Abstract
The translational control of oncoprotein expression is implicated in many cancers. Here we report an eIF4A RNA helicase-dependent mechanism of translational control that contributes to oncogenesis and underlies the anticancer effects of silvestrol and related compounds. For example, eIF4A promotes T-cell acute lymphoblastic leukaemia development in vivo and is required for leukaemia maintenance. Accordingly, inhibition of eIF4A with silvestrol has powerful therapeutic effects against murine and human leukaemic cells in vitro and in vivo. We use transcriptome-scale ribosome footprinting to identify the hallmarks of eIF4A-dependent transcripts. These include 5′ untranslated region (UTR) sequences such as the 12-nucleotide guanine quartet (CGG)4 motif that can form RNA G-quadruplex structures. Notably, among the most eIF4A-dependent and silvestrol-sensitive transcripts are a number of oncogenes, superenhancer-associated transcription factors, and epigenetic regulators. Hence, the 5′ UTRs of select cancer genes harbour a targetable requirement for the eIF4A RNA helicase.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Wendel, H. G. et al. Survival signalling by Akt and eIF4E in oncogenesis and cancer therapy. Nature 428, 332–337 (2004)
Mamane, Y., Petroulakis, E., LeBacquer, O. & Sonenberg, N. mTOR, translation initiation and cancer. Oncogene 25, 6416–6422 (2006)
Rajasekhar, V. K. et al. Oncogenic Ras and Akt signaling contribute to glioblastoma formation by differential recruitment of existing mRNAs to polysomes. Mol. Cell 12, 889–901 (2003)
Jackson, R. J., Hellen, C. U. & Pestova, T. V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nature Rev. Mol. Cell Biol. 11, 113–127 (2010)
Lazaris-Karatzas, A., Montine, K. S. & Sonenberg, N. Malignant transformation by a eukaryotic initiation factor subunit that binds to mRNA 5′ cap. Nature 345, 544–547 (1990)
Ruggero, D. et al. The translation factor eIF-4E promotes tumor formation and cooperates with c-Myc in lymphomagenesis. Nature Med. 10, 484–486 (2004)
Wendel, H. G. et al. Dissecting eIF4E action in tumorigenesis. Genes Dev. 21, 3232–3237 (2007)
Mavrakis, K. J. et al. Tumorigenic activity and therapeutic inhibition of Rheb GTPase. Genes Dev. 22, 2178–2188 (2008)
Parsyan, A. et al. mRNA helicases: the tacticians of translational control. Nature Rev. Mol. Cell Biol. 12, 235–245 (2011)
Schatz, J. H. et al. Targeting cap-dependent translation blocks converging survival signals by AKT and PIM kinases in lymphoma. J. Exp. Med. 208, 1799–1807 (2011)
Bordeleau, M. E. et al. Therapeutic suppression of translation initiation modulates chemosensitivity in a mouse lymphoma model. J. Clin. Invest. 118, 2651–2660 (2008)
Marintchev, A. et al. Topology and regulation of the human eIF4A/4G/4H helicase complex in translation initiation. Cell 136, 447–460 (2009)
Meijer, H. A. et al. Translational repression and eIF4A2 activity are critical for microRNA-mediated gene regulation. Science 340, 82–85 (2013)
Ingolia, N. T., Ghaemmaghami, S., Newman, J. R. & Weissman, J. S. Genome-wide analysis in vivo of translation with nucleotide resolution using ribosome profiling. Science 324, 218–223 (2009)
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012)
Hsieh, A. C. et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 485, 55–61 (2012)
Palomero, T. et al. Mutational loss of PTEN induces resistance to NOTCH1 inhibition in T-cell leukemia. Nature Med. 13, 1203–1210 (2007)
Weng, A. P. et al. c-Myc is an important direct target of Notch1 in T-cell acute lymphoblastic leukemia/lymphoma. Genes Dev. 20, 2096–2109 (2006)
Zenatti, P. P. et al. Oncogenic IL7R gain-of-function mutations in childhood T-cell acute lymphoblastic leukemia. Nature Genet. 43, 932–939 (2011)
Pear, W. S. et al. Exclusive development of T cell neoplasms in mice transplanted with bone marrow expressing activated Notch alleles. J. Exp. Med. 183, 2283–2291 (1996)
Rong, L. et al. Control of eIF4E cellular localization by eIF4E-binding proteins, 4E-BPs. RNA 14, 1318–1327 (2008)
Rodrigo, C. M., Cencic, R., Roche, S. P., Pelletier, J. & Porco, J. A. Synthesis of rocaglamide hydroxamates and related compounds as eukaryotic translation inhibitors: synthetic and biological studies. J. Med. Chem. 55, 558–562 (2012)
Choo, A. Y., Yoon, S. O., Kim, S. G., Roux, P. P. & Blenis, J. Rapamycin differentially inhibits S6Ks and 4E-BP1 to mediate cell-type-specific repression of mRNA translation. Proc. Natl Acad. Sci. USA 105, 17414–17419 (2008)
Anders, S., Reyes, A. & Huber, W. Detecting differential usage of exons from RNA-seq data. Genome Res. 22, 2008–2017 (2012)
Drewe, P. et al. Accurate detection of differential RNA processing. Nucleic Acids Res. 41, 5189–5198 (2013)
Meyuhas, O. Synthesis of the translational apparatus is regulated at the translational level. Eur. J. Biochem. 267, 6321–6330 (2000)
Pelletier, J. & Sonenberg, N. Internal initiation of translation of eukaryotic mRNA directed by a sequence derived from poliovirus RNA. Nature 334, 320–325 (1988)
Bailey, T. L. DREME: motif discovery in transcription factor ChIP-seq data. Bioinformatics 27, 1653–1659 (2011)
Bugaut, A. & Balasubramanian, S. 5′-UTR RNA G-quadruplexes: translation regulation and targeting. Nucleic Acids Res. 40, 4727–4741 (2012)
Booy, E. P. et al. The RNA helicase RHAU (DHX36) unwinds a G4-quadruplex in human telomerase RNA and promotes the formation of the P1 helix template boundary. Nucleic Acids Res. 40, 4110–4124 (2012)
Chakraborty, P. & Grosse, F. Human DHX9 helicase preferentially unwinds RNA-containing displacement loops (R-loops) and G-quadruplexes. DNA Repair (Amst.) 10, 654–665 (2011)
Van Vlierberghe, P. et al. ETV6 mutations in early immature human T cell leukemias. J. Exp. Med. 208, 2571–2579 (2011)
Sadlish, H. et al. Evidence for a functionally relevant rocaglamide binding site on the eIF4A-RNA complex. ACS Chem. Biol. 8, 1519–1527 (2013)
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013)
Penn, L. J., Brooks, M. W., Laufer, E. M. & Land, H. Negative autoregulation of c-myc transcription. EMBO J. 9, 1113–1121 (1990)
Kumari, S., Bugaut, A., Huppert, J. L. & Balasubramanian, S. An RNA G-quadruplex in the 5′ UTR of the NRAS proto-oncogene modulates translation. Nature Chem. Biol. 3, 218–221 (2007)
Shahid, R., Bugaut, A. & Balasubramanian, S. The BCL-2 5′ untranslated region contains an RNA G-quadruplex-forming motif that modulates protein expression. Biochemistry 49, 8300–8306 (2010)
Morris, M. J., Negishi, Y., Pazsint, C., Schonhoft, J. D. & Basu, S. An RNA G-quadruplex is essential for cap-independent translation initiation in human VEGF IRES. J. Am. Chem. Soc. 132, 17831–17839 (2010)
Huppert, J. L., Bugaut, A., Kumari, S. & Balasubramanian, S. G-quadruplexes: the beginning and end of UTRs. Nucleic Acids Res. 36, 6260–6268 (2008)
Moerke, N. J. et al. Small-molecule inhibition of the interaction between the translation initiation factors eIF4E and eIF4G. Cell 128, 257–267 (2007)
Kentsis, A., Topisirovic, I., Culjkovic, B., Shao, L. & Borden, K. L. Ribavirin suppresses eIF4E-mediated oncogenic transformation by physical mimicry of the 7-methyl guanosine mRNA cap. Proc. Natl Acad. Sci. USA 101, 18105–18110 (2004)
Jean, G., Kahles, A., Sreedharan, V. T., De Bona, F. & Ratsch, G. RNA-Seq read alignments with PALMapper. Curr. Protoc. Bioinformatics 32, 11.6.1–11.6.37 (2010)
Flicek, P. et al. Ensembl 2013. Nucleic Acids Res. 41, D48–D55 (2013)
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013)
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Katz, Y., Wang, E. T., Airoldi, E. M. & Burge, C. B. Analysis and design of RNA sequencing experiments for identifying isoform regulation. Nature Methods 7, 1009–1015 (2010)
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011)
Hofacker, I. L. Vienna RNA secondary structure server. Nucleic Acids Res. 31, 3429–3431 (2003)
Fratta, P. et al. C9orf72 hexanucleotide repeat associated with amyotrophic lateral sclerosis and frontotemporal dementia forms RNA G-quadruplexes. Sci. Rep. 2, 1016 (2012)
Greenfield, N. J. Using circular dichroism collected as a function of temperature to determine the thermodynamics of protein unfolding and binding interactions. Nature Protocols 1, 2527–2535 (2007)
Mavrakis, K. J. et al. A cooperative microRNA-tumor suppressor gene network in acute T-cell lymphoblastic leukemia (T-ALL). Nature Genet. 43, 673–678 (2011)
Shochat, C. et al. Gain-of-function mutations in interleukin-7 receptor-α (IL7R) in childhood acute lymphoblastic leukemias. J. Exp. Med. 208, 901–908 (2011)
Zuurbier, L. et al. The significance of PTEN and AKT aberrations in pediatric T-cell acute lymphoblastic leukemia. Haematologica 97, 1405–1413 (2012)
Menten, B. et al. arrayCGHbase: an analysis platform for comparative genomic hybridization microarrays. BMC Bioinformatics 6, 124 (2005)
Galicia-Vázquez, G., Cencic, R., Robert, F., Agenor, A. Q. & Pelletier, J. A cellular response linking eIF4AI activity to eIF4AII transcription. RNA 18, 1373–1384 (2012)
Acknowledgements
We thank the members of A.L.W.’s thesis committee: N. Rosen, A. M. Brown and S. W. Lowe. For reagents and advice we thank J. T. Barata, W. S. Pear, R. Cencic, S. Shuman, J. Cools, A. A. Ferrando, C. S. Fraser, N. J. Lajkiewicz, A. Luz, J. F. Glickman, C. Y. Park, P. Yellen, A. Heguy, K. Huberman and A. Viale. H.-G.W. is a Scholar of the Leukemia and Lymphoma Society. This research was supported by National Cancer Institute R01-CA142798-01 (H.-G.W.), the Leukemia Research Foundation (H.-G.W.), the Experimental Therapeutics Center (H.-G.W.), the American Cancer Society 10284 (H.-G.W.), European Union grant no. PITN-GA-2012-316861 (Y.Z.), the Fund for Scientific Research FWO Flanders (J.V.d.M. and P.R.), grants G.0198.08 and G.0869.10N (F.S.), the GOA-UGent 12051203 (F.S.), Stichting tegen Kanker (F.S.), the Belgian Program of Interuniversity Poles of Attraction (F.S.), the Belgian Foundation Against Cancer (F.S.), the American Cancer Society PF-11-077-01-CDD (C.M.R.), the Lymphoma Research Foundation (J.H.S.), National Institutes of Health grants GM-067041 and GM-073855 (J.A.P.), and the Canadian Institutes of Health Research MOP-10653 (J.P.).
Author information
Authors and Affiliations
Contributions
A.L.W. performed in vivo and treatment studies; K.S. performed ribosome footprinting and RNA structure studies; Y.Z. and P.D. analysed footprint data; V.K.R., V.R.S., K.J.M., M.J., J.E.R., J.H.S., C.Z., J.T.-F. and M.A.K. contributed to experiments; C.M.R. prepared (±)-CR-31-B; E.d.S. directed murine drug toxicity experiments; J.V.d.M., P.R. and F.S. generated genomic data on T-ALL; J.A.P. Jr and J.P. advised on all aspects of the study; G.R. supervised computational analyses; H.-G.W. designed the study and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Translational activation in T-ALL.
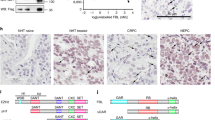
a–c, Diagram of mutations in human T-ALL affecting PTEN (a), IL7R (b) and NOTCH1 (c). d, Immunoblots of lysates from ICN-driven murine leukaemia with the additional indicated construct, probed as indicated. e, Immunoblots of lysates from 3T3 cells with empty vector or sheIF4A and probed as indicated. f, Representative fluorescence-activated cell sorting (FACS) profiles measuring levels of the indicated markers in murine leukaemia. g, Surface marker expression on murine leukaemic cells of the indicated genotype (plus and minus signs indicate <50% or ≥50% positive cells). h, Lysates of murine leukaemia expressing ICN and either empty vector or eIF4A1 and probed as indicated. i, Representative histology detailing the pathological appearance of murine T-ALLs harbouring the indicated genes and stained as indicated.
Extended Data Figure 2 Silvestrol and the synthetic analogue (±)-CR-31-B are effective against T-ALL.
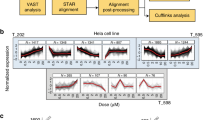
a, b, Chemical structure of silvestrol (a) and (±)-CR-31-B (b). c, Dual luciferase reporter assay, shown are relative levels of each firefly (cap-dependent) and Renilla (IRES-dependent) luciferase upon treatment with silvestrol or (±)-CR-31-B. Mean and s.d. are shown, n = 3 biological replicates. d, IC50 values for silvestrol and (±)-CR-31-B in a panel of human T-ALL primary patient samples and cell lines. Mean and s.d. are shown, n = 4 biological replicates. e, Silvestrol’s effect on murine T-ALLs with the indicated genetic lesions; curves are mean of triplicates and differences between the genotypes did not reach significance. f, Kaplan–Meier analysis showing time to leukaemia development after systemic transplantation of MOHITO cells in Balb/c mice followed by treatment on 7 consecutive days (treatments are indicated by red arrows) with either silvestrol (0.5 mg kg−1, red line, n = 5) or vehicle (black line, n = 5). g, KOPT-K1 xenograft studies. Shown is the tumour volume during and after systemic treatment with (±)-CR-31-B or vehicle (i.p. injection, 0.2 mg kg−1 on days indicated by red arrows). Mean and s.d. are shown, n = 6 biological replicates. h, Tumour volume upon i.p. treatment with vehicle or silvestrol (0.5 mg kg−1 on days indicated by red arrows). Mean and s.d. are shown, n = 3 biological replicates.
Extended Data Figure 3 Toxicity studies with (±)-CR-31-B.
a–j, Mean and standard deviation are shown, n = 2 biological replicates. a, Animal weights during and after (±)-CR-31-B treatment (i.p. injection, 0.2 mg kg−1 on days indicated by red arrows). Red, CR; black, vehicle. b–d, Counts of white blood cells (b), red cells (c) and platelets (d) 14 days after cessation of (±)-CR-31-B treatment. Blue lines indicate the species- and strain-specific reference range. NS, not significant, n = 2 biological replicates. e, Representative histology of gastrointestinal tract (small intestine) on the indicated days during (n = 4) and after (n = 2) (±)-CR-31-B treatment. f–j, Serum levels of alanine aminotransferase (ALT) (f), aspartate transaminase (AST) (g), albumin (h), total bilirubin (i) and creatinine (j) 2 weeks after cessation of treatment with (±)-CR-31-B or vehicle. Blue lines indicate the species- and strain-specific reference range. NS, not significant.
Extended Data Figure 4 Ribosome profiling quality control data and effects on translation.
a, b, Read counts by length of mapped sequence before and after filtering rRNA, linker reads, non-coding RNAs, short mapped sequences (‘noisy’ reads; see text and Methods for details). n = 2 biological replicates. c, d, Read length frequency histograms and mapping analysis of ribosome footprint data after quality control filtering for vehicle-treated cells (c) or silvestrol-treated cells (d). n = 2 biological replicates. e, Silvestrol-induced changes in total RNA (log2 fold change RPKM) and ribosome-protected RNA (RF). n = 2 biological replicates. f, Histogram of the ribosome footprint intensity of all genes (measured as unique RPM) for silvestrol- and vehicle-treated cells, indicating that silvestrol-affected mRNAs were broadly distributed (see text for details). n = 2 biological replicates. g, Mean fluorescence intensity of incorporated AHA in newly synthesized proteins in KOPT-K1 cells treated with vehicle (DMSO), silvestrol (Silv.; 25 nM), or cycloheximide (CHX; 100 nM) for the indicated time period. n = 3 biological replicates. h, Polyribosome profiles of silvestrol- (25 nM) or vehicle (DMSO)-treated KOPT-K1 cells showing OD254 nm absorption across the ribosome-containing fractions. n = 3 biological replicates. i, Ribosome density for transcripts across control and silvestrol samples (ribosomal footprint (RF) reads per RPKM). n = 2 biological replicates. The correlation (R2 = 0.94) indicates a broad effect on translation and transcripts with significantly differential changes in ribosome density are indicated as red and blue dots. j, Length comparison of 5′ UTRs of TE up genes and a background gene set. Asterisks indicate mean, n = 2 biological replicates. k, Percentage of TE up genes and background genes containing the indicated sequence motifs. *P < 0.001, n = 2 biological replicates.
Extended Data Figure 5 Analysis of genes with differential ribosomal distribution rDiff.
a, Representation of ribosome coverage for 826 transcripts with significant changes in distribution between silvestrol (red) and vehicle (black)—corresponding to the rDiff positive gene list after filtering out genes with 5′ UTR length <20 nucleotides. Both ribosome footprint coverage and transcript length are normalized for comparison. Translation start and stop sites are indicated by blue lines, n = 826. b, c, Ribosomal distribution plots, as in a, showing how silvestrol affects ribosome distribution in all TE up genes (b) (n = 182 after filtering out genes with 5′ UTR length <20 nucleotides) and all TE down genes (c) (n = 276 after filtering out genes with 5′ UTR length <20 nucleotides). d, Length comparison of 5′ UTRs of genes with significantly altered ribosomal distribution (rDiff positive, red) and background genes (black). Asterisks indicate mean value, n = 826. e, Percentage of rDiff-positive genes and background genes containing the indicated sequence motifs. *P < 0.05, n = 2 biological replicates. f, g, The rDiff-positive genes are enriched for the indicated 12-nucleotide (f) and 9-nucleotide (g) consensus motifs.
Extended Data Figure 6 Circular dichroism and characterization of eIF4A.
a, Bar graph indicating the prevalence of each sequence motif from the rDiff data set and its predicted likelihood of forming G-quadruplex (GQ) structures (red). b, Circular dichroism spectra scan of 9-nucleotide (mer) motif with a 5-nucleotide flank taken from the actual 5′ UTR of the indicated genes, folded in KCl. c, Circular dichroism spectra scan of 12-nucleotide motif and mutant folded in sodium phosphate buffer without KCl; note the y-axis scale. d, Relative amounts of Renilla luciferase (normalized to firefly), expressed from the G-quadruplexes (red bars) or control construct (black bars), treated with 8 nM pateamine A (Pat. A) or 50 nM hippuristanol (Hipp.) for 24 h. *P < 0.05, n = 3 biological replicates and n = 2 technical replicates. e, Analysis of mRNA expression of the indicated RNA helicases in normal T cells and T-ALL cells. *P < 0.05, n = 57 biological replicates32. f, Relative amounts of Renilla luciferase expressed from the G-quadruplex construct in 3T3 cells and normalized to IRES/firefly with either empty vector or the indicated genes, treated with silvestrol (25 nM) for 24 h. Mean and s.d. are shown, n = 3 biological replicates, n = 2 technical replicates. NS, not significant.
Extended Data Figure 7 Silvestrol-sensitive transcripts.
a, Distribution of ribosomal footprints for the indicated genes, n = 2 biological replicates. Red, silvestrol; black, vehicle; purple dots, 9-nucleotide (mer) motifs; blue dots, 12-nucleotide motif. b, Gene ontology classification for genes in the TE down group with G-quadruplex (GQ), 12-nucleotide and 9-nucleotide motifs. c, Venn diagram illustrating the overlap between TE and/or rDiff genes and reported superenhancers in T-ALL cell lines34.
Extended Data Figure 8 Immunoblots and mRNA expression.
a, Lysates from human T-ALL lines treated with (±)-CR-31-B (25 nM, 24 h) and probed as indicated. b, Lysates from Jurkat cells treated with escalating doses of silvestrol and probed as indicated. c, mRNA levels for the indicated genes treated with vehicle (DMSO, black) or silvestrol (red, 25 nM) for 45 min. Mean and s.d. are shown, n = 2 biological replicates. d–g, Immunoblots of lysates from murine T-ALL cells expressing either vector control or IRES-MYC (d), IRES-CCND3 T283A (e), IRES-ICN (f) or IRES-BCL2 (g) and probed as indicated.
Supplementary information
Supplementary Table 1
Summary of re-sequencing data showing mutation data observed in 36 T-ALL samples. (XLS 41 kb)
Supplementary Table 2
The multi-panel table provides results of a detailed analysis of the toxic effects of CR treatment in non-tumour bearing c57/B6 mice. a) Body and organ weights; b) Individual haematology; c) Bone marrow and spleen cytology; d) Individual chemistry. (XLS 51 kb)
Supplementary Table 3
a) Genes with decreased Translational Efficiency (TE down); b) Genes with increased Translational Efficiency (TE up); c) Background genes with no change in Translational Efficiency (TE background); d) Complete list of genes that showed a significant change in RF distribution across the length of their transcript (rDiff positive gene set); e) Genes that are both TE down and rDiff positive. (XLSX 389 kb)
Supplementary Table 4
a-b) Complete lists of TE Down genes that harbour the 12-mer (a) or 9-mer (b) in their 5’UTRs; c-d) Complete lists of rDiff genes that harbour the 12-mer (c) or 9-mer (d) in their 5’UTRs; Prevalence of the TE Down 12-mer motif; e) TE Down genes with one or more predicted G-quadruplex structures; f) rDiff genes with one or more predicted G-quadruplex structures; g-h) Overlap of TE Down 12-mer motif (g) or 9-mer motif (h) with predicted G-quadruplexes. i-j) Overlap of rDiff 12-mer motif (i) or 9-mer motif (j) with predicted G-quadruplexes; k) Nucleotide-level depiction of the overlap between 9-mer motifs and G-quadruplexes. The motif is located at the positions marked X in the sequence. The middle row shows the structure. + represents that it is part of a G-quadruplex, ) ( represents a bond with another nucleotide. (XLSX 107 kb)
Supplementary Table 5
RNA oligos used for Circular Dichroism (CD) and thermal denaturation analysis. Note: 12-mers or 9-mers are shown in red. (XLSX 32 kb)
Supplementary Table 6
Overlap of TE down or rDiff genes with reported super-enhancer associated genes in the T-ALL cell lines DND41, JURKAT, or RPMI-840234. '+' indicates presence of one or more 12-mers, 9-mers, or predicted G-quadruplexes in the 5’UTR. (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Wolfe, A., Singh, K., Zhong, Y. et al. RNA G-quadruplexes cause eIF4A-dependent oncogene translation in cancer. Nature 513, 65–70 (2014). https://doi.org/10.1038/nature13485
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13485
This article is cited by
-
Ribosome profiling: a powerful tool in oncological research
Biomarker Research (2024)
-
KRAS, MYC, and ARF6: inseparable relationships cooperatively promote cancer malignancy and immune evasion
Cell Communication and Signaling (2023)
-
Bioactive nutraceuticals as G4 stabilizers: potential cancer prevention and therapy—a critical review
Naunyn-Schmiedeberg's Archives of Pharmacology (2023)
-
5′-tiRNA-Gln inhibits hepatocellular carcinoma progression by repressing translation through the interaction with eukaryotic initiation factor 4A-I
Frontiers of Medicine (2023)
-
Identification of eIF6 as a prognostic factor that drives tumor progression and predicts arsenic trioxide efficacy in lung adenocarcinoma
Molecular Biology Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.