Abstract
Neurotransmitter-gated ion channels of the Cys-loop receptor family mediate fast neurotransmission throughout the nervous system. The molecular processes of neurotransmitter binding, subsequent opening of the ion channel and ion permeation remain poorly understood. Here we present the X-ray structure of a mammalian Cys-loop receptor, the mouse serotonin 5-HT3 receptor, at 3.5 Å resolution. The structure of the proteolysed receptor, made up of two fragments and comprising part of the intracellular domain, was determined in complex with stabilizing nanobodies. The extracellular domain reveals the detailed anatomy of the neurotransmitter binding site capped by a nanobody. The membrane domain delimits an aqueous pore with a 4.6 Å constriction. In the intracellular domain, a bundle of five intracellular helices creates a closed vestibule where lateral portals are obstructed by loops. This 5-HT3 receptor structure, revealing part of the intracellular domain, expands the structural basis for understanding the operating mechanism of mammalian Cys-loop receptors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Maricq, A. V., Peterson, A. S., Brake, A. J., Myers, R. M. & Julius, D. Primary structure and functional expression of the 5HT3 receptor, a serotonin-gated ion channel. Science 254, 432–437 (1991)
Lummis, S. C. R. 5-HT3 receptors. J. Biol. Chem. 287, 40239–40245 (2012)
Thompson, A. J., Lester, H. A. & Lummis, S. C. R. The structural basis of function in Cys-loop receptors. Q. Rev. Biophys. 43, 449–499 (2010)
Corringer, P.-J. et al. Structure and pharmacology of pentameric receptor channels: from bacteria to brain. Structure 20, 941–956 (2012)
Walstab, J., Rappold, G. & Niesler, B. 5-HT3 receptors: role in disease and target of drugs. Pharmacol. Ther. 128, 146–169 (2010)
Changeux, J.-P. 50 years of allosteric interactions: the twists and turns of the models. Nature Rev. Mol. Cell Biol. 14, 819–829 (2013)
Schmauder, R., Kosanic, D., Hovius, R. & Vogel, H. Correlated optical and electrical single-molecule measurements reveal conformational diffusion from ligand binding to channel gating in the nicotinic acetylcholine receptor. ChemBioChem 12, 2431–2434 (2011)
daCosta, C. J. B. & Baenziger, J. E. Gating of pentameric ligand-gated ion channels: structural insights and ambiguities. Structure 21, 1271–1283 (2013)
Auerbach, A. The gating isomerization of neuromuscular acetylcholine receptors. J. Physiol. (Lond.) 588, 573–586 (2010)
Unwin, N. Refined structure of the nicotinic acetylcholine receptor at 4 Å resolution. J. Mol. Biol. 346, 967–989 (2005)
Unwin, N. & Fujiyoshi, Y. Gating movement of acetylcholine receptor caught by plunge-freezing. J. Mol. Biol. 422, 617–634 (2012)
Hilf, R. J. C. & Dutzler, R. Structure of a potentially open state of a proton-activated pentameric ligand-gated ion channel. Nature 457, 115–118 (2009)
Bocquet, N. et al. X-ray structure of a pentameric ligand-gated ion channel in an apparently open conformation. Nature 457, 111–114 (2009)
Prevost, M. S. et al. A locally closed conformation of a bacterial pentameric proton-gated ion channel. Nature Struct. Mol. Biol. 19, 642–649 (2012)
Sauguet, L. et al. Structural basis for ion permeation mechanism in pentameric ligand-gated ion channels. EMBO J. 32, 728–741 (2013)
Sauguet, L. et al. Crystal structures of a pentameric ligand-gated ion channel provide a mechanism for activation. Proc. Natl Acad. Sci. USA 111, 966–971 (2014)
Hilf, R. J. C. & Dutzler, R. X-ray structure of a prokaryotic pentameric ligand-gated ion channel. Nature 452, 375–379 (2008)
Spurny, R. et al. Pentameric ligand-gated ion channel ELIC is activated by GABA and modulated by benzodiazepines. Proc. Natl Acad. Sci. USA 109, E3028–E3034 (2012)
Hibbs, R. E. & Gouaux, E. Principles of activation and permeation in an anion-selective Cys-loop receptor. Nature 474, 54–60 (2011)
Hassaïne, G. et al. Large scale expression and purification of the mouse 5-HT3 receptor. Biochim. Biophys. Acta 1828, 2544–2552 (2013)
Haeger, S. et al. An intramembrane aromatic network determines pentameric assembly of Cys-loop receptors. Nature Struct. Mol. Biol. 17, 90–98 (2010)
Muyldermans, S. Nanobodies: natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013)
Livesey, M. R., Cooper, M. A., Lambert, J. J. & Peters, J. A. Rings of charge within the extracellular vestibule influence ion permeation of the 5-HT3A receptor. J. Biol. Chem. 286, 16008–16017 (2011)
Hansen, S. B., Wang, H. L., Taylor, P. & Sine, S. M. An ion selectivity filter in the extracellular domain of Cys-loop receptors reveals determinants for ion conductance. J. Biol. Chem. 283, 36066–36070 (2008)
Moroni, M., Meyer, J. O., Lahmann, C. & Sivilotti, L. G. In glycine and GABAA channels, different subunits contribute asymmetrically to channel conductance via residues in the extracellular domain. J. Biol. Chem. 286, 13414–13422 (2011)
Reeves, D. C., Goren, E. N., Akabas, M. H. & Lummis, S. C. Structural and electrostatic properties of the 5–HT3 receptor pore revealed by substituted cysteine accessibility mutagenesis. J. Biol. Chem. 276, 42035–42042 (2001)
Panicker, S., Cruz, H., Arrabit, C. & Slesinger, P. A. Evidence for a centrally located gate in the pore of a serotonin-gated ion channel. J. Neurosci. 22, 1629–1639 (2002)
Beckstein, O. & Sansom, M. S. P. A hydrophobic gate in an ion channel: the closed state of the nicotinic acetylcholine receptor. Phys. Biol. 3, 147–159 (2006)
Wang, H., Cheng, X., Taylor, P., McCammon, J. & Sine, S. Control of cation permeation through the nicotinic receptor channel. PLOS Comput. Biol. 4, e41 (2008)
Corringer, P. J. et al. Mutational analysis of the charge selectivity filter of the alpha7 nicotinic acetylcholine receptor. Neuron 22, 831–843 (1999)
Thompson, A. J. & Lummis, S. C. R. A single ring of charged amino acids at one end of the pore can control ion selectivity in the 5-HT3 receptor. Br. J. Pharmacol. 140, 359–365 (2003)
Cymes, G. D. & Grosman, C. The unanticipated complexity of the selectivity-filter glutamates of nicotinic receptors. Nature Chem. Biol. 8, 975–981 (2012)
Zuber, B. & Unwin, N. Structure and superorganization of acetylcholine receptor-rapsyn complexes. Proc. Natl Acad. Sci. USA 110, 10622–10627 (2013)
Bouzat, C., Bren, N. & Sine, S. M. Structural basis of the different gating kinetics of fetal and adult acetylcholine receptors. Neuron 13, 1395–1402 (1994)
Kelley, S. P., Dunlop, J. I., Kirkness, E. F., Lambert, J. J. & Peters, J. A. A cytoplasmic region determines single-channel conductance in 5-HT3 receptors. Nature 424, 321–324 (2003)
Peters, J. A. et al. Novel structural determinants of single channel conductance and ion selectivity in 5-hydroxytryptamine type 3 and nicotinic acetylcholine receptors. J. Physiol. (Lond.) 588, 587–596 (2010)
McKinnon, N., Bali, M. & Akabas, M. H. 5-HT3 receptor ion size selectivity is a property of the transmembrane channel, not the cytoplasmic vestibule portals. J. Gen. Physiol. 138, 453–466 (2011)
Carland, J. E. et al. Mutagenic analysis of the intracellular portals of the human 5–HT3A receptor. J. Biol. Chem. 288, 31592–31601 (2013)
Kozuska, J. L. et al. Impact of intracellular domain flexibility upon properties of activated human 5–HT3 receptors. Br. J. Pharmacol. 171, 1617–1628 (2014)
Brejc, K. et al. Crystal structure of an ACh-binding protein reveals the ligand-binding domain of nicotinic receptors. Nature 411, 269–276 (2001)
Hansen, S. B. et al. Structures of Aplysia AChBP complexes with nicotinic agonists and antagonists reveal distinctive binding interfaces and conformations. EMBO J. 24, 3635–3646 (2005)
Kesters, D. et al. Structural basis of ligand recognition in 5-HT3 receptors. EMBO Rep. 14, 49–56 (2013)
Huang, S. et al. Complex between α-bungarotoxin and an α7 nicotinic receptor ligand-binding domain chimaera. Biochem. J. 454, 303–310 (2013)
Beene, D. L. et al. Cation-π interactions in ligand recognition by serotonergic (5-HT3A) and nicotinic acetylcholine receptors: the anomalous binding properties of nicotine. Biochemistry 41, 10262–10269 (2002)
Zhong, W. et al. From ab initio quantum mechanics to molecular neurobiology: A cation–π binding site in the nicotinic receptor. Proc. Natl Acad. Sci. USA 95, 12088–12093 (1998)
Miles, T. F., Bower, K. S., Lester, H. A. & Dougherty, D. A. A coupled array of noncovalent interactions impacts the function of the 5-HT 3A serotonin receptor in an agonist-specific way. ACS Chem. Neurosci. 3, 753–760 (2012)
Hu, X.-Q., Zhang, L., Stewart, R. R. & Weight, F. F. Arginine 222 in the pre-transmembrane domain 1 of 5-HT3A receptors links agonist binding to channel gating. J. Biol. Chem. 278, 46583–46589 (2003)
Lee, W. Y. & Sine, S. M. Principal pathway coupling agonist binding to channel gating in nicotinic receptors. Nature 438, 243–247 (2005)
Mukhtasimova, N. & Sine, S. M. Nicotinic receptor transduction zone: invariant arginine couples to multiple electron-rich residues. Biophys. J. 104, 355–367 (2013)
Li, T. et al. Cell-penetrating anti-GFAP VHH and corresponding fluorescent fusion protein VHH-GFP spontaneously cross the blood–brain barrier and specifically recognize astrocytes: application to brain imaging. FASEB J. 26, 3969–3979 (2012)
Muller, N., Girard, P., Hacker, D. L., Jordan, M. & Wurm, F. M. Orbital shaker technology for the cultivation of mammalian cells in suspension. Biotechnol. Bioeng. 89, 400–406 (2005)
Aimon, S. et al. Functional reconstitution of a voltage-gated potassium channel in giant unilamellar vesicles. PLoS ONE 6, e25529 (2011)
Montes, L.-R., Alonso, A., Goñi, F. M. & Bagatolli, L. A. Giant unilamellar vesicles electroformed from native membranes and organic lipid mixtures under physiological conditions. Biophys. J. 93, 3548–3554 (2007)
Hovius, R. et al. Characterization of a mouse serotonin 5-HT3 receptor purified from mammalian cells. J. Neurochem. 70, 824–834 (1998)
Conrath, K. et al. Camelid nanobodies raised against an integral membrane enzyme, nitric oxide reductase. Protein Sci. 18, 619–628 (2009)
Karlsson, R., Katsamba, P. S., Nordin, H., Pol, E. & Myszka, D. G. Analyzing a kinetic titration series using affinity biosensors. Anal. Biochem. 349, 136–147 (2006)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
McCoy, A. J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D 63, 32–41 (2007)
Schröder, G. F., Levitt, M. & Brunger, A. T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D 68, 352–367 (2012)
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT. Acta Crystallogr. D 60, 2210–2221 (2004)
Herget, S., Ranzinger, R., Maass, K. & Lieth, C.-W. V. D. GlycoCT-a unifying sequence format for carbohydrates. Carbohydr. Res. 343, 2162–2171 (2008)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
The PyMOL Molecular Graphics System. Version 1.6, Schrödinger, LLC. (2013)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996)
Koehl, P. & Delarue, M. AQUASOL: An efficient solver for the dipolar Poisson-Boltzmann-Langevin equation. J. Chem. Phys. 132, 064101 (2010)
Ho, B. K. & Gruswitz, F. HOLLOW: generating accurate representations of channel and interior surfaces in molecular structures. BMC Struct. Biol. 8, 49 (2008)
Chovancova, E. et al. CAVER 3.0: a tool for the analysis of transport pathways in dynamic protein structures. PLOS Comput. Biol. 8, e1002708 (2012)
Calimet, N. et al. From the Cover: A gating mechanism of pentameric ligand-gated ion channels. Proc. Natl Acad. Sci. USA 110, E3987–E3996 (2013)
Ulens, C. et al. Structural determinants of selective alpha-conotoxin binding to a nicotinic acetylcholine receptor homolog AChBP. Proc. Natl Acad. Sci. USA 103, 3615–3620 (2006)
Purohit, P. & Auerbach, A. Acetylcholine receptor gating at extracellular transmembrane domain interface: the ‘pre-M1’ linker. J. Gen. Physiol. 130, 559–568 (2007)
Dang, H., England, P. M., Farivar, S. S., Dougherty, D. A. & Lester, H. A. Probing the role of a conserved M1 proline residue in 5-hydroxytryptamine3 receptor gating. Mol. Pharmacol. 57, 1114–1122 (2000)
Acknowledgements
Research in the laboratory of H.V. was supported by the Swiss National Science Foundation (grant 31003A-133141; SystemsX/CINA program), internal funds of the Ecole Polytechnique Fédérale de Lausanne, the European Community’s Seventh Framework Program FP7/2007-2013 under Grant 211800, the Human Frontier Science Program (postdoc fellowship to H.N.). Additional support was provided by The Agence Nationale de la Recherche (VenomPicoScreen project, grant ANR-11-RPIB-022-04). We thank S. Thurnheer and the expression facility, F. Pojer and the crystallization facility at the EPFL, D. Blot, the HTXLab and the SLS and ESRF beamlines staff. We are particularly appreciative of discussions with H.V. laboratory members and E. Pebay-Peyroula. We thank the company Theranyx for the discovery and production of the functional nanobody used for the structure resolution.
Author information
Authors and Affiliations
Contributions
H.V. initiated the project and was responsible for overall project management and strategy. G.H. designed the research for the expression, purification and nanobody aspects. C.D., G.H. and H.N. performed purification. A.D. and G.H. made the nanobodies. C.D., G.H., H.N., L.G., R.W., M.B.T., R.H. and C.M. performed biochemical, biophysical and functional characterization of the receptor and/or of the complexes with nanobodies. A.G. and H.S. performed electron microscopy characterization. X.-D.L., T.T., C.D. and H.N. performed crystallization and data collection. H.N. solved the structure. H.N. and F.P. analysed the structure. H.N. wrote the manuscript with help of H.V. and F.P. All authors discussed results and commented on manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.H. declares employment in the company Theranyx, which sells 5-HT3 receptors and other membrane receptors.
Extended data figures and tables
Extended Data Figure 1 Limited proteolysis preserves the 5-HT3 receptor structural and functional integrity.
All data shown were obtained with purified 5-HT3 receptor submitted to limited digestion in the presence of trypsin. a, Size-exclusion chromatography profile on a Superose 6 10/300 column with a major peak of the pentameric protein–detergent complex. b, SDS–PAGE of the peak pooled fractions presents two bands corresponding to the N-terminal and C-terminal fragments of the receptor. c, Single-particle analysis using negative-stained transmission electron microscopy of the same fraction depicts the structural envelope of an intact split receptor (left, electron micrograph scale bar: 50 nm; right, selection of class averages, box sizes: 38.5 nm). d, After reconstitution in GUVs (middle panel), the split receptor exhibits single-channel activities provoked by serotonin present in the patch pipette (left panel) with a single-channel conductance of ∼30 pS (right panel). e, The split receptor in detergent (blue) or reconstituted in liposomes (black) presents equivalent [3H]GR65630 binding profiles (Kd = 0.3 nM in both cases) compared to the wild-type receptor in cell membranes20 (red). VHH15 was found to be a potent inhibitor of the 5-HT3 receptor. f, Schematic of the protocol used to quantify the inhibition of 5-HT3 receptor by the nanobody using automated two-electrode voltage-clamp recording on Xenopus oocytes. g, Representative recordings of serotonin-evoked activation after VHH incubation (red traces) or after buffer incubation (grey traces, negative control). The time of incubation with VHH or the buffer is not scaled as only 20 s of recording is shown. h, Inhibition of the serotonin signal upon different nanobody concentrations. The Hill equation was used to fit the data (red line) leading to a half maximal inhibitory concentration of 29 nM. 11 oocytes have been tested. i, Validation of the VHH15 inhibitory effect on the HEK 293 cells expressing the 5-HT3 receptor used for overexpression. The pulse train contains serotonin applications (orange: 5 μM, 30 s) and VHH15 application (red: 100 nM, 3 min). j, Analysis of the interaction of VHH15 with the 5-HT3 receptor by surface plasmon resonance. Experimental sensorgram (dashed curve) was recorded in single-cycle mode56 with sequential injection of increasing concentrations of 5HT-3 receptors (3, 9, 27, 81 and 243 nM) on the VHH15-functionalized surface. The sensorgram was fitted with a 1:1 binding model including mass transfer and drift correction (solid line) yielding an association rate constant kon = 6.2 × 104 M−1 s−1 and a dissociation rate constant koff = 4.7 × 10−6 s−1 with a derived equilibrium dissociation constant  . k, Ligand binding for VHH15 competing against [3H]GR65630. Samples were incubated with the indicated concentrations of VHH15 before measuring the specific [3H]GR65630 binding. Normalized VHH15 competition binding isotherms lead to IC50 value of ∼2 nM. Competition binding affinity of VHH15 calculated according to the Cheng–Prusoff equation is in the range of hundreds of pM.
. k, Ligand binding for VHH15 competing against [3H]GR65630. Samples were incubated with the indicated concentrations of VHH15 before measuring the specific [3H]GR65630 binding. Normalized VHH15 competition binding isotherms lead to IC50 value of ∼2 nM. Competition binding affinity of VHH15 calculated according to the Cheng–Prusoff equation is in the range of hundreds of pM.
Extended Data Figure 2 Crystallographic statistics, topology and sequence.
a, Data collection and refinement statistics table. b, r.m.s.d. after superimposition with homologous structures. Values of r.m.s.d. (Å) are computed for the core Cα atoms (which are defined in the bottom table) after superimposition. The upper diagonal (blue) is based on monomer superimposition, while the lower diagonal (pink) is based on pentamer superimposition. Values corresponding to GLIC resting state at the monomeric level are averaged over one representative (first chain) of each copy. c, Topology scheme of the extracellular domain and (d) of the entire 5-HT3 receptor. e, Sequence of the mouse 5-HT3A receptor in the crystal structure. Secondary structures and important loops are indicated with a colour code consistent with main-text figures. Prominent residues of the neurotransmitter site binding loops are highlighted (loop A, red; loop B, green; loop C, yellow; loop D, purple; loop E, blue; loop F, orange; and loop G, khaki). Pore-facing residues with their prime numbering (which allow comparison within the family) are boxed in red. Prominent residues defining the conductance in the intracellular domain are boxed in blue (the so-called RRR or QDA residues) while residues tightening the bundle of intracellular MA helices at their tips are boxed in yellow. Glycosylation sites are depicted in green in panels b and c. The mouse receptor used for this study is a variant compared to the consensus sequence as it contains an alanine insertion in the M2–M3 loop, which is highlighted by the pink box. Notably, this insertion is present in the consensus human receptor.
Extended Data Figure 3 Quality of the electron density maps.
a, Top view of a slab of the transmembrane domain, shown as ribbon (colour code as in Fig. 1c) with blue mesh representing the maximum-likelihood 2mFo−DFc electron density map contoured at a level of 1σ. b, Side view of one subunit of the 5-HT3 receptor as ribbon with the same density map. c, Enlarged views of the complementary subunit (top) (at the binding site level viewed from the principal subunit) and of the binding site (bottom). The main chain is represented as ribbon with side chains as sticks, and overlaid with the same density map. d, Enlarged view of the extracellular side of M4, which is not well defined in electron density maps (contoured at 1σ in blue and 1.5σ in yellow). e, Enlarged views of the intracellular post-M3 loop and MX helix (both green) and the triplet of arginines of MA (inset). Only Cα atoms of the MX helix were modelled. f, Stereo view of one subunit, similar to panel b.
Extended Data Figure 4 The constriction of the extracellular vestibule.
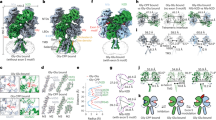
a, The structures of the extracellular vestibule of GLIC (pale green), GluCl (dark green) and Torpedo nAChR (pink) are superimposed to that of the 5-HT3 receptor (blue). For clarity, only a small stretch of the β4–β5 loop is represented as ribbon for homologous structures. The side chains of D105 and K108 of the 5-HT3 receptor, as well as the side chains of charged residues of the loop in superimposed structures, are represented as sticks. D97 of the Torpedo nAChR, which was shown to be an important determinant of conductance24, is labelled. b, View parallel to the membrane plane of the same region. c, Sequence alignment for the superimposed structures, around D105 and K108. d, Ion-density distribution, for cations (left) and anions (right), integrated radially around the pore axis. The protein density and membrane are in grey.
Extended Data Figure 5 The intracellular domain.
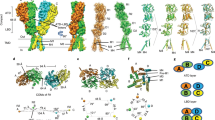
a, View parallel to the membrane plane of the intracellular domain in a surface representation. b, Cross-section of the intracellular domain in a surface representation. c, Cross-section with residues lining the vestibule represented as spheres. d, Cross-section in ribbon, stick and surface representation. e, Two orthogonal views of an obstructed portal in the intracellular domain. The protein is represented as ribbon with sticks for residues lining the narrow tunnel at the portal level. The maximum-likelihood 2mFo−DFc electron density map contoured at a level of 1σ is also represented. Mutations of D417, E42139 and R416, R420 and R42435 all influence the channel conductance. f, Stereo views depicting the position of the three key arginines in the intracellular domain, as seen from the intracellular vestibule and from the pore axis.
Extended Data Figure 6 Comparison of the MA and M4 helices of the 5-HT3 and nACh receptors.
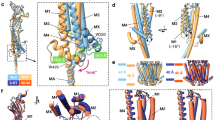
a, Superimposition of the 5-HT3 receptor (grey) and Torpedo nAChR (orange, α-subunit, 2BG9 chain D, r.m.s.d. = 3.7 Å), the 5 MA–M4 helices are shown as ribbon (for the superimposed subunit) or cartoon. b, Enlarged view of the superimposed subunit, showing as sticks the arginine triplet, the conserved W426 and D434 of the 5-HT3 receptor and equivalent residues of the α-nACh receptor. There is an offset of −1 residue at the level of W426 and +1 residue at the level of D434 (the change in offset is due to the break of the helical structure in Torpedo nAChR between MA and M4). c, Superimposition of all Torpedo nAChR subunits with the 5-HT3 receptor. The conserved tryptophan and aspartate residues are at equivalent positions in all Torpedo nAChR subunits. d, Sequence alignment showing the conserved positions in purple. Bars indicate the offset in structure superpositions. e, f, Two orthogonal views of the superimposition with proteins depicted as cartoons, illustrating the global conformations of the intracellular domain.
Extended Data Figure 7 The neurotransmitter binding site.
a, Cartoon representation of a binding site at the interface between two subunits, viewed parallel to the membrane plane. Binding loops are indicated. b, Same view with solvent-accessible electrostatic potential mapped on the molecular surface, colour coded from red to blue (−10 to +10 kT e−1, respectively). c, Same view with the surface removed for loop C, highlighting the electronegative binding site interior. d, The principal subunit Cα atoms were superimposed for epibatidine-bound AChBP24 (Protein Data Bank accession 2BYQ, blue, r.m.s.d. = 1.07 Å) and α-conotoxin-bound AChBP73 (Protein Data Bank accession 2C9T, grey, r.m.s.d. = 1.04 Å) on the extracellular domain of the 5-HT3 receptor (orange). e, Top view of a slab (black line) of the superimposed structure. The loop C of the 5-HT3 receptor adopts an intermediate conformation compared to the contracted loop of epibatidine-bound AChBP and the open loop of α-conotoxin-bound AChBP. f, View of the complementary subunits when principal subunits are superimposed, as in a and b. The β-sheet of the 5-HT3 receptor clearly deviates from these of the AChBP, indicating a different quaternary organization. g, Top view of two 5-HT3 receptor subunits with one VHH15 (cartoon and surface), with the neurotransmitter binding cavity represented as molecular surface (pink). The VHH15 covers the binding site, which remains accessible from below. h, Residues lining the neurotransmitter binding site are depicted on this scheme of the site, together with a sequence alignment, for each loop, of the mouse 5-HT3 receptor with one human receptor of each subfamily.
Extended Data Figure 8 Conserved residues close to the extracellular/transmembrane domains interface.
a–c, Charged residues just above the transmembrane domain. Based on the observations that a charged residue in the pre-M1is important for gating47,48,74, we represent the charged residues of one subunit located at the same altitude as R218 of the pre-M1, on X-ray structures of full-length Cys-loop receptors. In addition to this pre-M1 positively charged residue, there are two or three negatively charged residues, one in the β1–β2 loop (which can form a salt bridge with the former, E53 in the 5-HT3 receptor, D32 in GLIC, D43 in GluCl and E30 in ELIC), a conserved residue in the Cys-loop (motif FPFD145 in the 5-HT3 receptor, D122 in GLIC, D140 in GluCl and D122 in ELIC), and another conserved residue in cationic eukaryotic receptors in β8–β9 (motif 186-EW in the 5-HT3 receptor, also present in ELIC 159-EW). Interestingly, a structurally conserved aromatic residue, depicted as yellow spheres, is located just above the level of the aforementioned charged residues. Three levels are clearly depicted in panel b, with from top to bottom (that is, from site to pore) the aromatic residue, the charged residues and the conserved FPF motif of the Cys-loop. This structural organization is common to Cys-loop receptors of known structures. c, The superimposition of the 5-HT3 receptor with closed and apparently open structures suggests that the location of the sandwich structure might be characteristic of the state of the channel. Comparing the two panels, we note that the 5-HT3 receptor aromatic residue (yellow spheres) and Cys-loop motif overlay with those of the closed structures, while those of the open structures are shifted downwards and radially (arrow in d). We also note that this observation does not correspond to an equivalent domain–domain organization, as evidenced by the clear deviation of the GluCl and open GLIC core atoms of the extracellular domain (green arrows in d) and by Extended Data Fig. 8a–c. Superimpositions were performed on Cα core atoms of the M1 to M3 helices of chain A. d, M1 helical structure is broken at the conserved proline P230. View of one subunit as a ribbon with an enlarged view of M1 (thicker ribbon), which depicts αC(n)–αC(n+4) distances for the extracellular part of M1. Conserved residues R217, P230 and F242 are shown as sticks. In line with the structure, it was shown that P230 can be substituted only with non-natural amino-acids lacking hydrogen bond donor activity75.
Extended Data Figure 9 Comparison of the 5-HT3 receptor with other Cys-loop receptor structures.
a, Ribbon view of the superimposition of known structures on one subunit’s extracellular domain. This representation highlights the spread of domain–domain orientations: extracellular domains are well superimposed while transmembrane domains are spread. Structures are superimposed on their extracellular domain core Cα atoms. b, Enlarged view of the interfacial extremity of the extracellular domain, where deviations are maximal at the pre-M1 residues. Overall there is very good superimposition of the extracellular domain cores, but deviations are found at this lower level. c, Top view of the transmembrane domain of one subunit. d, Top view of the extracellular domain. The arrow indicates the direction of motion from closed to open to 5-HT3 receptor structure. e, Top view of superimposed transmembrane domains. Structures in panels d and e are superimposed using the transmembrane domain core Cα atoms of all 5 subunits. f, Side views of two superimposed cores, with a vertical pore axis (dotted line). Structures are superimposed using the transmembrane domain core Cα atoms of subunit A. g, Close-up view of the extracellular parts, in two equivalent images for clarity, one with the 5-HT3 receptor overlaid with closed structures, the second with the 5-HT3 receptor overlaid with apparently open structures.
Extended Data Figure 10 Conserved motifs of residues can be used for structural comparison.
a, Zoomed view of two archetypal motifs, conserved throughout the Cys-loop receptor family: the ‘stabilizing box’ (top) and the ‘Cys-loop region’ (bottom). The stabilizing box is made of residues belonging to or neighbouring loops defining the orthosteric binding site: motifs WXPD (loop A), S-Φ-X-H/Y (loop B; Φ being a tryptophan and H/Y a histidine in the 5-HT3 receptor), W (loop D) and GXV (loop E). The ‘Cys-loop region’ is made of residues belonging to or neighbouring loops defining the extracellular domain (ECD)–transmembrane domain (TMD) interface: Φ-P-Φ-D (loop β6–β7), R (pre-M1) and Φ (loop β8–β9); the latter being member of the GEW motif in nACh and 5-HT3 receptors. These sets of residues, shown as yellow sticks, define a core with little configurational variation among known structures of Cys-loop receptors, and can thus be considered as defining a ‘rigid rod’ (the ‘rod’ can be seen as the union of the Van der Waals balls shown as dotted spheres). b, The two rigid rods are represented as yellow surfaces for the 5-HT3 receptor, along with another conserved residue, F242, located close to the intracellular end of the M1 segment. Notably, at the quaternary level, this last residue seems to be superimposable from one structure to the other. It can thus be used to define a common framework into which structures from different species can be compared. The segment joining this residue to the conserved arginine of the pre-M1 is shown as a broken line, while the tetragon joining αC atoms of D97, V123, F142 and W187 is shown with plain lines. c, Rotated view of the 5-HT receptor, now viewed from the extracellular medium, further defining the geometric elements used to compare structures. Additional segments joining the five αCs of F242 are shown as plain lines, along with a broken line representing the segment joining the conserved Leu 9′ of M2 to the conserved Pro of loop M2–M3. d, In the framework defined above, the tetragons of the 5-HT3 receptor, GluCl, GLIC (in its apparently open form) and ELIC, are represented viewed parallel to the membrane (left) and perpendicular to the membrane (right). It appears from this representation that there is a gradual motion when going from ELIC (closed) to GLIC (open), GluCl (open) and finally to the 5-HT3 receptor. This motion can be approximated as an anti-clockwise rotation of the ECD with respect to the lower TMD when viewed from the extracellular medium. Notably, the segment used to represent the transmembrane pore status seems to follow the motion of the ECD, but in a more all-or-nothing way since ELIC is more distinguishable from the other structures than the latter between themselves. Overall, the reductionist representation of structures by their conserved motifs minimizes the bias of inter-species comparison and may present an interesting way to classify conformations. It suggests implicitly that the motion sampled by the structures compared here results both in different shapes for the neurotransmitter binding pocket and in different states for the transmembrane channel pore.
Supplementary information
Supplementary Information
This file contains a multiple sequence alignment. (PDF 4637 kb)
Rights and permissions
About this article
Cite this article
Hassaine, G., Deluz, C., Grasso, L. et al. X-ray structure of the mouse serotonin 5-HT3 receptor. Nature 512, 276–281 (2014). https://doi.org/10.1038/nature13552
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13552
This article is cited by
-
An original potentiating mechanism revealed by the cryo-EM structures of the human α7 nicotinic receptor in complex with nanobodies
Nature Communications (2023)
-
Generation of nanobodies acting as silent and positive allosteric modulators of the α7 nicotinic acetylcholine receptor
Cellular and Molecular Life Sciences (2023)
-
Structure, dynamics and lipid interactions of serotonin receptors: excitements and challenges
Biophysical Reviews (2021)
-
Molecular mechanism of setron-mediated inhibition of full-length 5-HT3A receptor
Nature Communications (2019)
-
The roles of aromatic residues in the glycine receptor transmembrane domain
BMC Neuroscience (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.