotide polymorphisms and genotypes are provided as .vcf files. Scripts and executable files are freely available from the Tools section of the PopPhyl website.
Extended data figures and tables
Extended Data Figure 1 Phylogenetic tree of metazoan orders and the position of the taxa analysed in this study.
The tree topology is consistent with the NCBI taxonomy. Red arrows identify 25 orders that were sampled. Five gastropod species from two distinct families (Calyptraeidae (Crepidula fornicata, C. plana and Bostrycapulus aculeatus) and Physidae (Physa acuta and P. gyrina)) are not represented because they lacked any assignment to an order in current taxonomy.
Extended Data Figure 2 Correlations between genetic diversity and life history variables.
Blue indicates a positive relationship, red a negative one; colour intensity is proportional to Pearson’s correlation coefficient.
Extended Data Figure 3 Family-level phylogenetic tree (31 families included).
The scale is in million years of divergence.
Extended Data Figure 5 Relationship between the πs/propagule-size r2 and the number of sampled loci.
Extended Data Figure 6 Phylum distribution of the first BLAST hit in four representative species.
a, Common vole (Microtus arvalis). b, Glanville fritillary butterfly (Melitaea cinxia). c, Blue mussel (Mytilus edulis). d, Earthworm (Allolobophora chlorotica).
Extended Data Figure 7 Correlation between πn/πs and maximum longevity (P < 10−8, r2 = 0.54).
Only species with at least four individuals are included.
Supplementary information
Supplementary Information
This file contains additional theoretical developments and modelling. (PDF 810 kb)
Supplementary Data
This file contains Supplementary Table 1. (XLS 87 kb)
Supplementary Data
This file contains Supplementary Table 2. (XLS 36 kb)
Supplementary Data
This file contains Supplementary Table 3. (PDF 139 kb)
Supplementary Data
This file contains Supplementary Table 4. (XLS 8 kb)
Supplementary Data
This file contains Supplementary Table 5. (XLS 9 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Romiguier, J., Gayral, P., Ballenghien, M. et al. Comparative population genomics in animals uncovers the determinants of genetic diversity. Nature 515, 261–263 (2014). https://doi.org/10.1038/nature13685
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13685
This article is cited by
-
Remnant kenngoor (Phascogale calura) retain genetic connectivity and genetic diversity in a highly fragmented landscape
Conservation Genetics (2024)
-
Population genetic structure of invasive apple snails Pomacea maculata in Louisiana
Aquatic Ecology (2024)
-
Gene expression response of the non-target gastropod Physella acuta to Fenoxycarb, a juvenile hormone analog pesticide
Scientific Reports (2023)
-
Population transcriptomics uncover the relative roles of positive selection and differential expression in Batrachium bungei adaptation to the Qinghai–Tibetan plateau
Plant Cell Reports (2023)
-
Whole genome re-sequencing uncovers significant population structure and low genetic diversity in the endangered clouded apollo (Parnasssius mnemosyne) in Sweden
Conservation Genetics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.