Abstract
Ligation of tRNAs with their cognate amino acids, by aminoacyl-tRNA synthetases, establishes the genetic code. Throughout evolution, tRNAAla selection by alanyl-tRNA synthetase (AlaRS) has depended predominantly on a single wobble base pair in the acceptor stem, G3•U70, mainly on the kcat level. Here we report the crystal structures of an archaeal AlaRS in complex with tRNAAla with G3•U70 and its A3•U70 variant. AlaRS interacts with both the minor- and the major-groove sides of G3•U70, widening the major groove. The geometry difference between G3•U70 and A3•U70 is transmitted along the acceptor stem to the 3′-CCA region. Thus, the 3′-CCA region of tRNAAla with G3•U70 is oriented to the reactive route that reaches the active site, whereas that of the A3•U70 variant is folded back into the non-reactive route. This novel mechanism enables the single wobble pair to dominantly determine the specificity of tRNA selection, by an approximate 100-fold difference in kcat.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
20 November 2018
In Fig. 1b of this Article, a U was inadvertently inserted after G15 in the D loop. The original Article has not been corrected.
References
Giegé, R., Sissler, M. & Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 26, 5017–5035 (1998)
Beuning, P. J. & Musier-Forsyth, K. Transfer RNA recognition by aminoacyl-tRNA synthetases. Biopolymers 52, 1–28 (1999)
Pütz, J., Puglisi, J. D., Florentz, C. & Giegé, R. Additive, cooperative and anti-cooperative effects between identity nucleotides of a tRNA. EMBO J. 12, 2949–2957 (1993)
Weygand-Durasević, I., Rogers, M. J. & Söll, D. Connecting anticodon recognition with the active site of Escherichia coli glutaminyl-tRNA synthetase. J. Mol. Biol. 240, 111–118 (1994)
Rogers, M. J., Adachi, T., Inokuchi, H. & Söll, D. Functional communication in the recognition of tRNA by Escherichia coli glutaminyl-tRNA synthetase. Proc. Natl Acad. Sci. USA 91, 291–295 (1994)
Uter, N. T. & Perona, J. J. Long-range intramolecular signaling in a tRNA synthetase complex revealed by pre-steady-state kinetics. Proc. Natl Acad. Sci. USA 101, 14396–14401 (2004)
Hou, Y. M. & Schimmel, P. A simple structural feature is a major determinant of the identity of a transfer RNA. Nature 333, 140–145 (1988)
McClain, W. H. & Foss, K. Changing the identity of a tRNA by introducing a G-U wobble pair near the 3′ acceptor end. Science 240, 793–796 (1988)
de Duve, C. Transfer RNAs: the second genetic code. Nature 333, 117–118 (1988)
Park, S. J., Hou, Y. M. & Schimmel, P. A single base pair affects binding and catalytic parameters in the molecular recognition of a transfer RNA. Biochemistry 28, 2740–2746 (1989)
Hou, Y. M. & Schimmel, P. Evidence that a major determinant for the identity of a transfer RNA is conserved in evolution. Biochemistry 28, 6800–6804 (1989)
Shi, J. P., Francklyn, C., Hill, K. & Schimmel, P. A nucleotide that enhances the charging of RNA minihelix sequence variants with alanine. Biochemistry 29, 3621–3626 (1990)
Beuning, P. J., Gulotta, M. & Musier-Forsyth, K. Atomic group “mutagenesis” reveals major groove fine interactions of a tRNA synthetase with an RNA helix. J. Am. Chem. Soc. 119, 8397–8402 (1997)
Musier-Forsyth, K. & Schimmel, P. Functional contacts of a transfer RNA synthetase with 2′-hydroxyl groups in the RNA minor groove. Nature 357, 513–515 (1992)
Beuning, P. J. et al. Efficient aminoacylation of the tRNAAla acceptor stem: dependence on the 2:71 base pair. RNA 8, 659–670 (2002)
Francklyn, C., Shi, J. P. & Schimmel, P. Overlapping nucleotide determinants for specific aminoacylation of RNA microhelices. Science 255, 1121–1125 (1992)
Francklyn, C. & Schimmel, P. Aminoacylation of RNA minihelices with alanine. Nature 337, 478–481 (1989)
Varani, G. & McClain, W. H. The ĠU wobble base pair. A fundamental building block of RNA structure crucial to RNA function in diverse biological systems. EMBO Rep. 1, 18–23 (2000)
Musier-Forsyth, K. et al. Specificity for aminoacylation of an RNA helix: an unpaired, exocyclic amino group in the minor groove. Science 253, 784–786 (1991)
Chang, K. Y., Varani, G., Bhattacharya, S., Choi, H. & McClain, W. H. Correlation of deformability at a tRNA recognition site and aminoacylation specificity. Proc. Natl Acad. Sci. USA 96, 11764–11769 (1999)
Ueda, H. et al. X-ray crystallographic conformational study of 5′-O-[N-(L-alanyl)-sulfamoyl]adenosine, a substrate analogue for alanyl-tRNA synthetase. Biochim. Biophys. Acta 1080, 126–134 (1991)
Fukunaga, R. & Yokoyama, S. Crystallization and preliminary X-ray crystallographic study of alanyl-tRNA synthetase from the archaeon Archaeoglobus fulgidus. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 63, 224–228 (2007)
Dignam, J. D. et al. Allosteric interaction of nucleotides and tRNAala with E. coli alanyl-tRNA synthetase. Biochemistry 50, 9886–9900 (2011)
Jasin, M., Regan, L. & Schimmel, P. Modular arrangement of functional domains along the sequence of an aminoacyl tRNA synthetase. Nature 306, 441–447 (1983)
Beebe, K., Ribas De Pouplana, L. & Schimmel, P. Elucidation of tRNA-dependent editing by a class II tRNA synthetase and significance for cell viability. EMBO J. 22, 668–675 (2003)
Swairjo, M. A. et al. Alanyl-tRNA synthetase crystal structure and design for acceptor-stem recognition. Mol. Cell 13, 829–841 (2004)
Guo, M. et al. Paradox of mistranslation of serine for alanine caused by AlaRS recognition dilemma. Nature 462, 808–812 (2009)
Naganuma, M., Sekine, S., Fukunaga, R. & Yokoyama, S. Unique protein architecture of alanyl-tRNA synthetase for aminoacylation, editing, and dimerization. Proc. Natl Acad. Sci. USA 106, 8489–8494 (2009)
Sokabe, M. et al. The structure of alanyl-tRNA synthetase with editing domain. Proc. Natl Acad. Sci. USA 106, 11028–11033 (2009)
Eriani, G., Delarue, M., Poch, O., Gangloff, J. & Moras, D. Partition of tRNA synthetases into two classes based on mutually exclusive sets of sequence motifs. Nature 347, 203–206 (1990)
Cusack, S. Eleven down and nine to go. Nature Struct. Biol. 2, 824–831 (1995)
Ruff, M. et al. Class II aminoacyl transfer RNA synthetases: crystal structure of yeast aspartyl-tRNA synthetase complexed with tRNAAsp. Science 252, 1682–1689 (1991)
Biou, V., Yaremchuk, A., Tukalo, M. & Cusack, S. The 2.9 Å crystal structure of T. thermophilus seryl-tRNA synthetase complexed with tRNASer. Science 263, 1404–1410 (1994)
Ramos, A. & Varani, G. Structure of the acceptor stem of Escherichia coli tRNAAla: role of the G3·U70 base pair in synthetase recognition. Nucleic Acids Res. 25, 2083–2090 (1997)
Mueller, U., Schübel, H., Sprinzl, M. & Heinemann, U. Crystal structure of acceptor stem of tRNAAla from Escherichia coli shows unique ĠU wobble base pair at 1.16 Å resolution. RNA 5, 670–677 (1999)
Miller, W. T., Hou, Y. M. & Schimmel, P. Mutant aminoacyl-tRNA synthetase that compensates for a mutation in the major identity determinant of its tRNA. Biochemistry 30, 2635–2641 (1991)
Guo, M. et al. The C-Ala domain brings together editing and aminoacylation functions on one tRNA. Science 325, 744–747 (2009)
Zhang, C. M., Perona, J. J., Ryu, K., Francklyn, C. & Hou, Y. M. Distinct kinetic mechanisms of the two classes of Aminoacyl-tRNA synthetases. J. Mol. Biol. 361, 300–311 (2006)
Guth, E., Connolly, S. H., Bovee, M. & Francklyn, C. S. A substrate-assisted concerted mechanism for aminoacylation by a class II aminoacyl-tRNA synthetase. Biochemistry 44, 3785–3794 (2005)
McClain, W. H., Chen, Y. M., Foss, K. & Schneider, J. Association of transfer RNA acceptor identity with a helical irregularity. Science 242, 1681–1684 (1988)
Gabriel, K., Schneider, J. & McClain, W. H. Functional evidence for indirect recognition of G•U in tRNAAla by alanyl-tRNA synthetase. Science 271, 195–197 (1996)
McClain, W. H., Jou, Y. Y., Bhattacharya, S., Gabriel, K. & Schneider, J. The reliability of in vivo structure-function analysis of tRNA aminoacylation. J. Mol. Biol. 290, 391–409 (1999)
Nakagawa, H. et al. Crystallographic and mutational studies on the tRNA thiouridine synthetase TtuA. Proteins 81, 1232–1244 (2013)
Yamashita, S. et al. Structures of the first and second double-stranded RNA-binding domains of human TAR RNA-binding protein. Protein Sci. 20, 118–130 (2011)
Collaborative Computational Project, No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Brunger, A. T. Version 1.2 of the Crystallography and NMR system. Nature Protocols 2, 2728–2733 (2007)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Sankaranarayanan, R. et al. The structure of threonyl-tRNA synthetase-tRNAThr complex enlightens its repressor activity and reveals an essential zinc ion in the active site. Cell 97, 371–381 (1999)
Yaremchuk, A., Tukalo, M., Grøtli, M. & Cusack, S. A succession of substrate induced conformational changes ensures the amino acid specificity of Thermus thermophilus prolyl-tRNA synthetase: comparison with histidyl-tRNA synthetase. J. Mol. Biol. 309, 989–1002 (2001)
Goldgur, Y. et al. The crystal structure of phenylalanyl-tRNA synthetase from Thermus thermophilus complexed with cognate tRNAPhe. Structure 5, 59–68 (1997)
Nozawa, K. et al. Pyrrolysyl-tRNA synthetase-tRNAPyl structure reveals the molecular basis of orthogonality. Nature 457, 1163–1167 (2009)
Fukunaga, R. & Yokoyama, S. Structural insights into the first step of RNA-dependent cysteine biosynthesis in archaea. Nature Struct. Mol. Biol. 14, 272–279 (2007)
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25, 4876–4882 (1997)
Robertus, J. D. et al. Structure of yeast phenylalanine tRNA at 3 Å resolution. Nature 250, 546–551 (1974)
Kim, S. H. et al. Three-dimensional tertiary structure of yeast phenylalanine transfer RNA. Science 185, 435–440 (1974)
Sekine, S. et al. ATP binding by glutamyl-tRNA synthetase is switched to the productive mode by tRNA binding. EMBO J. 22, 676–688 (2003
Acknowledgements
We are grateful to the staff of the beam line AR-NW12A at the Photon Factory and the beam lines BL32XU and BL41XU at SPring-8 for their assistance during data collection. This work was supported by the Targeted Proteins Research Program, and the Platform for Drug Discovery, Informatics, and Structural Life Science from the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan. This work was also supported by a Grant-in-Aid for Scientific Research (No. 20247008) from the Japan Society for the Promotion of Science and MEXT, and the Global COE Program (Integrative Life Science Based on the Study of Biosignaling Mechanisms) from MEXT. Research support was also provided by National Institutes of Health grants GM015539 and GM023562 to P.S. and NS085092 to X.-L.Y.
Author information
Authors and Affiliations
Contributions
M.N., S.S. and S.Y. designed the research. M.N. and S.S. performed the structural analysis. M.N., Y.E.C., M.G., X.-L.Y., H.G., and Y.-M.H. performed the enzyme kinetics analyses. M.N., S.S., P.S., and S.Y. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Conformational differences between the tRNA-bound and tRNA-free subunits of A. fulgidus AlaRS.
a, Rotation of the C-terminal domain. The C-terminal domain of subunit A is inclined ∼28° towards the outer corner of the L-shaped tRNA, relative to subunit B, and thus the globular subdomain stacks with the outer corner of the tRNA. b, c, Mid2 reorientation. α13 is straight with tRNAAla (b) and kinked without tRNAAla (c).
Extended Data Figure 2 The positions of tRNAs relative to the aminoacylation domains of AlaRS and other class II aaRSs depend on the specific spatial arrangements of their tRNA-binding domain(s).
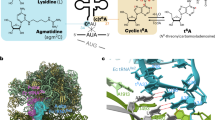
a–d, Class II aaRS dimers bound to tRNAs. A. fulgidus AlaRS, yeast AspRS, E. coli ThrRS, and T. thermophilus SerRS are shown as surface models, with their cognate tRNAs displayed as tube models. For each aaRS, one of the two tRNA-binding aminoacylation domains (ADs) and its bound tRNA are coloured green and yellow, respectively, whereas the other AD and its bound tRNA are coloured light green and grey, respectively. In each of the aaRSs, except for A. fulgidus AlaRS, the tRNA-binding domain (RBD) corresponding to the green AD is coloured red, and that of the other subunit is coloured raspberry. A. fulgidus AlaRS has two RBDs (RBD1 and RBD2), coloured in the same manner as the RBDs of the other aaRSs. e, The position of tRNAAla relative to the AD of AlaRS (displayed as yellow and green surface models, respectively) is completely different from the well conserved positions of tRNAAsp, tRNAThr, tRNAPro, tRNASer, tRNAPhe, tRNAPyl, and tRNACys (brown tube models) shown in the same positions relative to the ADs of AspRS32, ThrRS53, ProRS54, SerRS33, PheRS55, PylRS56, and SepRS57, respectively (PDB IDs: 1ASY, 1QF6, 1H4Q, 1SER, 1EIY, 2ZNI and 2DU3, respectively).
Extended Data Figure 3 Summary of tRNA–protein interactions.
a, Schematics of tRNA-protein interactions. The tRNA backbones and bases are coloured as indicated. The G3•U70 base pair and the anticodon bases are highlighted by red and pink squares, respectively. The hydrogen bonds between the nucleotides and the AlaRS main chain are indicated with blue arrows. Those between the nucleotides and the AlaRS side chains are depicted with magenta arrows. The stacking interactions are indicated with black dashed lines. b, The tRNAAla interaction with the α11 helix. The α11 helix interacts with the backbone phosphate groups of the 3′ half of the acceptor stem. The side chains of Tyr 364 and Arg 371 hydrogen bond with C72 and C71, respectively. The side chain of Arg 375 hydrogen bonds with U70. c, Interactions between the tRNAAla outer corner region and the globular subdomain. The conserved Gly-rich segment 870KGSGGGR878 stacks with the hinge region of the T- and D-loops. The side chain of Arg 876 stacks and hydrogen bonds with the base and the backbone phosphate group, respectively, of G20. The side chain of Asn 823 hydrogen bonds with the backbone phosphate group of G20. d, The tRNAAla interactions with the α13 helix. The α13 helix interacts with the backbone phosphate groups of the D arm (U12-C13-A14-G15) and that of U70. The side chain of Thr 422 in α13 hydrogen bonds with U70. The side chains of Arg 428 and Arg 432 hydrogen bond with C13. The side chain of Arg 436 hydrogen bonds with A14 and G15.
Extended Data Figure 4 Nucleotide interactions with the conserved residues of AlaRS.
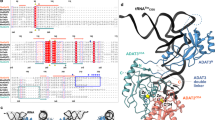
a, b, |Fo–Fc| omit electron density maps contoured at 3σ, superimposed on the refined models of G3•U70 (a) and A3•U70 (b). c–g, The interactions of G1•C72 (c), G2•C71 (d), C4•G69 (e), U5•A68 (f), and A73 (g) with AlaRS. The amino-acid side chains and the nucleotides are shown as grey and beige stick models, respectively. Hydrogen bonds are indicated with dashed lines. h, A73 stacking with G1 and C72. i, Alignments of the AlaRS sequences. A total of 249 AlaRS sequences were initially aligned with the ClustalW program58, and then were manually adjusted based on the structural information. The sequences of the A. fulgidus, P. horikoshii, Homo sapiens, E. coli, and A. aeolicus AlaRSs are shown. The highly-conserved residues are boxed in red. The residues that are involved the acceptor stem interactions in A. fulgidus AlaRS are boxed in blue. Secondary structure information is shown above the A. fulgidus sequence. The full-sequence alignments were reported previously28.
Extended Data Figure 5 The routes of the CCA regions of tRNAAla/GU and tRNAAla/AU.
a–c, tRNAAla/GU. d–f, tRNAAla/AU. tRNA is shown in stick models (a, d), cartoon models (b, e), and surface models (c, f). The |Fo–Fc| omit electron density maps contoured at 3σ are superimposed on the refined models of the CCA regions of tRNAAla/GU (a) and tRNAAla/AU (d). The CCA region of tRNAAla/GU is in the reactive route (b, c), while that of tRNAAla/AU is in the non-reactive route (e, f). The CCA region runs under the linker (b, c, e, f). g, h, Heterologous superimpositions of the |2Fo–Fc| electron density maps contoured at 1σ of tRNAAla/AU (cyan, g) and tRNAAla/GU (pink, h) on the stick models of the structures of the acceptor stems and the CCA regions of tRNAAla/GU (pink, g) and tRNAAla/AU (cyan, h), respectively, which clearly demonstrate the conformational difference between the two tRNAs. i, The |Fo–Fc| omit electron density maps for tRNAAla/AU, contoured at 2.5σ, superimposed on the refined models of the C74 s of tRNAAla/GU (pink) and tRNAAla/AU (cyan). j, k, The similarity in the spatial positions between E220/193GGG195 of A. fulgidus AlaRS (j) and D174 of E. coli AlaRS (k), marked in red on the surface models of the aminoacylation and tRNA-recognition domains with the tube models of tRNAAla (the crystal structure and the docking model, respectively). l, Misorientation of the CCA region of tRNAAla/WC on the D450A mutant of AlaRS.
Extended Data Figure 6 Major-groove widening of the tRNAAla acceptor stem.
a, The phosphate–phosphate distances of G1–C66, C2–C65, G3–G64, G4–C63, A5–C62, and U6–C61 in tRNAPhe (refs. 59, 60). b, The phosphate–phosphate distances of G1–C66, G2–C65, G3–G64, C4–C63, U5–C62, and C6–C61 in the wild-type tRNAAla (tRNAAla/GU) complexed with AlaRS. The phosphate–phosphate distances between G1–C66, G2–C65, and G3–G64 across the major groove of tRNAAla/GU are extended by 6.3, 2.9, and 0.9 Å, respectively, as compared to those of tRNAPhe. Thus, AlaRS clearly widens the major groove of the acceptor stem of tRNAAla/GU. c, The phosphate–phosphate distances of G1–C66, G2–C65, A3–G64, C4–C63, U5–C62, and C6–C61 in the variant tRNAAla/AU complexed with AlaRS. The major-groove of the tRNAAla/AU acceptor stem is slightly wider than that in the wild-type complex structure. Therefore, the replacement of G3•U70 by A3•U70 does not seem to impair the major-groove widening by AlaRS. d, The phosphate–phosphate distances of U1–U66, C2–G65, C3–C64, G4–C63, U5–C62, and G6–C61 in tRNAAsp bound to AspRS32 (PDB ID: 1ASY). e, The phosphate–phosphate distances of G1–U66, C2–U65, C3–A64, G4–U63, A5–C62, and U6–C61 in tRNAThr bound to ThrRS53 (PDB ID: 1QF6). f, The phosphate–phosphate distances of G1–U66, G2–C65, C3–C64, C4–C63, C5–C62, and C6–C61 in tRNAGlu bound to GluRS61 (PDB ID: 1G59). g, The AlaRS-induced widening of the tRNAAla acceptor stem. Without widening, AlaRS clashes with A67-G68. h, Widening of the acceptor stems of tRNAAla/GU (pink tube models) and tRNAAla/AU (cyan tube models).
Extended Data Figure 7 Torsion angles of nucleotides in the acceptor stems of tRNAAla/GU and tRNAAla/AU.
a, b, Tables for torsion angles of nucleotides in the acceptor stems of tRNAAla/GU (a) and tRNAAla/AU (b). c, Conformational changes of G3 and G69 relative to the A-form RNAs.
Extended Data Figure 8 tRNAAla/AU inhibits aminoacylation of human tRNAAla/GU by human AlaRS.
In vitro transcribed human tRNAAla/AU was added to aminoacylation reactions containing 2 μM human tRNAAla and 50 nM human AlaRS, in pH 6.0 reaction buffer at room temperature. The dose-dependent inhibition of aminoacylation of tRNAAla/GU by excess tRNAAla/AU shows that the ultimate discrimination against A3•U70 is via kcat.
Rights and permissions
About this article
Cite this article
Naganuma, M., Sekine, Si., Chong, Y. et al. The selective tRNA aminoacylation mechanism based on a single G•U pair. Nature 510, 507–511 (2014). https://doi.org/10.1038/nature13440
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13440
This article is cited by
-
Relaxed sequence constraints favor mutational freedom in idiosyncratic metazoan mitochondrial tRNAs
Nature Communications (2020)
-
Chimeric design of pyrrolysyl-tRNA synthetase/tRNA pairs and canonical synthetase/tRNA pairs for genetic code expansion
Nature Communications (2020)
-
G:U-Independent RNA Minihelix Aminoacylation by Nanoarchaeum equitans Alanyl-tRNA Synthetase: An Insight into the Evolution of Aminoacyl-tRNA Synthetases
Journal of Molecular Evolution (2020)
-
ANKRD16 prevents neuron loss caused by an editing-defective tRNA synthetase
Nature (2018)
-
Utilization of rare codon-rich markers for screening amino acid overproducers
Nature Communications (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.