Abstract
The isolation of human monoclonal antibodies is providing important insights into the specificities that underlie broad neutralization of HIV-1 (reviewed in ref. 1). Here we report a broad and extremely potent HIV-specific monoclonal antibody, termed 35O22, which binds a novel HIV-1 envelope glycoprotein (Env) epitope. 35O22 neutralized 62% of 181 pseudoviruses with a half-maximum inhibitory concentration (IC50) <50 μg ml−1. The median IC50 of neutralized viruses was 0.033 μg ml−1, among the most potent thus far described. 35O22 did not bind monomeric forms of Env tested, but did bind the trimeric BG505 SOSIP.664. Mutagenesis and a reconstruction by negative-stain electron microscopy of the Fab in complex with trimer revealed that it bound to a conserved epitope, which stretched across gp120 and gp41. The specificity of 35O22 represents a novel site of vulnerability on HIV Env, which serum analysis indicates to be commonly elicited by natural infection. Binding to this new site of vulnerability may thus be an important complement to current monoclonal-antibody-based approaches to immunotherapies, prophylaxis and vaccine design.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Electron Microscopy Data Bank
GenBank/EMBL/DDBJ
Protein Data Bank
Data deposits
The 35O22 heavy and light chain plasmids and antibody have been submitted to the NIH-AIDS reagent program. The nucleotide sequence of 35O22 and its variants have been submitted to GenBank under accession numbers KM001872–KM001879 (heavy chain) and KM001880–KM001887 (light chain). The Env nucleotide sequences of the patient (N152) plasma virus are deposited in GenBank under accession numbers KM516886–KM516897. Coordinates and structure factors for 35O22 Fab have been deposited with the Protein Data Bank under accession code 4TOY. The reconstruction of BG505 SOSIP.664 in complex with 35O22 Fab has been deposited in the Electron Microscopy Data Base under accession code EMD-2672.
Change history
05 November 2014
Minor changes were made to the author list, affiliations, Acknowledgements, Author Contributions and Methods.
References
Kwong, P. D. & Mascola, J. R. Human antibodies that neutralize HIV-1: identification, structures, and B cell ontogenies. Immunity 37, 412–425 (2012)
Doria-Rose, N. A. et al. Breadth of human immunodeficiency virus-specific neutralizing activity in sera: clustering analysis and association with clinical variables. J. Virol. 84, 1631–1636 (2010)
Sather, D. N. et al. Factors associated with the development of cross-reactive neutralizing antibodies during human immunodeficiency virus type 1 infection. J. Virol. 83, 757–769 (2009)
Walker, L. M. et al. A limited number of antibody specificities mediate broad and potent serum neutralization in selected HIV-1 infected individuals. PLoS Pathog. 6, e1001028 (2010)
Simek, M. D. et al. Human immunodeficiency virus type 1 elite neutralizers: individuals with broad and potent neutralizing activity identified by using a high-throughput neutralization assay together with an analytical selection algorithm. J. Virol. 83, 7337–7348 (2009)
Gray, E. S. et al. Antibody specificities associated with neutralization breadth in plasma from human immunodeficiency virus type 1 subtype C-infected blood donors. J. Virol. 83, 8925–8937 (2009)
Walker, L. M. et al. Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326, 285–289 (2009)
Wu, X. et al. Rational design of envelope identifies broadly neutralizing human monoclonal antibodies to HIV-1. Science 329, 856–861 (2010)
Huang, J. et al. Isolation of human monoclonal antibodies from peripheral blood B cells. Nature Protocols 8, 1907–1915 (2013)
Tiller, T. et al. Efficient generation of monoclonal antibodies from single human B cells by single cell RT-PCR and expression vector cloning. J. Immunol. Methods 329, 112–124 (2008)
Scheid, J. F. et al. Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature 458, 636–640 (2009)
Bonsignori, M. et al. Two distinct broadly neutralizing antibody specificities of different clonal lineages in a single HIV-1-infected donor: implications for vaccine design. J. Virol. 86, 4688–4692 (2012)
Walker, L. M. et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477, 466–470 (2011)
Burton, D. R. et al. Efficient neutralization of primary isolates of HIV-1 by a recombinant human monoclonal antibody. Science 266, 1024–1027 (1994)
Muster, T. et al. A conserved neutralizing epitope on gp41 of human immunodeficiency virus type 1. J. Virol. 67, 6642–6647 (1993)
Scharf, L. et al. Antibody 8ANC195 reveals a site of broad vulnerability on the HIV-1 envelope spike. Cell Rep. 7, 785–795 (2014)
Falkowska, E. et al. Broadly neutralizing HIV antibodies define a glycan-dependent epitope on the prefusion conformation of gp41 on cleaved envelope trimers. Immunity 40, 657–668 (2014)
Blattner, C. et al. Structural delineation of a quaternary, cleavage-dependent epitope at the gp41-gp120 interface on intact HIV-1 Env trimers. Immunity 40, 669–680 (2014)
Zhang, M. Y. et al. Identification and characterization of a broadly cross-reactive HIV-1 human monoclonal antibody that binds to both gp120 and gp41. PLoS ONE 7, e44241 (2012)
Huang, J. et al. Broad and potent neutralization of HIV-1 by a gp41-specific human antibody. Nature 491, 406–412 (2012)
Scheid, J. F. et al. Sequence and structural convergence of broad and potent HIV antibodies that mimic CD4 binding. Science 333, 1633–1637 (2011)
Klein, F. et al. Somatic mutations of the immunoglobulin framework are generally required for broad and potent HIV-1 neutralization. Cell 153, 126–138 (2013)
Haynes, B. F. et al. Cardiolipin polyspecific autoreactivity in two broadly neutralizing HIV-1 antibodies. Science 308, 1906–1908 (2005)
Mouquet, H. et al. Polyreactivity increases the apparent affinity of anti-HIV antibodies by heteroligation. Nature 467, 591–595 (2010)
Doores, K. J. & Burton, D. R. Variable loop glycan dependency of the broad and potent HIV-1-neutralizing antibodies PG9 and PG16. J. Virol. 84, 10510–10521 (2010)
Sanders, R. W. et al. A next-generation cleaved, soluble HIV-1 Env Trimer, BG505 SOSIP.664 gp140, expresses multiple epitopes for broadly neutralizing but not non-neutralizing antibodies. PLoS Pathog. 9, e1003618 (2013)
Ringe, R. P. et al. Cleavage strongly influences whether soluble HIV-1 envelope glycoprotein trimers adopt a native-like conformation. Proc. Natl Acad. Sci. USA 110, 18256–18261 (2013)
Georgiev, I. S. et al. Delineating antibody recognition in polyclonal sera from patterns of HIV-1 isolate neutralization. Science 340, 751–756 (2013)
Pancera, M. & Wyatt, R. Selective recognition of oligomeric HIV-1 primary isolate envelope glycoproteins by potently neutralizing ligands requires efficient precursor cleavage. Virology 332, 145–156 (2005)
Crooks, E. T. et al. Characterizing anti-HIV monoclonal antibodies and immune sera by defining the mechanism of neutralization. Hum. Antibodies 14, 101–113 (2005)
Tong, T., Crooks, E. T., Osawa, K. & Binley, J. M. HIV-1 virus-like particles bearing pure env trimers expose neutralizing epitopes but occlude nonneutralizing epitopes. J. Virol. 86, 3574–3587 (2012)
Li, M. et al. Human immunodeficiency virus type 1 env clones from acute and early subtype B infections for standardized assessments of vaccine-elicited neutralizing antibodies. J. Virol. 79, 10108–10125 (2005)
Chakrabarti, B. K. et al. Direct antibody access to the HIV-1 membrane-proximal external region positively correlates with neutralization sensitivity. J. Virol. 85, 8217–8226 (2011)
Binley, J. M. et al. Redox-triggered infection by disulfide-shackled human immunodeficiency virus type 1 pseudovirions. J. Virol. 77, 5678–5684 (2003)
Chuang, G. Y. et al. Residue-level prediction of HIV-1 antibody epitopes based on neutralization of diverse viral strains. J. Virol. 87, 10047–10058 (2013)
Pancera, M. et al. Soluble mimetics of human immunodeficiency virus type 1 viral spikes produced by replacement of the native trimerization domain with a heterologous trimerization motif: characterization and ligand binding analysis. J. Virol. 79, 9954–9969 (2005)
Yasmeen, A. et al. Differential binding of neutralizing and non-neutralizing antibodies to native-like soluble HIV-1 Env trimers, uncleaved Env proteins, and monomeric subunits. Retrovirology (in the press) (2014)
McLellan, J. S. et al. Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nature 480, 336–343 (2011)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Macromol. Crystallogr. A 276, 307–326 (1997)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. 60, 2126–2132 (2004)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Kong, L. et al. Supersite of immune vulnerability on the glycosylated face of HIV-1 envelope glycoprotein gp120. Nature Struct. Mol. Biol. 20, 796–803 (2013)
Thornburg, N. J. et al. Human antibodies that neutralize respiratory droplet transmissible H5N1 influenza viruses. J. Clin. Invest. 123, 4405–4409 (2013)
Julien, J. P. et al. Crystal structure of a soluble cleaved HIV-1 envelope trimer. Science 342, 1477–1483 (2013)
Liu, J., Bartesaghi, A., Borgnia, M. J., Sapiro, G. & Subramaniam, S. Molecular architecture of native HIV-1 gp120 trimers. Nature 455, 109–113 (2008)
Acknowledgements
We thank J. P. Moore for experiments using the BG505 SOSIP trimer, discussions of experimental data, and review of this manuscript. We thank K. Lloyd. R. Parks, J. Eudailey and J. Blinn for performing autoantibody assays. We also thank S. Moir for providing patient samples. This project has been funded in part with federal funds from the Intramural Research Programs of NIAID, National Institutes of Health. This work was also supported by the NIH HIVRAD grant P01 AI082362, and by the Aids Fonds Netherlands, grants 2011032 and 2012041. R.W.S. is a recipient of a Vidi grant from the Netherlands Organization for Scientific Research (NWO) and a Starting Investigator Grant from the European Research Council (ERC-StG-2011-280829-SHEV). J.H.L., Y.F., R.T.W. and A.W. are supported through the International AIDS Vaccine Initiative. J.M.B. is supported by NIH grants AI93278 and AI84714. Use of sector 22 (Southeast Region Collaborative Access team) at the Advanced Photon Source was supported by the US Department of Energy, Basic Energy Sciences, Office of Science, under contract number W-31-109-Eng-38. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government.
Author information
Authors and Affiliations
Contributions
M.C., J.H., B.H.K., M.P., J.R.M., P.D.K., J.M.B., R.T.W., R.W.S. and A.B.W. each contributed to the design of the study, analysis of the data, and preparation of this manuscript. J.H. and B.H.K. performed B-cell sorting, antibody cloning, neutralization assays, cell surface staining and epitope mapping assays. H.I. performed the sequencing and analysis of the patient plasma virus. L.L. cultured B cells and assisted with recovery of IgG genes. N.A.D.-R. performed the 35O22 biotinylation and provided data of PG9, PG16, PGT121 and PGT128 breadth and potency. P.-J.K. performed SPR experiments. R.D., K.S., A.T.d.l.P. and P.P. performed the ELISA BG505 SOSIP.664 mutants. M.J.v.G. performed neutralization assays with HIVLAI viruses. M.A. and B.F.H. performed the autoreactivity assays. I.S.G., A.D. and G.-Y.C. performed serological analysis. M.P. and P.D.K. performed 35O22 crystallographic analysis. J.H.L. and A.B.W. performed the negative stain EM studies. J.M.B. and T.T. planned and performed the antibody competition, and HIV VLP entry studies. Y.F. and R.T.W. planned and performed the washout and kinetic studies. S.A.M. led the clinical care of the patients. R.T.B. screened the B-cell culture supernatants for neutralization activity.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Analysis of 35O22 autoreactivity.
a, Reactivity of 35O22 with HEP-2 epithelial cells. 2F5 was used as a positive control and 17b as a negative control. Antibody concentration was 25 μg ml−1. All pictures are shown at 400× magnification. b, SPR analysis of 35O22 binding to anionic phospholipids. 35O22 was injected over PC-CLP liposomes or PC-PS liposomes immobilized on the BIAcore L1 sensor chip. 4E10 and 2F5 were used as positive controls and 13H1 as a negative control. c, Reactivity of 35O22 with autoantigens detected in Luminex assay. 4E10 was used as a positive control. Synagis, an anti-RSV monoclonal antibody, was used as a negative control. SSA, Sjogren’s syndrome antigen A; SSB, Sjogren’s syndrome antigen B; Sm, Smith antigen; RNP, ribonucleoprotein; Scl 70, scleroderma 70; Jo1, antigen; CentrB, centromere B. A positive response is >120 units.
Extended Data Figure 2 Neutralization similarities between 35O22 and other HIV-1 bNAbs.
a, Correlation (Spearman) between the neutralization potencies of 35O22 and the indicated antibody against 172 pseudoviruses. Representatives from all four major sites of vulnerability are shown. Resistant strains corresponding to values of >50 μg ml−1 are plotted as 50. b, Neutralization-based clustering of bNabs over a set of 172 diverse HIV-1 strains. A putative epitope-specific clustering cutoff is shown as a dashed line. Antibodies are coloured according to the respective target site of vulnerability: red, CD4bs; blue, glycan-V3; green, V1V2; light blue, MPER; purple, other. 35O22 (yellow) clusters separately from all other antibodies, indicating a novel mechanism of neutralization. c, 35O22 competition with other bNAbs on HIVJRFL VLPs with the trimer stabilizing SOS mutations in an ELISA assay. Biotin–bNAbs were titrated into the ELISA at increasing concentrations in the presence of excess (10 μg ml−1) cold competitor neutralizing antibodies. Values in the table indicate percentage binding of biotin-nAbs in the presence of cold competitor.
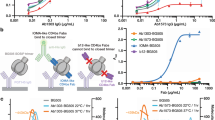
Extended Data Figure 3 35O22 binds to N-linked glycans.
a, Neutralization by 35O22 plateaus below 80% against several pseudoviruses. b, Neutralization activity of monoclonal antibodies against JRCSF pseudoviruses generated in the presence of glycosidase inhibitors, such as kifunensine (25 μM), NB-DNJ (500 μM) or swainsonine (20 μM). Error bars denote one standard error of the mean (s.e.m.).
Extended Data Figure 4 Neutralization of 35O22 against pseudovirus mutants known to knock out activity against known glycan-specific antibodies.
a, Neutralization of 35O22 against JRCSF or KER2018.11 with or without the N160K mutation. PG9 and PG16 were used as positive controls. b, Neutralization of 35O22 against N332A mutants of JRCSF. PGT121 was used as a positive control. c, Neutralization of 35O22 against N234S, T236K and N276D mutants of 3337.V2.C6. 8ANC195 was used as a positive control. Error bars denote one standard error of the mean (s.e.m.).
Extended Data Figure 5 Binding specificity of 35O22.
a, ELISA binding of indicated monoclonal antibodies to HIVYU2 gp140 foldon trimer, gp120 and gp41 monomers. b, c, ELISA binding of gp120 (b) and gp140 (c) monomers from different HIV-1 subtypes.
Extended Data Figure 6 35O22 Fab features.
a, 35O22 is seen looking down on the combining site from the viewpoint of antigen in ribbons with the CDR coloured as in Fig. 3a. Insets (bottom row) show structural details of the framework 3 insertion and disulphides in CDR L1 and CDR L3 with electron density 2Fo − Fc contoured at 1σ. b, Superposition of HIVBaL gp160 negative stain (yellow surface) with the negative stain reconstruction of soluble BG505 SOSIP in complex with 35O22 (grey surface) gives an estimation of the viral membrane location relative to 35O22 antibody as shown in Fig. 3b.
Extended Data Figure 7 Binding site of 35O22 on the HIV Env trimer.
Binding site of 35O22 (red) relative to those of PGT151 (blue) or 8ANC195 (green) are shown.
Extended Data Figure 8 A new site of HIV-1 vulnerability at the interface of gp120 and gp41 and prevalence of targeting.
a, Dominant sites of vulnerability to neutralizing antibody elicited by natural infection, shown in the context of an EM tomogram from the BaL viral spike. The viral membrane is positioned at the top of the spike. It is unclear if 35O22 and MPER antibodies bind to this form of the viral spike, and approximate locations for these are shown in dotted outlines. b, Viral spike from the soluble BG505 SOSIP context, shown in the same orientation as a, with gp120 surface coloured by conservation from 0% to 100%, from 4,265 HIV-1 strains (white to purple for protomer 1 with scale shown, white to blue for protomer 2, and white to orange for protomer 3), with glycans shown in green when present in more than 90% of strains, in grey when present in 30–90% of strains and not shown otherwise. c, 35O22-identified site of HIV-1 vulnerability comprises both conserved amino acids and a cluster of glycans, including N88 from gp120 and N625 from gp41. N230 and N241 are not present in BG505 strain. The 35O22 epitope is shown by a yellow dotted line. d, Neutralization fingerprints for 35O22 and for antibodies encompassing ten different epitope specificities representing the other four known major sites of Env vulnerability were used to interrogate the serum specificities of 34 HIV-infected patients. Values (with proportional colour intensities) predict the fraction of serum neutralization that can be attributed to each antibody specificity. Possible 35O22-like signals were predicted for 13 of the sera (values >0.2), while strong signals were observed in 3 of the sera (values >0.3). A panel of 21 HIV-1 strains was used in the neutralization analysis and for computing serum breadth. e, Sites of HIV-1 vulnerability to neutralizing antibody outlined by a yellow line. Prevalence in a 34-donor cohort and critical glycans are indicated.
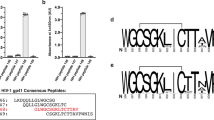
Extended Data Figure 9 Autologous virus Env sequences and the impact of variants on 35O22 neutralization.
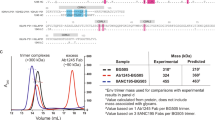
a, A total of 12 single-genome amplicons from plasma of patient N152 were sequenced. Donor Env sequences together with the reference sequences of JRCSF and LAI are aligned. Amino acids critical for 35O22 neutralization of JRCSF and LAI are labelled in yellow. Differences between autologous and JRCSF sequences are labelled in green. b, 35O22 neutralization of JRCSF pseudovirus or variants containing the autologous virus mutations from patient N152. Error bars denote one standard error of the mean.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-8. (PDF 670 kb)
Rights and permissions
About this article
Cite this article
Huang, J., Kang, B., Pancera, M. et al. Broad and potent HIV-1 neutralization by a human antibody that binds the gp41–gp120 interface. Nature 515, 138–142 (2014). https://doi.org/10.1038/nature13601
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13601
This article is cited by
-
Absolute quantitation of binding antibodies from clinical samples
npj Vaccines (2024)
-
Inactivation of cell-free HIV-1 by designing potent peptides based on mutations in the CD4 binding site
Medical & Biological Engineering & Computing (2024)
-
Conformational antigenic heterogeneity as a cause of the persistent fraction in HIV-1 neutralization
Retrovirology (2023)
-
Structures and immune recognition of Env trimers from two Asia prevalent HIV-1 CRFs
Nature Communications (2023)
-
Profound structural conservation of chemically cross-linked HIV-1 envelope glycoprotein experimental vaccine antigens
npj Vaccines (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.