Abstract
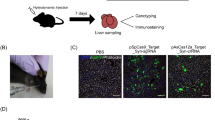
The study of cancer genes in mouse models has traditionally relied on genetically-engineered strains made via transgenesis or gene targeting in embryonic stem cells1. Here we describe a new method of cancer model generation using the CRISPR/Cas (clustered regularly interspaced short palindromic repeats/CRISPR-associated proteins) system in vivo in wild-type mice. We used hydrodynamic injection to deliver a CRISPR plasmid DNA expressing Cas9 and single guide RNAs (sgRNAs)2,3,4 to the liver that directly target the tumour suppressor genes Pten (ref. 5) and p53 (also known as TP53 and Trp53) (ref. 6), alone and in combination. CRISPR-mediated Pten mutation led to elevated Akt phosphorylation and lipid accumulation in hepatocytes, phenocopying the effects of deletion of the gene using Cre–LoxP technology7,8. Simultaneous targeting of Pten and p53 induced liver tumours that mimicked those caused by Cre–loxP-mediated deletion of Pten and p53. DNA sequencing of liver and tumour tissue revealed insertion or deletion mutations of the tumour suppressor genes, including bi-allelic mutations of both Pten and p53 in tumours. Furthermore, co-injection of Cas9 plasmids harbouring sgRNAs targeting the β-catenin gene and a single-stranded DNA oligonucleotide donor carrying activating point mutations led to the generation of hepatocytes with nuclear localization of β-catenin. This study demonstrates the feasibility of direct mutation of tumour suppressor genes and oncogenes in the liver using the CRISPR/Cas system, which presents a new avenue for rapid development of liver cancer models and functional genomics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
BioProject
Data deposits
Data generated during the work are deposited at NCBI BioProject under accession code PRJNA252101.
References
Van Dyke, T. & Jacks, T. Cancer modeling in the modern era: progress and challenges. Cell 108, 135–144 (2002)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013)
Song, M. S., Salmena, L. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor. Nature Rev. Mol. Cell Biol. 13, 283–296 (2013)
Feldser, D. M. et al. Stage-specific sensitivity to p53 restoration during lung cancer progression. Nature 468, 572–575 (2010)
Horie, Y. et al. Hepatocyte-specific Pten deficiency results in steatohepatitis and hepatocellular carcinomas. J. Clin. Invest. 113, 1774–1783 (2004)
Stiles, B. et al. Liver-specific deletion of negative regulator Pten results in fatty liver and insulin hypersensitivity. Proc. Natl Acad. Sci. USA 101, 2082–2087 (2004)
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nature Biotechnol. 31, 827–832 (2013)
Mali, P., Esvelt, K. M. & Church, G. M. Cas9 as a versatile tool for engineering biology. Nature Methods 10, 957–963 (2013)
Sander, J. D. & Joung, J. K. CRISPR-Cas systems for editing, regulating and targeting genomes. Nature Biotechnol. 32, 347–355 (2014)
Fellmann, C. & Lowe, S. W. Stable RNA interference rules for silencing. Nature Cell Biol. 16, 10–18 (2013)
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013)
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013)
Li, W., Teng, F., Li, T. & Zhou, Q. Simultaneous generation and germline transmission of multiple gene mutations in rat using CRISPR-Cas systems. Nature Biotechnol. 31, 684–686 (2013)
Li, D. et al. Heritable gene targeting in the mouse and rat using a CRISPR-Cas system. Nature Biotechnol. 31, 681–683 (2013)
Shen, B. et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res. 23, 720–723 (2013)
Wu, Y. et al. Correction of a genetic disease in mouse via use of CRISPR-Cas9. Cell Stem Cell 13, 659–662 (2013)
Niu, Y. et al. Generation of gene-modified cynomolgus monkey via Cas9/RNA-mediated gene targeting in one-cell embryos. Cell 156, 836–843 (2014)
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nature Biotechnol. 32, 551–553 (2014)
Liu, F., Song, Y. & Liu, D. Hydrodynamics-based transfection in animals by systemic administration of plasmid DNA. Gene Ther. 6, 1258–1266 (1999)
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nature Biotechnol. 31, 822–826 (2013)
Mali, P. et al. Cas9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nature Biotechnol. 31, 833–838 (2013)
Ran, F. A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013)
Ong, C. K. et al. Exome sequencing of liver fluke-associated cholangiocarcinoma. Nature Genet. 44, 690–693 (2012)
Malina, A. et al. Repurposing CRISPR/Cas9 for in situ functional assays. Genes Dev. 27, 2602–2614 (2013)
Katz, S. F. et al. Disruption of Trp53 in livers of mice induces formation of carcinomas with bilineal differentiation. Gastroenterology 142, 1229–1239 (2012)
Moon, R. T., Kohn, A. D., De Ferrari, G. V. & Kaykas, A. Wnt and β-catenin signalling: diseases and therapies. Nature Rev. Genet. 5, 691–701 (2004)
Tward, A. D. et al. Distinct pathways of genomic progression to benign and malignant tumors of the liver. Proc. Natl Acad. Sci. USA 104, 14771–14776 (2007)
Xue, W. et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 445, 656–660 (2007)
Zender, L. et al. Identification and validation of oncogenes in liver cancer using an integrative oncogenomic approach. Cell 125, 1253–1267 (2006)
Xue, W. et al. Response and resistance to NF-κB inhibitors in mouse models of lung adenocarcinoma. Cancer Discov. 1, 236–247 (2011)
Chen, S. et al. Global microRNA depletion suppresses tumor angiogenesis. Genes Dev. 28, 1054–1067 (2014)
Acknowledgements
We thank D. McFadden, N. Dimitrova, E. Snyder, A. Farago, M. Muzumdar, F. Sanchez-Rivera, J. Doench, L. Cong and S. Levine for discussions and for sharing reagents. We thank the Koch Institute Swanson Biotechnology Center (SBC) for technical support, specifically the Hope Babette Tang (1983) Histology Facility and K. Cormier. This work was supported by grants 2-PO1-CA42063, RO1-EB000244, RO1-CA115527 and RO1-CA132091 from the National Institutes of Health and supported in part by Cancer Center Support (core) grant P30-CA14051 from the National Cancer Institute. This work was supported, in part, by NIH Grant R01-CA133404 and Casimir-Lambert Fund to P.A.S. H.Y. is supported by 5-U54-CA151884-04 NIH Centers for Cancer Nanotechnology Excellence and the Harvard-MIT Center of Cancer Nanotechnology Excellence. S.C. is a Damon Runyon Fellow (DRG-2117-12). W.X. was supported by fellowships from the American Association for Cancer Research and the Leukemia Lymphoma Society and is currently supported by grant 1K99CA169512. T.J. is a Howard Hughes Medical Institute (HHMI) Investigator, the David H. Koch Professor of Biology, and a Daniel K. Ludwig Scholar.
Author information
Authors and Affiliations
Contributions
W.X., S.C., H.Y. and T.J. designed the study. W.X., S.C., H.Y., T.T., W.C. and G.Y. performed experiments and analysed data. D.G.C. and R.B. performed histology and evaluations. T.P., N.S.J., F.Z. and D.A.G. provided reagents and conceptual advice. W.X., S.C., H.Y., P.A.S. and T.J. wrote the manuscript with comments from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Representative Pten indels in sgPten-treated 3T3 cells.
Mouse 3T3 cells were co-transfected with sgPten and a GFP plasmid. The highest 20% of GFP-positive cells were sorted to enrich for cells expressing sgPten. Deep sequencing of the Pten locus revealed 36.4% Pten indels in this context (Supplementary Table 5), presumably due to the more efficient delivery of sgPten via cell culture transfection and sorting. Red arrowheads denote predicted Cas9 cutting sites. Black or purple bars in grey sequencing reads indicate deletions or insertions, respectively. Other colours indicate SNPs. a, Pten PCR region. b, Zoom in view. n = 1 DNA sample. c, Distribution of Pten indel length.
Extended Data Figure 2 CRISPR generates Pten-negative hepatocytes in vivo.
a, b, Low-magnification images of Pten IHC in sgGFP- (a) and sgPten-treated (b) mice. Scale bar is 100 μm. c, d, IHC on serial sections from sgPten-treated mice. Black arrows denote cells with negative Pten staining and positive pAkt staining. White arrowhead denotes cells with intermediate Pten staining, potentially indicating heterozygous Pten mutation or multi-nucleated hepatocytes with partial Pten loss. Insets show high-magnification IHC images. Scale bar is 100 μm. n = 5 mice. The frequency of Pten-deficient cells is probably a reflection of the transduction efficiency following hydrodynamic injection and the time required to achieve mutation. A recent study by our groups has shown that ∼17% of hepatocytes were Flag–Cas9 positive as indicated by IHC 24 h after hydrodynamic injection, only 1.4% of cells on day 7, and less than 0.3% at one month20. Given that Cas9-mediated genome editing usually takes more than 48 h (ref. 2), the fraction of hepatocytes that productively express Cas9 and an sgRNA after hydrodynamic injection is estimated to be less than 17%.
Extended Data Figure 3 sgPten induces lipid accumulation in the liver.
FVB mice were injected with sgGFP or sgPten (n = 5). 2 months later, liver sections were stained for Oil Red O, a marker for lipid accumulation. Scale bars are 50 μm.
Extended Data Figure 4 sgPten generated indels at the Pten locus in the liver.
a, Representative indel frequency. Base pair position denotes position along the Pten reference sequence. b, Representative Pten indel frequency in sgGFP mice. Note the low mutant allele frequency compared to a. sgPten samples show indels peaking at the predicted Cas9 cutting site whereas sgGFP indels distribute randomly. c, Representative indel frequency in Cas9D10A + sgPten.2/3-treated mice. For all panels */+x denotes insertion of x nucleotides, */−x denotes deletion of x nucleotides.
Extended Data Figure 6 Assessing off-target cutting of sgPten.
a, Top 30 potential off-target sites for sgPten in the mouse genome. Score is likelihood of off-target binding. Only site 4 is in the exon region of NR_045386, which is a long non-coding RNA. b, Surveyor assay in sgGFP (−) and sgPten (+) treated liver genomic DNA. Pten and Pten off-targets sites 1, 2, 3 and 4 were PCR amplified. Predicted size of uncut and cut bands are indicated. Red arrowheads indicate Surveyor-nuclease-cleaved Pten PCR products. The data are representative of two independent liver samples.
Extended Data Figure 7 Representative p53 indels in sgp53 treated 3T3 cells.
Red arrowheads denote predicted Cas9 cutting sites. Black or purple bars in grey sequencing reads indicate deletions or insertions, respectively. a, p53 PCR region. b, Zoom in view. n = 1 DNA sample.
Extended Data Figure 8 Analysing sgp53-treated livers.
a, Histology of sgp53-treated livers. Scale bars, 50 μm, n = 3 mice. b, p53 indel frequency was measured by MiSeq at day 14. Error bars are s.d., n = 2 mice.
Extended Data Figure 9 sgPten- and sgp53-generated indels in the liver.
sgPten and sgp53 were co-injected into FVB mice. Representative analysis of MiSeq is shown. n = 2 mice. a–c, Pten locus. d–f, p53 locus. a, d, Indel frequency. */+ indicates insertions and */– indicates deletions. Base pair position denotes position along the Pten or p53 reference sequences. Arrowheads denote predicted Cas9 cutting sites. b, e, Distribution of indel length. c, f, Distribution of indel frame phase. Frame phase of indels was calculated as the length of indels modulus 3.
Extended Data Figure 10 CRISPR introduces β-catenin mutations in the liver.
a, Low-magnification images of glutamine synthetase (GS) IHC as in Fig. 4c. b, Frequency of Ctnnb1 deep sequencing reads with all four G nucleotides. The rate of β-catenin donor integration was calculated as donor allele frequency. n = 2 mice.
Supplementary information
TITSupplementary InformationLE
This file contains Supplementary Tables 1-3, legends for Supplementary Tables 4-8 (see separate excel files) and Supplementary Sequences. (PDF 206 kb)
Supplementary Table 4
This file shows deep sequencing data for Pten and p53 indels in the liver. (XLSX 446 kb)
Supplementary Table 5
This file shows deep sequencing data for Pten and p53 indels in 3T3 cells. (XLSX 48 kb)
Supplementary Table 6
This file shows deep sequencing data for sgPten off-target analysis. (XLSX 18 kb)
Supplementary Table 7
This file shows sequences of Pten and p53 alleles in sgPten+sgp53 liver tumors. (XLSX 17 kb)
Supplementary Table 8
This file shows deep sequencing data for β-Catenin indels in the liver. (XLSX 89 kb)
Rights and permissions
About this article
Cite this article
Xue, W., Chen, S., Yin, H. et al. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature 514, 380–384 (2014). https://doi.org/10.1038/nature13589
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13589
This article is cited by
-
Doxycycline-dependent Cas9-expressing pig resources for conditional in vivo gene nullification and activation
Genome Biology (2023)
-
A potential paradigm in CRISPR/Cas systems delivery: at the crossroad of microalgal gene editing and algal-mediated nanoparticles
Journal of Nanobiotechnology (2023)
-
An overview of mouse models of hepatocellular carcinoma
Infectious Agents and Cancer (2023)
-
Small extrachromosomal circular DNA harboring targeted tumor suppressor gene mutations supports intratumor heterogeneity in mouse liver cancer induced by multiplexed CRISPR/Cas9
Genome Medicine (2023)
-
Activation of melanocortin-1 receptor signaling in melanoma cells impairs T cell infiltration to dampen antitumor immunity
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.