Abstract
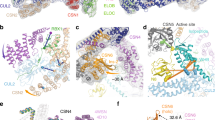
Ubiquitination is a crucial cellular signalling process, and is controlled on multiple levels. Cullin–RING E3 ubiquitin ligases (CRLs) are regulated by the eight-subunit COP9 signalosome (CSN). CSN inactivates CRLs by removing their covalently attached activator, NEDD8. NEDD8 cleavage by CSN is catalysed by CSN5, a Zn2+-dependent isopeptidase that is inactive in isolation. Here we present the crystal structure of the entire ∼350-kDa human CSN holoenzyme at 3.8 Å resolution, detailing the molecular architecture of the complex. CSN has two organizational centres: a horseshoe-shaped ring created by its six proteasome lid–CSN–initiation factor 3 (PCI) domain proteins, and a large bundle formed by the carboxy-terminal α-helices of every subunit. CSN5 and its dimerization partner, CSN6, are intricately embedded at the core of the helical bundle. In the substrate-free holoenzyme, CSN5 is autoinhibited, which precludes access to the active site. We find that neddylated CRL binding to CSN is sensed by CSN4, and communicated to CSN5 with the assistance of CSN6, resulting in activation of the deneddylase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Wei, N., Chamovitz, D. A. & Deng, X. W. Arabidopsis COP9 is a component of a novel signaling complex mediating light control of development. Cell 78, 117–124 (1994)
Wei, N. & Deng, X. W. The COP9 signalosome. Annu. Rev. Cell Dev. Biol. 19, 261–286 (2003)
Lyapina, S. et al. Promotion of NEDD-CUL1 conjugate cleavage by COP9 signalosome. Science 292, 1382–1385 (2001)
Schwechheimer, C. et al. Interactions of the COP9 signalosome with the E3 ubiquitin ligase SCFTIR1 in mediating auxin response. Science 292, 1379–1382 (2001)
Zhou, C. et al. The fission yeast COP9/signalosome is involved in cullin modification by ubiquitin-related Ned8p. BMC Biochem. 2, 7 (2001)
Lydeard, J. R., Schulman, B. A. & Harper, J. W. Building and remodelling Cullin–RING E3 ubiquitin ligases. EMBO Rep. 14, 1050–1061 (2013)
Soucy, T. A. et al. An inhibitor of NEDD8-activating enzyme as a new approach to treat cancer. Nature 458, 732–736 (2009)
Cope, G. A. et al. Role of predicted metalloprotease motif of Jab1/Csn5 in cleavage of Nedd8 from Cul1. Science 298, 608–611 (2002)
Fischer, E. S. et al. The molecular basis of CRL4DDB2/CSA ubiquitin ligase architecture, targeting, and activation. Cell 147, 1024–1039 (2011)
Enchev, R. I. et al. Structural basis for a reciprocal regulation between SCF and CSN. Cell Rep. 2, 616–627 (2012)
Emberley, E. D., Mosadeghi, R. & Deshaies, R. J. Deconjugation of Nedd8 from Cul1 is directly regulated by Skp1-F-box and substrate, and the COP9 signalosome inhibits deneddylated SCF by a noncatalytic mechanism. J. Biol. Chem. 287, 29679–29689 (2012)
Zimmerman, E. S., Schulman, B. A. & Zheng, N. Structural assembly of cullin-RING ubiquitin ligase complexes. Curr. Opin. Struct. Biol. 20, 714–721 (2010)
Furukawa, M., Zhang, Y., McCarville, J., Ohta, T. & Xiong, Y. The CUL1 C-terminal sequence and ROC1 are required for efficient nuclear accumulation, NEDD8 modification, and ubiquitin ligase activity of CUL1. Mol. Cell. Biol. 20, 8185–8197 (2000)
Podust, V. N. et al. A Nedd8 conjugation pathway is essential for proteolytic targeting of p27Kip1 by ubiquitination. Proc. Natl Acad. Sci. USA 97, 4579–4584 (2000)
Read, M. A. et al. Nedd8 modification of Cul-1 activates SCFβTrCP-dependent ubiquitination of IκBα. Mol. Cell. Biol. 20, 2326–2333 (2000)
Wu, K., Chen, A. & Pan, Z. Q. Conjugation of Nedd8 to CUL1 enhances the ability of the ROC1–CUL1 complex to promote ubiquitin polymerization. J. Biol. Chem. 275, 32317–32324 (2000)
Morimoto, M., Nishida, T., Honda, R. & Yasuda, H. Modification of cullin-1 by ubiquitin-like protein Nedd8 enhances the activity of SCFskp2 toward p27kip1. Biochem. Biophys. Res. Commun. 270, 1093–1096 (2000)
Sharon, M. et al. Symmetrical modularity of the COP9 signalosome complex suggests its multifunctionality. Structure 17, 31–40 (2009)
Wei, N., Serino, G. & Deng, X. W. The COP9 signalosome: more than a protease. Trends Biochem. Sci. 33, 592–600 (2008)
Zhao, R. et al. Subunit 6 of the COP9 signalosome promotes tumorigenesis in mice through stabilization of MDM2 and is upregulated in human cancers. J. Clin. Invest. 121, 851–865 (2011)
Lee, M. H., Zhao, R., Phan, L. & Yeung, S. C. Roles of COP9 signalosome in cancer. Cell Cycle 10, 3057–3066 (2011)
Echalier, A. et al. Insights into the regulation of the human COP9 signalosome catalytic subunit, CSN5/Jab1. Proc. Natl Acad. Sci. USA 110, 1273–1278 (2013)
Enchev, R. I., Schreiber, A., Beuron, F. & Morris, E. P. Structural insights into the COP9 signalosome and its common architecture with the 26S proteasome lid and eIF3. Structure 18, 518–527 (2010)
Beck, F. et al. Near-atomic resolution structural model of the yeast 26S proteasome. Proc. Natl Acad. Sci. USA 109, 14870–14875 (2012)
Lander, G. C. et al. Complete subunit architecture of the proteasome regulatory particle. Nature 482, 186–191 (2012)
Lasker, K. et al. Molecular architecture of the 26S proteasome holocomplex determined by an integrative approach. Proc. Natl Acad. Sci. USA 109, 1380–1387 (2012)
da Fonseca, P. C., He, J. & Morris, E. P. Molecular model of the human 26S proteasome. Mol. Cell 46, 54–66 (2012)
Matyskiela, M. E., Lander, G. C. & Martin, A. Conformational switching of the 26S proteasome enables substrate degradation. Nature Struct. Mol. Biol. 20, 781–788 (2013)
Siridechadilok, B., Fraser, C. S., Hall, R. J., Doudna, J. A. & Nogales, E. Structural roles for human translation factor eIF3 in initiation of protein synthesis. Science 310, 1513–1515 (2005)
Sun, C. et al. Functional reconstitution of human eukaryotic translation initiation factor 3 (eIF3). Proc. Natl Acad. Sci. USA 108, 20473–20478 (2011)
Hashem, Y. et al. Structure of the mammalian ribosomal 43S preinitiation complex bound to the scanning factor DHX29. Cell 153, 1108–1119 (2013)
Querol-Audi, J. et al. Architecture of human translation initiation factor 3. Structure 21, 920–928 (2013)
Wei, Z. et al. Crystal structure of human eIF3k, the first structure of eIF3 subunits. J. Biol. Chem. 279, 34983–34990 (2004)
Dessau, M. et al. The Arabidopsis COP9 signalosome subunit 7 is a model PCI domain protein with subdomains involved in COP9 signalosome assembly. Plant Cell 20, 2815–2834 (2008)
Pathare, G. R. et al. The proteasomal subunit Rpn6 is a molecular clamp holding the core and regulatory subcomplexes together. Proc. Natl Acad. Sci. USA 109, 149–154 (2012)
Zhang, H. et al. The crystal structure of the MPN domain from the COP9 signalosome subunit CSN6. FEBS Lett. 586, 1147–1153 (2012)
Boehringer, J. et al. Structural and functional characterization of Rpn12 identifies residues required for Rpn10 proteasome incorporation. Biochem. J. 448, 55–65 (2012)
Lee, J. H. et al. Crystal structure and versatile functional roles of the COP9 signalosome subunit 1. Proc. Natl Acad. Sci. USA 110, 11845–11850 (2013)
Khoshnevis, S. et al. Structural integrity of the PCI domain of eIF3a/TIF32 is required for mRNA recruitment to the 43S pre-initiation complexes. Nucleic Acids Res. 42, 4123–4139 (2014)
Worden, E. J., Padovani, C. & Martin, A. Structure of the Rpn11–Rpn8 dimer reveals mechanisms of substrate deubiquitination during proteasomal degradation. Nature Struct. Mol. Biol. 21, 220–227 (2014)
Pathare, G. R. et al. Crystal structure of the proteasomal deubiquitylation module Rpn8-Rpn11. Proc. Natl Acad. Sci. USA 111, 2984–2989 (2014)
Hofmann, K. & Bucher, P. The PCI domain: a common theme in three multiprotein complexes. Trends Biochem. Sci. 23, 204–205 (1998)
Ellisdon, A. M. & Stewart, M. Structural biology of the PCI-protein fold. BioArchitecture 2, 118–123 (2012)
Tran, H. J., Allen, M. D., Lowe, J. & Bycroft, M. Structure of the Jab1/MPN domain and its implications for proteasome function. Biochemistry 42, 11460–11465 (2003)
Ambroggio, X. I., Rees, D. C. & Deshaies, R. J. JAMM: a metalloprotease-like zinc site in the proteasome and signalosome. PLoS Biol. 2, e2 (2004)
Pick, E. et al. The minimal deneddylase core of the COP9 signalosome excludes the Csn6 MPN− domain. PLoS ONE 7, e43980 (2012)
Sato, Y. et al. Structural basis for specific cleavage of Lys 63-linked polyubiquitin chains. Nature 455, 358–362 (2008)
Duda, D. M. et al. Structural insights into NEDD8 activation of cullin-RING ligases: conformational control of conjugation. Cell 134, 995–1006 (2008)
Zheng, N. et al. Structure of the Cul1–Rbx1–Skp1–F boxSkp2 SCF ubiquitin ligase complex. Nature 416, 703–709 (2002)
Hao, B. et al. Structural basis of the Cks1-dependent recognition of p27Kip1 by the SCFSkp2 ubiquitin ligase. Mol. Cell 20, 9–19 (2005)
Cronin, C. N., Lim, K. B. & Rogers, J. Production of selenomethionyl-derivatized proteins in baculovirus-infected insect cells. Protein Sci. 16, 2023–2029 (2007)
Tsuge, T., Matsui, M. & Wei, N. The subunit 1 of the COP9 signalosome suppresses gene expression through its N-terminal domain and incorporates into the complex through the PCI domain. J. Mol. Biol. 305, 1–9 (2001)
Menon, S., Rubio, V., Wang, X., Deng, X. W. & Wei, N. Purification of the COP9 signalosome from porcine spleen, human cell lines, and Arabidopsis thaliana plants. Methods Enzymol. 398, 468–481 (2005)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Abrahams, J. P. & Leslie, A. G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D 52, 30–42 (1996)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. BUSTER Version 2.11.5 (Global Phasing, 2011)
Smart, O. S. et al. Exploiting structure similarity in refinement: automated NCS and target-structure restraints in BUSTER. Acta Crystallogr. D 68, 368–380 (2012)
Schröder, G. F., Levitt, M. & Brunger, A. T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011)
Nicholls, R. A., Long, F. & Murshudov, G. N. Low-resolution refinement tools in REFMAC5. Acta Crystallogr. D 68, 404–417 (2012)
Karplus, P. A. & Diederichs, K. Linking crystallographic model and data quality. Science 336, 1030–1033 (2012)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Marks, B. D. et al. Multiparameter analysis of a screen for progesterone receptor ligands: comparing fluorescence lifetime and fluorescence polarization measurements. Assay Drug Dev. Technol. 3, 613–622 (2005)
Estrin, E., Lopez-Blanco, J. R., Chacon, P. & Martin, A. Formation of an intricate helical bundle dictates the assembly of the 26S proteasome lid. Structure 21, 1624–1635 (2013)
Guevara, T. et al. Proenzyme structure and activation of astacin metallopeptidase. J. Biol. Chem. 285, 13958–13965 (2010)
Bode, W., Gomis-Ruth, F. X., Huber, R., Zwilling, R. & Stocker, W. Structure of astacin and implications for activation of astacins and zinc-ligation of collagenases. Nature 358, 164–167 (1992)
Pei, J., Kim, B. H. & Grishin, N. V. PROMALS3D: a tool for multiple protein sequence and structure alignments. Nucleic Acids Res. 36, 2295–2300 (2008)
Acknowledgements
This work was supported by the Novartis Research Foundation, the Swiss National Science Foundation (31003A_144020) and the European Research Council (MoBa-CS #260481). G.M.L. is the recipient of a Novartis (NIBR) presidential postdoctoral fellowship. Part of this work was performed at beamline X06DA and X10SA of the Swiss Light Source. We thank H. Wu for sharing the A. thaliana CSN1 coordinates before publication; H. Gut, J. Keusch and M. Jones for support; and C. Vonrhein and the BUSTER development group, R. Read, A. McCoy, T. Terwilliger, R. Nicholls, R. Kingston and K. Diederichs for technical advice. We are also grateful for the assistance of B. Martoglio, R. Assenberg, C. Logel and I. Bechtold in developing the PT22 CSN assays.
Author information
Authors and Affiliations
Contributions
G.M.L., M.R. and N.H.T. initiated the project. G.M.L. established the purification methods for CSN and CSN4, produced most proteins, performed the gel-based assays, and obtained the crystals. G.M.L. improved the crystals with input from R.D.B. G.M.L. and R.D.B. collected the diffraction data. R.D.B. carried out the crystallographic analyses and interpreted the results. S.C. performed electron microscopy analysis. D.H. carried out protein analysis. E.S.F. developed the binding and activity assays with input from U.H.; E.S.F. performed the assays and analysed the results. N.H.T. supervised all aspects of the project and analysed the results. R.D.B. and N.H.T. wrote the manuscript with important contributions from G.M.L., M.R., E.S.F. and S.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Protein complexes used in this study.
a–d, f, j, k, SDS–PAGE analysis of CSN variants used in the study. g, The N8CRL4ADDB2 substrate used for enzymatic measurements. a, The CSN5 E76A mutant catalytically inactivates CSN: activity of CSN (full-length) and CSN (CSN5(E76A)), a holocomplex carrying the active site mutant, was determined by fluorescence polarization measurements using a substrate with a PT22-labelled N8CRL4ADDB2. The decrease in signal for CSN (CSN5(E76A)) and in the buffer control is due to fluorophore photobleaching. Determination of steady-state kinetics using N8CRL4ADDB2 as substrate: initial rates observed following incubation of CSN or mutants thereof, with increasing concentrations of PT22-labelled N8CRL4ADDB2 substrate, as indicated on the abscissa. The fit of the observed initial velocities to the Michaelis–Menten equation is shown as a red line in b, endogenous CSN purified from HEK293T cells (CSN (HEK293T)); c, recombinant full-length CSN (CSN (full-length)); d, CSN with the boundaries used for crystallization (CSN); e, CSN (CSN6ΔMPN), lacking the CSN6 MPN domain (comprising residues 192–327); and f, CSN (CSN6Δloop), lacking residues 174–179 of the CSN6 Ins-2 loop. a–f, Data are the average of three technical replicates. c, e, f, Error bars show standard error of the mean (s.e.m.). l, Table of steady-state kinetic parameters (errors show ± s.e.m.). A previous study46 described a CSN variant where the mouse CSN6 MPN domain was deleted (retaining CSN6 residues 171–324). This construct when immunoprecipitated from cells with the remainder of CSN was found to be active. For human CSN (CSN6ΔMPN), detailed quantitative kinetic analysis revealed a ∼100-fold reduction in kcat compared with CSN (full-length). Interestingly, mouse CSN6 171–324 retains the residues involved in the CSN4–CSN6 interface including the Ins-2 loop. Determination of steady-state kinetics using ubiquitin-rhodamine 110 as substrate: CSN variants were assayed for proteolytic release of the rhodamine 110 fluorescent group from the C-terminal glycine of ubiquitin (ubiquitin-rhodamine), using fluorescence quenching as readout. h, Wild-type CSN was not sufficiently active on ubiquitin-rhodamine to determine Michaelis–Menten parameters. To benchmark wild-type CSN against CSN (CSN5(E104A)), we assessed the relative rates at a fixed concentration of 0.5 µM ubiquitin-rhodamine substrate and 1 nM CSN (CSN5(E76A)), CSN (full-length) (23.0 ± 2.9 fmol s−1) and CSN (CSN5(E104A)) (137.8 ± 2.2 fmol s−1), using the CSN (CSN5(E76A)) active site mutant as a control. i, j, k, Increasing concentrations of ubiquitin-rhodamine with CSN (CSN6Δloop) (i), CSN (CSN5(E104A)) (j) and CSN (CSN6Δloop, CSN5(E104A)) (k) double mutant were assayed. Fit of the initial velocities to the Michaelis–Menten equation is shown as a red line. h–k, Data are the average of three technical replicates. m, Table summarizing the activity of CSN variants on ubiquitin-rhodamine (errors show ± s.e.m.). Assayed protein concentrations and Vmax values are indicated.
Extended Data Figure 2 CSN structure solution and validation.
a, Table of differentially substituted SeMet crystals used for CSN structure determination. b, The final experimentally determined heavy atom positions for one CSN complex in the ASU are shown as spheres and coloured differently for each data set. The Zn2+ ion, which is found in the LLG maps calculated for all CSN data sets, is labelled. The models of CSN1 and CSN4, which are relatively poorly described by heavy atoms, were based on similar high-resolution structures. c, To avoid potential complications due to twinning in the 3.8 Å resolution data set (c343), the final model was also validated against the untwinned 4.1 Å (c318) data set. A full 2mFo − DFc simulated annealing composite omit map calculated for c318 by CNS and displayed at a contour level of 1 r.m.s.d. with a radius of 3 Å about the model. The map coefficients were sharpened by a B-factor of −80 Å2. d, Portion of the 2mFo − DFc electron density calculated for c318 surrounding the CSN4 and CSN2 WH subdomains (coloured sticks) sharpened by a B-factor of −80 Å2 and contoured at 1.5 r.m.s.d. at a radius of 3 Å about the displayed model. e–g, Close-up of the electron density for the CSN5 active site in the composite omit map shown in c; e, overlaid with an anomalous LLG map (pink mesh) calculated for c318 and contoured at 6 r.m.s.d., indicating the position of the zinc ion (peak height 10.3 r.m.s.d.); f, shown with a ribbon diagram (cyan) of the final model with the Zn2+ ion as a sphere (mauve); and g, the same as f, with sticks and two active site residues labelled.
Extended Data Figure 3 The overall fold of each CSN protein.
a–h, Cartoon representation of the individual subunits as found in CSN: CSN1 (a); CSN2 (b); CSN3 (c); CSN4 (d); CSN7a (e); CSN8 (f); CSN5 (g); and CSN6 (h). C-terminal helices (dark grey) are numbered in roman numerals. The WH subdomain of the PCI domain is coloured light grey. i–l, Structural comparisons with previously reported CSN structures. Superposition of CSN1 (red) and A. thaliana CSN1 (grey) (PDB accession 4LCT38; r.m.s.d. 0.8 Å (296 residues) with 50% sequence identity) (i); CSN7 (blue) and A. thaliana CSN7 (grey) (PDB accession 3CHM34; r.m.s.d. 1.4 Å (151 residues) with 36% sequence identity) (j); CSN5 (cyan) and human CSN5 (grey) (PDB accession 4F7O22; r.m.s.d. 2.2 Å (168 residues) with 100% sequence identity) (k); and CSN6 (orange) and D. melanogaster CSN6 (grey) (PDB accession 4E0Q36; r.m.s.d. 1.9 Å (87 residues) with 56% sequence identity) (l).
Extended Data Figure 4 Fit of the CSN model to reported electron microscopy maps of CSN, the 19S lid, and eIF3.
a, Isolated CSN (Electron Microscopy Data Bank (EMDB) accession 2176)10. The electron microscopy maps are represented as surfaces and CSN is styled in cylindrical cartoon. We note that CSN electron microscopy maps (EMDB accessions 1700 (ref. 23) and 2176) are largely inconsistent with each other (cross-correlation between the maps is ∼0.36). b, Table of CSN paralogues in 19S lid and eIF3. The CSN crystal structure explains features of the 19S lid and eIF3 electron microscopy maps. c, d, CSN fit to the isolated 19S proteasome lid25 (EMDB accession 1994) (c); and CSN fit to the map of the 19S lid bound to the proteasome in the absence of substrate24 (EMDB accession 2165) (d). The electron microscopy maps of 19S lid are represented as surface (light pink), with the base and the 20S proteasome shown in grey. e, Superposition of the CSN5 (cyan)–CSN6 (orange) MPN dimer with the RPN11 (green)–RPN8 (purple) MPN dimer from the 19S lid (PDB accession 4O8X40). The structures superpose well with an r.m.s.d. of 1.8 Å (719 residues). f, Structural comparison of the CSN helical bundle (coloured as in c) with the predicted model of the helical bundle in the 19S lid (grey) (PDB accession 3J47)69. g, h, CSN fit into the maps of isolated eIF3 complex (yellow)32 (EMDB accession 2166) (g); and to the eIF3 segment of the 43S preinitiation complex map31 (EMDB accession 5658) (h).
Extended Data Figure 5 Details of the PCI ring interactions and PCI ring capping features.
a, Location of the composite β-sheet within CSN. b, c, Close-up views of the interface between the CSN1–CSN3 (b) and the CSN2–CSN4 WH subdomains (c). d, CSN8 carries a helix-turn-β2 motif in the WH subdomain that is reduced by six residues (magenta), retracting the available interaction interface for further PCI interactions (see Supplementary Data). e, Superposition of PCI subunits, highlighting the position of conserved aromatic residues (Tyr/Phe shown as sticks) interconnecting PCI subunits. CSN7 lacks an equivalent residue. Interactions between subunits within CSN: f, g, Interactions between the CSN3 and CSN8 helical repeats (f); and the CSN7 helical repeats and the proximal end of the C-terminal helical bundle (g).
Extended Data Figure 6 The role of the PCI domain and the C-terminal helix in CSN assembly.
a–e, SDS–PAGE analysis of the effects of C-terminal helix and/or PCI domain deletions in CSN subunits: CSN1 (a); CSN2 (b); CSN3 (c); CSN4 (d); and CSN8 (e) on their ability to incorporate into CSN. All complexes contained one Strep(II)-tagged subunit (as indicated) and His6-tags on the other subunits. The cell lysate was divided and subjected to parallel Strep(II)-tag pull-downs using Strep-Tactin beads to probe complex integrity (top) and His6-tag pull-downs using Ni-NTA beads for expression controls (bottom). CSN1 (a) and CSN4 (d) are dependent on the presence of their C-terminal helix (CSN1 isoform 2 residues 466–527; and CSN4 364–406) for integration into CSN. The deletion of C-terminal helix of CSN3 (residues 364–423) impaired its ability to incorporate into the holoenzyme (c). For CSN2 (b) and CSN8 (e), deletion of their C-terminal helices (CSN2 residues 417–443 and CSN8 residues 159–209/166–209) had no effect on complex incorporation and integrity. f, SDS–PAGE analysis showing the inability of C-terminally truncated CSN5 to incorporate into CSN. The co-expressed complexes bore a Strep(II)-tag on CSN6 and His6-tags on the other subunits. g, The CSN (CSN (CSN6ΔMPN)) complex pulled down by the Strep(II)-tagged C-terminal fragment of CSN6 (residues 192–327) was stoichiometric. Arrows indicate truncated subunits. Topological knot between CSN5 and CSN6: h, CSN5 and CSN6 are tightly interlinked. Cartoon representation of the topological knot made by CSN5 (cyan) and CSN6 (orange) immediately before entering the helical bundle. The CSN6 loop (residues 208–214) that crosses CSN5 was disordered in most crystals and is not included in the CSN model.
Extended Data Figure 7 CSN5 integration into a preassembled seven-subunit CSN complex.
a, A preassembled seven-subunit CSN complex lacking CSN5 with a Strep(II)-tag on CSN6 and His6-tags on the other subunits (CSN 7mer) was incubated with His6-tagged CSN5 for 30 min, followed by pull-down using Strep-Tactin beads. CSN5 enters the seven-subunit CSN, reconstituting the complex. b, CSN5 incorporation over time measured by the catalytic activity of the reconstituted complex. 2 nM seven-subunit CSN was mixed with the indicated concentrations of CSN5 and incubated for 24 h at 4 °C. Catalytic activity was assessed using a fluorescence polarization-based assay with a PT22-labelled N8CRL4ADDB2 substrate. CSN5 enters the preassembled seven-subunit complex in a concentration-dependent manner. CSN5 relies on the other seven holoenzyme subunits for efficient integration: c, We examined the effect individual subunits have on the integration of CSN5 into CSN by pull-down assay. Complexes contained a Strep(II)-tag on CSN6 and His6-tags on the other subunits. When CSN6 was omitted, the Strep(II)-tag was placed on CSN8. Full-length CSN subunits were pulled-down from the same lysate using Strep-Tactin beads (top), and Ni-NTA beads (bottom). d, In-gel quantification of CSN5 incorporation into CSN by Strep-Tactin pull-down normalized against CSN1 or, if absent, relative to CSN4 normalized against CSN1 in the control lane.
Extended Data Figure 8 CSN5 autoinhibition in the CSN holoenzyme and isolated CSN5.
a, Cartoon representation of the active site of isolated CSN5 (PDB accession 4F7O)22. b, A cullin Lys-NEDD8 isopeptide conjugate modelled in the isolated CSN5 active site (see Fig. 4d). c, Superposition of the CSN5 Ins-1 loop conformations in CSN (cyan) and isolated CSN5 (cream). d, Active site of the zinc metalloprotease, pro-astacin (PDB accession 3LQ0)70, showing the zinc coordinating and active site residues (green), and the autoinhibitory residue, Asp 21, from its pro-region in sticks (orange). e, Removal of the pro-region bearing Asp 21 produces catalytically active astacin (PDB accession 1AST71). f, Structure-based sequence alignment (PROMALS3D72) of the JAMM motif region in CSN5 and its 19S lid paralogue, RPN11.
Extended Data Figure 9 Crystal structure of isolated CSN4 (residues 1–363) and model of CRL binding-induced conformational remodelling in CSN.
a, Overall fold of isolated CSN4 shown as cartoon (mauve) with the chain termini and the WH subdomain (light grey) labelled. The second orientation highlights the hinge loop (residues 291–298) (red) and indicates the conformationally variable N-terminal region (mauve background). b, CSN4 conformers in the CSN holoenzyme (purple) and the structure of isolated CSN4 (mauve), which matches the SCF-bound conformation of CSN4 (Extended Data Fig. 9a). c, CSN5 is autoinhibited in the context of the holoenzyme by Glu 104 in the Ins-1 loop (red sphere), which coordinates to the catalytic Zn2+ ion (closed lock). Binding of the neddylated SCFSKP2/CKS1 cullin RING ligase pulls the CSN4 N-terminal domains towards CUL1/RBX1. The CSN5–CSN6 dimer undergoes conformational rearrangement resulting in CSN5 activation (open lock). CSN5 activation results in proteolytic removal of NEDD8 from SCFSKP2/CKS1.
Supplementary information
Supplementary Information
This file contains Supplementary Data, a Supplementary Discussion and Supplementary References. (PDF 250 kb)
Rights and permissions
About this article
Cite this article
Lingaraju, G., Bunker, R., Cavadini, S. et al. Crystal structure of the human COP9 signalosome. Nature 512, 161–165 (2014). https://doi.org/10.1038/nature13566
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13566
This article is cited by
-
Dynamic molecular architecture and substrate recruitment of cullin3–RING E3 ligase CRL3KBTBD2
Nature Structural & Molecular Biology (2024)
-
Regulatory mechanisms and therapeutic potential of JAB1 in neurological development and disorders
Molecular Medicine (2023)
-
The COP9 signalosome reduces neuroinflammation and attenuates ischemic neuronal stress in organotypic brain slice culture model
Cellular and Molecular Life Sciences (2023)
-
PKD phosphorylation and COP9/Signalosome modulate intracellular Spry2 protein stability
Oncogenesis (2023)
-
Kelch-like protein 3 in human disease and therapy
Molecular Biology Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.