Abstract
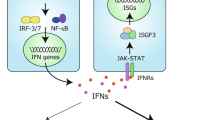
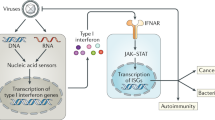
Mammalian cells possess mechanisms to detect and defend themselves from invading viruses. In the cytosol, the RIG-I-like receptors (RLRs), RIG-I (retinoic acid-inducible gene I; encoded by DDX58) and MDA5 (melanoma differentiation-associated gene 5; encoded by IFIH1) sense atypical RNAs associated with virus infection1,2. Detection triggers a signalling cascade via the adaptor MAVS that culminates in the production of type I interferons (IFN-α and β; hereafter IFN), which are key antiviral cytokines. RIG-I and MDA5 are activated by distinct viral RNA structures and much evidence indicates that RIG-I responds to RNAs bearing a triphosphate (ppp) moiety in conjunction with a blunt-ended, base-paired region at the 5′-end (reviewed in refs 1, 2, 3). Here we show that RIG-I also mediates antiviral responses to RNAs bearing 5′-diphosphates (5′pp). Genomes from mammalian reoviruses with 5′pp termini, 5′pp-RNA isolated from yeast L-A virus, and base-paired 5′pp-RNAs made by in vitro transcription or chemical synthesis, all bind to RIG-I and serve as RIG-I agonists. Furthermore, a RIG-I-dependent response to 5′pp-RNA is essential for controlling reovirus infection in cultured cells and in mice. Thus, the minimal determinant for RIG-I recognition is a base-paired RNA with 5′pp. Such RNAs are found in some viruses but not in uninfected cells, indicating that recognition of 5′pp-RNA, like that of 5′ppp-RNA, acts as a powerful means of self/non-self discrimination by the innate immune system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Goubau, D., Deddouche, S. & Reis e Sousa, C. Cytosolic sensing of viruses. Immunity 38, 855–869 (2013)
Schlee, M. Master sensors of pathogenic RNA–RIG-I like receptors. Immunobiology 218, 1322–1335 (2013)
Rehwinkel, J. & Reis e Sousa, C. Targeting the viral Achilles’ heel: recognition of 5′-triphosphate RNA in innate anti-viral defence. Curr. Opin. Microbiol. 16, 485–492 (2013)
Holm, G. H. et al. Retinoic acid-inducible gene-I and interferon-beta promoter stimulator-1 augment proapoptotic responses following mammalian reovirus infection via interferon regulatory factor-3. J. Biol. Chem. 282, 21953–21961 (2007)
Kato, H. et al. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J. Exp. Med. 205, 1601–1610 (2008)
Loo, Y.-M. et al. Distinct RIG-I and MDA5 signaling by RNA viruses in innate immunity. J. Virol. 82, 335–345 (2008)
Pichlmair, A. et al. Activation of MDA5 requires higher-order RNA structures generated during virus infection. J. Virol. 83, 10761–10769 (2009)
Lu, C. et al. The structural basis of 5′ triphosphate double-stranded RNA recognition by RIG-I C-terminal domain. Structure 18, 1032–1043 (2010)
Luo, D. et al. Structural insights into RNA recognition by RIG-I. Cell 147, 409–422 (2011)
Jiang, F. et al. Structural basis of RNA recognition and activation by innate immune receptor RIG-I. Nature 479, 423–427 (2011)
Wang, Y. et al. Structural and functional insights into 5′-ppp RNA pattern recognition by the innate immune receptor RIG-I. Nature Struct. Mol. Biol. 17, 781–787 (2010)
Dermody, T. S., Sherry, B. & Parker, J. S. L. in Fields Virology 6th edn ( Knipe, D. M. & Howley, P. M. ) 2, 1304–1346 (Wolters Kluwer Health/Lippincott Williams & Wilkins, 2013)
Rehwinkel, J. et al. RIG-I detects viral genomic RNA during negative-strand RNA virus infection. Cell 140, 397–408 (2010)
Baum, A., Sachidanandam, R. & García-Sastre, A. Preference of RIG-I for short viral RNA molecules in infected cells revealed by next-generation sequencing. Proc. Natl Acad. Sci. USA 107, 16303–16308 (2010)
Fujimura, T. & Esteban, R. Yeast double-stranded RNA virus L-A deliberately synthesizes RNA transcripts with 5′-diphosphate. J. Biol. Chem. 285, 22911–22918 (2010)
Fujimura, T. & Esteban, R. Cap-snatching mechanism in yeast L-A double-stranded RNA virus. Proc. Natl Acad. Sci. USA 108, 17667–17671 (2011)
Hornung, V. et al. 5′-triphosphate RNA is the ligand for RIG-I. Science 314, 994–997 (2006)
Schlee, M. et al. Recognition of 5′ triphosphate by RIG-I helicase requires short blunt double-stranded RNA as contained in panhandle of negative-strand virus. Immunity 31, 25–34 (2009)
Schmidt, A. et al. 5′-triphosphate RNA requires base-paired structures to activate antiviral signaling via RIG-I. Proc. Natl Acad. Sci. USA 106, 12067–12072 (2009)
Goldeck, M., Tuschl, T., Hartmann, G. & Ludwig, J. Efficient solid-phase synthesis of pppRNA by using product-specific labeling. Angew. Chem. Int. Edn Engl. 53, 4694–4698 (2014)
Vela, A., Fedorova, O., Ding, S. C. & Pyle, A. M. The thermodynamic basis for viral RNA detection by the RIG-I innate immune sensor. J. Biol. Chem. 287, 42564–42573 (2012)
Kohlway, A., Luo, D., Rawling, D. C., Ding, S. C. & Pyle, A. M. Defining the functional determinants for RNA surveillance by RIG-I. EMBO Rep. 14, 772–779 (2013)
Grunberg-Manago, M., Oritz, P. J. & Ochoa, S. Enzymatic synthesis of nucleic acidlike polynucleotides. Science 122, 907–910 (1955)
Decroly, E., Ferron, F., Lescar, J. & Canard, B. Conventional and unconventional mechanisms for capping viral mRNA. Nature Rev. Microbiol. 10, 51–65 (2012)
Gerlier, D. & Lyles, D. S. Interplay between innate immunity and negative-strand RNA viruses: towards a rational model. Microbiol. Mol. Biol. Rev. 75, 468–490 (2011)
Inaba, K. et al. Generation of large numbers of dendritic cells from mouse bone marrow cultures supplemented with granulocyte/macrophage colony-stimulating factor. J. Exp. Med. 176, 1693–1702 (1992)
Ran, F. A. et al. Genome engineering using the CRISPR-Cas9 system. Nature Protocols 8, 2281–2308 (2013)
Kobayashi, T., Ooms, L. S., Ikizler, M., Chappell, J. D. & Dermody, T. S. An improved reverse genetics system for mammalian orthoreoviruses. Virology 398, 194–200 (2010)
Barton, E. S., Connolly, J. L., Forrest, J. C., Chappell, J. D. & Dermody, T. S. Utilization of sialic acid as a coreceptor enhances reovirus attachment by multistep adhesion strengthening. J. Biol. Chem. 276, 2200–2211 (2001)
Virgin, H. W., Bassel-Duby, R., Fields, B. N. & Tyler, K. L. Antibody protects against lethal infection with the neurally spreading reovirus type 3 (Dearing). J. Virol. 62, 4594–4604 (1988)
Diaz-ruiz, J. R. & Kaper, J. M. Isolation of viral double-stranded RNAs using a LiCl fractionation procedure. Prep. Biochem. 8, 1–17 (1978)
Diebold, S. S. et al. Viral infection switches non-plasmacytoid dendritic cells into high interferon producers. Nature 424, 324–328 (2003)
Boehme, K. W., Frierson, J. M., Konopka, J. L., Kobayashi, T. & Dermody, T. S. The reovirus sigma1s protein is a determinant of hematogenous but not neural virus dissemination in mice. J. Virol. 85, 11781–11790 (2011)
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013)
Acknowledgements
We thank S. Akira and J. Tschopp (deceased) for gifts of mice and cells, as well as N. O’Reilly, the LRI Equipment Park (D. Phillips), and the LRI Protein Analysis and Proteomics Facility (R. George and S. Kjaer) for technical assistance. We also thank P. Maillard and K. Snelgrove for reading the manuscript, P. Tortora, G. Dehò and M. Freire for their insights on the synthesis of poly(I:C) and all members of the CRUK Immunobiology Laboratory for helpful discussions and comments. C.R.S., D.G., S.D. and A.G.V.V. are funded by Cancer Research UK, a prize from Fondation Bettencourt-Schueller, and a grant from the European Research Council (ERC Advanced Researcher Grant AdG-2010-268670). A.J.P. and T.S.D. are supported by Public Health Service award R37 AI038296 and the Elizabeth B. Lamb Center for Pediatric Research. T.F. is supported by the Fundación Ramón Areces. G.H., M.S. and W.B. are supported by the Deutsche Forschungsgemeinschaft (http://www.dfg.de; SFB670 to M.S., W.B. and G.H., DFG SCHL1930/1-1 to M.S., SFB704 to G.H. and W.B., SFB832 and KFO177 to G.H.). G.H. and M.S. are supported by the DFG Excellence Cluster ImmunoSensation. G.H. is supported by the German Center of Infectious Disease (DZIF).
Author information
Authors and Affiliations
Contributions
D.G., M.S., S.D., A.J.P., T.S.D., M.G., W.B., J.L., G.H. and C.R.S. designed experiments and analysed the data. D.G., M.S., S.D., A.J.P., T.F., A.G.V.V., J.R., J.A.I., T.Z., C.S., M.G., J.L. performed experiments. D.G., M.S., A.J.P., T.S.D., G.H. and C.R.S. wrote the manuscript. G.H. and C.R.S. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 RNA from reovirus and L-A virus induce a RIG-I-dependent IFN response that requires 5′-diphosphates.
a, Total RNA purified from reoT1L particles (vRNA) was treated or not with calf intestinal phosphatase (± CIP). RNA integrity was verified by gel electrophoresis (left panel) or transfected into HEK293 cells to determine its capacity to stimulate the IFN-β promoter using a reporter assay (right panel). b, L, M, and S reoT1L genome segments were isolated by gel fractionation and treated or not with CIP. An aliquot of the treated samples was electrophoresed in a 0.8% agarose gel to validate RNA integrity (left panel), whereas another was transfected into HEK293 cells to determine its capacity to stimulate the IFN-β promoter using a reporter assay (right panel). c, d, Total L-A RNA as well as gel-purified L-A genomes and transcripts (as in Fig. 1i) were transfected into RIG-I+/− or RIG-I−/− MEFs (c) and MDA5+/+ or MDA5−/− DCs (d). After incubation for 16 h, the relative expression (RE) of ifit1 over gapdh (c) or murine IFN-α levels (d) were determined. Water and ppp-IVT-RNA99nt, poly(dA:dT), or RNA isolated from Vero cells infected with encephalomyocarditis virus (Vero-EMCV-RNA) were included as controls (* = none detected). All experiments were performed at least twice; one representative experiment is shown.
Extended Data Figure 2 RIG-I associates with stimulatory RNA following reovirus infection.
This experiment was conducted exactly as in Fig. 2a but using strain reoT3D.
Extended Data Figure 3 Characterization of guanosine sources and IVT-RNA25nt.
a, Representative LC-MS spectra of GMP, GDP, and GTP sources used for the preparation of IVT-RNAs in Fig. 3. Asterisks indicate the expected mass-to-charge ratio (m/z) of the different guanosines. b, IVT-RNA25nt were generated as depicted in Fig. 3a using GMP, GTP or GMP spiked with GTP (GMP + 10% GTP) before being annealed to AS RNA and tested using the IFN-β promoter reporter assay following transfection into HEK293 cells. c, Spectra of 5′pp-RNA24nt and 5′ppp-RNA24nt following MALDI ToF characterization (a.i., absolute intensity). Ions with two charges (m2+) appear exactly at half the expected (m+) mass/charge (m/z) ratio.
Extended Data Figure 4 Phosphatase treatment of poly(I:C) affects RIG-I but not MDA5-dependent IFN-responses.
a, Schematic representation of inosinic acid or cytidylic acid homopolymer synthesis from inosine 5′-diphosphate or cytidine 5′-diphosphate through the action of polynucleotide phosphorylase, which when annealed form the synthetic dsRNA analogue poly(I:C). Whether the synthesized polynucleotides carry a 5′ di- or monophosphate or a mixture of both is unclear. b, IFN-pre-treated MDA5−/− or RIG-I−/− immortalized MEFs were transfected with poly(I:C) ± CIP. IFN induction was quantified 16 h later using an IFN-β promoter reporter assay. c, Poly(I:C) was first cleaved with RNase III for 1 or 5 min before being treated or not with CIP (+/−). Samples were subjected to gel electrophoresis to verify digestion (left panel) or transfected into IFN-pre-treated MDA5−/− MEFs (right panel). Cells were harvested 16 h post-transfection, and IFN-responses were assessed by RT-qPCR for ifit1 expression. Water and ppp-IVT-RNA99nt were included as controls. RE, relative expression. All experiments were performed at least twice; one representative experiment is shown. For PCR data, the mean ( ± s.d.) of triplicate technical replicates is shown.
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 and further text relating to Supplementary Figure 1. (PDF 444 kb)
Rights and permissions
About this article
Cite this article
Goubau, D., Schlee, M., Deddouche, S. et al. Antiviral immunity via RIG-I-mediated recognition of RNA bearing 5′-diphosphates. Nature 514, 372–375 (2014). https://doi.org/10.1038/nature13590
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13590
This article is cited by
-
Crosstalk between vault RNAs and innate immunity
Molecular Biology Reports (2024)
-
Global trends in research of melanoma differentiation-associated gene 5: a bibliometric analysis from 2002 to 2022
Clinical Rheumatology (2024)
-
Exploiting RIG-I-like receptor pathway for cancer immunotherapy
Journal of Hematology & Oncology (2023)
-
Mitochondrial E3 ligase MARCH5 is a safeguard against DNA-PKcs-mediated immune signaling in mitochondria-damaged cells
Cell Death & Disease (2023)
-
Novel insights into double-stranded RNA-mediated immunopathology
Nature Reviews Immunology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.