Abstract
The pluripotency factor Lin28 inhibits the biogenesis of the let-7 family of mammalian microRNAs1,2,3,4. Lin28 is highly expressed in embryonic stem cells and has a fundamental role in regulation of development5, glucose metabolism6 and tissue regeneration7. Overexpression of Lin28 is correlated with the onset of numerous cancers8, whereas let-7, a tumour suppressor, silences several human oncogenes5. Lin28 binds to precursor let-7 (pre-let-7) hairpins9, triggering the 3′ oligo-uridylation activity of TUT4 and TUT7 (refs 10, 11, 12). The oligoU tail added to pre-let-7 serves as a decay signal, as it is rapidly degraded by Dis3l2 (refs 13, 14), a homologue of the catalytic subunit of the RNA exosome. The molecular basis of Lin28-mediated recruitment of TUT4 and TUT7 to pre-let-7 and its subsequent degradation by Dis3l2 is largely unknown. To examine the mechanism of Dis3l2 substrate recognition we determined the structure of mouse Dis3l2 in complex with an oligoU RNA to mimic the uridylated tail of pre-let-7. Three RNA-binding domains form an open funnel on one face of the catalytic domain that allows RNA to navigate a path to the active site different from that of its exosome counterpart. The resulting path reveals an extensive network of uracil-specific interactions spanning the first 12 nucleotides of an oligoU-tailed RNA. We identify three U-specificity zones that explain how Dis3l2 recognizes, binds and processes uridylated pre-let-7 in the final step of the Lin28–let-7 pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Viswanathan, S. R., Daley, G. Q. & Gregory, R. I. Selective blockade of microRNA processing by Lin28. Science 320, 97–100 (2008)
Heo, I. et al. Lin28 mediates the terminal uridylation of let-7 precursor MicroRNA. Mol. Cell 32, 276–284 (2008)
Rybak, A. et al. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nature Cell Biol. 10, 987–993 (2008)
Newman, M. A., Thomson, J. M. & Hammond, S. M. Lin-28 interaction with the Let-7 precursor loop mediates regulated microRNA processing. RNA 14, 1539–1549 (2008)
Thornton, J. E. & Gregory, R. I. How does Lin28 let-7 control development and disease? Trends Cell Biol. 22, 474–482 (2012)
Zhu, H. et al. The Lin28/let-7 axis regulates glucose metabolism. Cell 147, 81–94 (2011)
Shyh-Chang, N. et al. Lin28 enhances tissue repair by reprogramming cellular metabolism. Cell 155, 778–792 (2013)
Urbach, A. et al. Lin28 sustains early renal progenitors and induces Wilms tumor. Genes Dev. 28, 971–982 (2014)
Nam, Y., Chen, C., Gregory, R. I., Chou, J. J. & Sliz, P. Molecular basis for interaction of let-7 microRNAs with Lin28. Cell 147, 1080–1091 (2011)
Hagan, J. P., Piskounova, E. & Gregory, R. I. Lin28 recruits the TUTase Zcchc11 to inhibit let-7 maturation in mouse embryonic stem cells. Nature Struct. Mol. Biol. 16, 1021–1025 (2009)
Heo, I. et al. TUT4 in concert with Lin28 suppresses microRNA biogenesis through pre-microRNA uridylation. Cell 138, 696–708 (2009)
Thornton, J. E., Chang, H. M., Piskounova, E. & Gregory, R. I. Lin28-mediated control of let-7 microRNA expression by alternative TUTases Zcchc11 (TUT4) and Zcchc6 (TUT7). RNA 18, 1875–1885 (2012)
Chang, H. M., Triboulet, R., Thornton, J. E. & Gregory, R. I. A role for the Perlman syndrome exonuclease Dis3l2 in the Lin28–let-7 pathway. Nature 497, 244–248 (2013)
Ustianenko, D. et al. Mammalian DIS3L2 exoribonuclease targets the uridylated precursors of let-7 miRNAs. RNA 19, 1632–1638 (2013)
Liu, Q., Greimann, J. C. & Lima, C. D. Reconstitution, activities, and structure of the eukaryotic RNA exosome. Cell 127, 1223–1237 (2006)
Dziembowski, A., Lorentzen, E., Conti, E. & Seraphin, B. A single subunit, Dis3, is essentially responsible for yeast exosome core activity. Nature Struct. Mol. Biol. 14, 15–22 (2007)
Tomecki, R. et al. The human core exosome interacts with differentially localized processive RNases: hDIS3 and hDIS3L. EMBO J. 29, 2342–2357 (2010)
Astuti, D. et al. Germline mutations in DIS3L2 cause the Perlman syndrome of overgrowth and Wilms tumor susceptibility. Nature Genet. 44, 277–284 (2012)
Malecki, M. et al. The exoribonuclease Dis3L2 defines a novel eukaryotic RNA degradation pathway. EMBO J. 32, 1842–1854 (2013)
Lubas, M. et al. Exonuclease hDIS3L2 specifies an exosome-independent 3′-5′ degradation pathway of human cytoplasmic mRNA. EMBO J. 32, 1855–1868 (2013)
Chang, H., Lim, J., Ha, M. & Kim, V. N. TAIL-seq: Genome-wide determination of poly(A) tail length and 3′ end modifications. Mol. Cell 53, 1044–1052 (2014)
Choi, Y. S., Patena, W., Leavitt, A. D. & McManus, M. T. Widespread RNA 3′-end oligouridylation in mammals. RNA 18, 394–401 (2012)
Lorentzen, E., Basquin, J., Tomecki, R., Dziembowski, A. & Conti, E. Structure of the active subunit of the yeast exosome core, Rrp44: diverse modes of substrate recruitment in the RNase II nuclease family. Mol. Cell 29, 717–728 (2008)
Makino, D. L., Baumgartner, M. & Conti, E. Crystal structure of an RNA-bound 11-subunit eukaryotic exosome complex. Nature 495, 70–75 (2013)
Bonneau, F., Basquin, J., Ebert, J., Lorentzen, E. & Conti, E. The yeast exosome functions as a macromolecular cage to channel RNA substrates for degradation. Cell 139, 547–559 (2009)
Frazão, C. et al. Unravelling the dynamics of RNA degradation by ribonuclease II and its RNA-bound complex. Nature 443, 110–114 (2006)
Barbas, A. et al. Determination of key residues for catalysis and RNA cleavage specificity: one mutation turns RNase II into a “SUPER-ENZYME”. J. Biol. Chem. 284, 20486–20498 (2009)
Reis, F. P. et al. Modulating the RNA processing and decay by the exosome: altering Rrp44/Dis3 activity and end-product. PLoS ONE 8, e76504 (2013)
Rajkowitsch, L. et al. RNA chaperones, RNA annealers and RNA helicases. RNA Biol. 4, 118–130 (2007)
Bieniossek, C., Richmond, T. J. & Berger, I. MultiBac: multigene baculovirus-based eukaryotic protein complex production. Curr. Protoc. Protein Sci. Ch. 5, Unit 5 20. (2008)
Li, M. Z. & Elledge, S. J. Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nature Methods 4, 251–256 (2007)
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010)
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D 67, 293–302 (2011)
Bunkóczi, G. & Read, R. J. Improvement of molecular-replacement models with Sculptor. Acta Crystallogr. D 67, 303–312 (2011)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Terwilliger, T. C. et al. Iterative model building, structure refinement and density modification with the PHENIX AutoBuild wizard. Acta Crystallogr. D 64, 61–69 (2008)
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D 56, 965–972 (2000)
Cowtan, K. The Buccaneer software for automated model building. 1. Tracing protein chains. Acta Crystallogr. D 62, 1002–1011 (2006)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, 375–383 (2007)
Price, S. R., Ito, N., Oubridge, C., Avis, J. M. & Nagai, K. Crystallization of RNA-protein complexes. I. Methods for the large-scale preparation of RNA suitable for crystallographic studies. J. Mol. Biol. 249, 398–408 (1995)
Shechner, D. M. & Bartel, D. P. The structural basis of RNA-catalyzed RNA polymerization. Nature Struct. Mol. Biol. 18, 1036–1042 (2011)
Acknowledgements
We thank C. M. Hammell for comments on this manuscript and members of the Joshua-Tor laboratory for comments and suggestions. We thank A. Héroux for help at the National Synchrotron Light Source, which is supported by Department of Energy, Office of Basic Energy Sciences. We also thank the Protein Core Facility at Columbia University. This work was supported by the Watson School of Biological Sciences (to J.W. and L.J.), the Louis Morin Charitable Trust and the Robertson Research Fund of Cold Spring Harbor Laboratory (to L.J.). L.J. is an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.R.F., J.W. and L.J. designed and C.R.F and J.W. conducted all experiments. All authors contributed to data analysis and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Substrate specificity of Dis3l2.
a, Dis3l2 exonuclease assays were conducted with 5′ radio-labelled RNA substrates at 30 °C over a 16 min time course and products resolved by denaturing urea PAGE. Two representative gels, one from an exonuclease assay with U15 that was used to quantify Dis3l2 enzymatic activity and another for deoxyU15, showing no detectable activity under the initial rate conditions tested. b, The initial rate plots for each substrate used to calculate specific activity as shown in Fig. 1a. Exonuclease assays were conducted in triplicate and quantified with ImageJ. The concentration of enzyme used to measure the initial rate is indicated on each plot. The amount of substrate degraded (fmol) was plotted against time and the initial rate (slope) was determined with linear regression using GraphPad Prism 6. Mean ± s.d. (n = 3) are shown. c, Representative slot blot filter-binding assay of Dis3l2 with U15. Dis3l2–RNA complexes were captured on the top nitrocellulose membrane and unbound RNA to the bottom nylon membrane. d, The equilibrium dissociation constant (Kd) for each substrate in Fig. 1b was determined by plotting the fraction RNA bound against the concentration of free Dis3l2 and fit by nonlinear regression with GraphPad Prism 6. The mean of three independently measured replicates with error bars representing ± s.d. are shown with the Kd.
Extended Data Figure 2 Sequence alignment of mouse and human Dis3l2, yeast Rrp44 and human Dis3.
Dis3l2 is composed of two cold shock domains (CSD1, residues 49–240, α1, β1–β5, and CSD2, residues 241–324, β6–β10, α2) at its N terminus followed by a catalytic RNB domain (residues 325–765, α3–α17, β11–β21) and flanked by an S1 domain (residues 766–856, β22–β29). The secondary structure elements deduced from the structure of mouse Dis3l2 are shown on top of the sequences, colour-coded by domain as in Fig. 1a. The segments of mouse Dis3l2 that were truncated to facilitate crystallization are outlined in a black box. Conserved amino acid residues are coloured blue. Blue diamonds indicate residues interacting with the backbone of U14 RNA and red diamonds denote conserved active-site residues. Residues in Dis3l2 that interact with U14 bases are shaded purple.
Extended Data Figure 3 Substrate specificity of truncated Dis3l2 and electron density map for U14 RNA.
a, Comparison of specific activity (nt min−1 molecule−1) of truncated Dis3l2 construct resembling the one used in crystallization, but without the D389N mutation with the same set of substrates analysed for wild-type Dis3l2. Truncated Dis3l2 had comparable levels of activity and displayed the same substrate preference as wild-type Dis3l2. The specific activity was calculated from the initial rate plots, in the same way as described in Fig. 1a and Extended Data Fig. 1. The mean ± s.d. (n = 3) are shown. b, The final model of Dis3l2 is shown as a transparent white cartoon. The final model of U14 is rendered as yellow sticks. Initial molecular replacement phases were improved by density modification with Resolve32 and automated model building with Buccaneer33, as implemented by the AutoBuild wizard in Phenix31. The unbiased density modified electron density map before inclusion of RNA is shown contoured at 1σ. Despite the crystallization of Dis3l2 in complex with U13, unambiguous density allowed modelling of U14 (details are discussed in the Methods). Clear electron density accounted for all 14 nucleotides, except some disorder contributed to weak electron density surrounding U5, precluding its accurate placement in density.
Extended Data Figure 4 Dis3l2 domain interface analysis.
a, Analysis of the CSD1–RNB interface. The conformation of CSD1 is stabilized by two protein–protein interactions with the RNB (K240 with D739 and D91 with T613) and an RNA-mediated interaction with the RNB through U6–U7 and α11. b, Analysis of the CSD2–RNB interface. CSD2 is intimately associated with CSD1, but also forms an interface with the RNB through α3 (S242 with E337 and K319 with E332). c, Analysis of the S1–RNB interface. S1 is part of a large hydrophobic interface with RNB (α11, α13 and α17, β18 and β19). The S1 domain also forms an interface with the RNB through interactions with U8 (also see Fig. 3d) d, CSD1 is further stabilized through an RNA-mediated interaction with S1. The backbone phosphate of U4 bridges K78 of CSD1 and R792 of S1.
Extended Data Figure 5 OligoU length preference of Dis3l2.
a, Specific activity (nt min−1 molecule−1) of Dis3l2 with increasing length oligoU (U9, U11, U13 and U15). The calculation of specific activity was conducted as in Fig. 1a. Mean ± s.d. (n = 3) are shown. b, The initial rate plots for each substrate used to calculate specific activity in panel a. The amount of substrate degraded (fmol) was plotted against time and the initial rate (slope) was determined with linear regression using GraphPad Prism 6. Mean ± s.d. (n = 3) are shown. c, Equilibrium dissociation constants (Kd) for increasing oligoU length. The Kd ± s.d. determined from three independent replicates is shown. d, The equilibrium dissociation constant (Kd) for each substrate in panel c was determined by plotting the fraction RNA bound versus the concentration of free Dis3l2 and fit by nonlinear regression with GraphPad Prism 6. The mean of three independently measured replicates with error bars representing ± s.d. are shown with the Kd.
Extended Data Figure 6 Dis3l2 has a conserved active site and an RNA path that resembles RNase II.
a, The active site of the Dis3l2–U14 complex. Conserved active site residues are shown as green sticks and U13 and U14 are shown as orange sticks. A single Mg2+ ion (purple sphere) is modelled in the active site. As proposed for RNase II, Dis3l2 may utilize a two Mg2+ ion mechanism during catalysis. b, Superposition of Dis3l2 and yeast Rrp44 (PDB 2VNU) active sites. Dis3l2 side chains and the Mg2+ ion are coloured the same as in panel a, and yeast Rrp44 residues are shown as grey sticks. c, Dis3l2–U14 complex in an identical layout as Fig. 2a. A wide-open funnel created by the CSD lobe and S1 allows RNA to access the top of the RNB. d, The structure of E. coli RNase II22 (PDB 2IX1). The path of RNA in RNase II more closely resembles that in Dis3l2, compared to yeast Rrp44, although narrow along its length, underscoring its ability to accommodate only single-stranded RNA substrates. e, Perpendicular side and top views of the electrostatic surface potential of Dis3l2 (contoured at ±5 kT e−1; white is neutral, blue is positive and red is negative). A much wider positively charged funnel on the top of the RNB supports the ability of Dis3l2 to degrade structured RNA substrates. f, Electrostatic surface potential of the single-strand-specific RNase II, in the same configuration as panel c. A narrow RNA binding channel can only accommodate single-stranded RNA.
Extended Data Figure 7 Dis3l2 discriminates U over A and C.
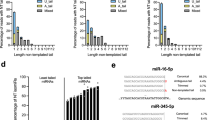
a, Dis3l2 selects oligoU-tailed substrates by way of an extensive U-specific interaction network along most of its binding path. In U-zone 1, Dis3l2 makes both main-chain and side-chain-mediated hydrogen bonds with uracil. The hydrogen bond network with U14 is disrupted for modelled A14 and C14 RNAs. Dis3l2 residues are shown as sticks colour-coded by domain as in Fig. 1c. RNA is shown as orange sticks. Hydrogen bond pairs are shown as dashed lines and disrupted hydrogen bonds are shown as curved red lines. b, U-zone 2, with the same layout as panel a. c, U-zone 3.
Extended Data Figure 8 Initial rate plots of Dis3l2 U-zone mutants.
a, Initial rate plots for U-zone 1 used to calculate specific activity as shown in Fig. 3f. Assays were conducted under the same conditions as described for U15 in Extended Data Fig. 1, where reactions contained 0.5 nM enzyme and 100 nM 5′ radio-labelled U15. Mean ± s.d. (n = 3) are shown. b, Initial rate plots for U-zone 2, conducted as described for panel a. c, Initial rate plots for U-zone 3, conducted as described for panel a.
Extended Data Figure 9 Binding affinity of Dis3l2 U-zone mutants.
a, Slot blot filter-binding assay was used to measure the binding affinity of selected Dis3l2 mutants with U15. The equilibrium dissociation constant (Kd) for U-zone 1 mutants as shown in Fig. 3g was determined by plotting the fraction RNA bound versus the concentration of free protein and fit by nonlinear regression with GraphPad Prism 6. The mean of three independently measured replicates with error bars representing ± s.d. are shown with the Kd. b, The equilibrium dissociation constant (Kd) for U-zone 2 mutants as shown in Fig. 3g, and measured as described for panel a. c, The Kd for U-zone 3 as shown in Fig. 3g, and measured as described for panel a.
Rights and permissions
About this article
Cite this article
Faehnle, C., Walleshauser, J. & Joshua-Tor, L. Mechanism of Dis3l2 substrate recognition in the Lin28–let-7 pathway. Nature 514, 252–256 (2014). https://doi.org/10.1038/nature13553
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13553
This article is cited by
-
microRNAs in action: biogenesis, function and regulation
Nature Reviews Genetics (2023)
-
DIS3L2 knockdown impairs key oncogenic properties of colorectal cancer cells via the mTOR signaling pathway
Cellular and Molecular Life Sciences (2023)
-
TENT2, TUT4, and TUT7 selectively regulate miRNA sequence and abundance
Nature Communications (2022)
-
RNA modifications: importance in immune cell biology and related diseases
Signal Transduction and Targeted Therapy (2022)
-
piRNAs Interact with Cold-Shock Domain-Containing RNA Binding Proteins and Regulate Neuronal Gene Expression During Differentiation
Molecular Neurobiology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.