Abstract
Spontaneous changes in the reading frame of translation are rare (frequency of 10−3 to 10−4 per codon)1, but can be induced by specific features in the messenger RNA (mRNA). In the presence of mRNA secondary structures, a heptanucleotide ‘slippery sequence’ usually defined by the motif X XXY YYZ, and (in some prokaryotic cases) mRNA sequences that base pair with the 3′ end of the 16S ribosomal rRNA (internal Shine–Dalgarno sequences), there is an increased probability that a specific programmed change of frame occurs, wherein the ribosome shifts one nucleotide backwards into an overlapping reading frame (−1 frame) and continues by translating a new sequence of amino acids2,3. Despite extensive biochemical and genetic studies, there is no clear mechanistic description for frameshifting. Here we apply single-molecule fluorescence to track the compositional and conformational dynamics of individual ribosomes at each codon during translation of a frameshift-inducing mRNA from the dnaX gene in Escherichia coli. Ribosomes that frameshift into the −1 frame are characterized by a tenfold longer pause in elongation compared to non-frameshifted ribosomes, which translate through unperturbed. During the pause, interactions of the ribosome with the mRNA stimulatory elements uncouple EF-G catalysed translocation from normal ribosomal subunit reverse-rotation, leaving the ribosome in a non-canonical intersubunit rotated state with an exposed codon in the aminoacyl-tRNA site (A site). tRNALys sampling and accommodation to the empty A site and EF-G action either leads to the slippage of the tRNAs into the −1 frame or maintains the ribosome into the 0 frame. Our results provide a general mechanistic and conformational framework for −1 frameshifting, highlighting multiple kinetic branchpoints during elongation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
20 August 2014
Extended Data Fig. 4 and the legend for Extended Data Fig. 3 have been updated.
References
Jenner, L. B., Demeshkina, N., Yusupova, G. & Yusupov, M. Structural aspects of messenger RNA reading frame maintenance by the ribosome. Nature Struct. Mol. Biol. 17, 555–560 (2010)
Tinoco, I., Jr, Kim, H. K. & Yan, S. Frameshifting dynamics. Biopolymers 99, 1147–1166 (2013)
Tsuchihashi, Z. & Kornberg, A. Translational frameshifting generates the gamma subunit of DNA polymerase III holoenzyme. Proc. Natl Acad. Sci. USA 87, 2516–2520 (1990)
Plant, E. P. et al. The 9-Å solution: how mRNA pseudoknots promote efficient programmed −1 ribosomal frameshifting. RNA 9, 168–174 (2003)
Baranov, P. V., Gesteland, R. F. & Atkins, J. F. P-site tRNA is a crucial initiator of ribosomal frameshifting. RNA 10, 221–230 (2004)
Horsfield, J. A., Wilson, D. N., Mannering, S. A., Adamski, F. M. & Tate, W. P. Prokaryotic ribosomes recode the HIV-1 gag-pol-1 frameshift sequence by an E/P site post-translocation simultaneous slippage mechanism. Nucleic Acids Res. 23, 1487–1494 (1995)
Lopinski, J. D., Dinman, J. D. & Bruenn, J. A. Kinetics of ribosomal pausing during programmed −1 translational frameshifting. Mol. Cell. Biol. 20, 1095–1103 (2000)
Jacks, T., Madhani, H. D., Masiarz, F. R. & Varmus, H. E. Signals for ribosomal frameshifting in the Rous sarcoma virus gag-pol region. Cell 55, 447–458 (1988)
Namy, O., Moran, S. J., Stuart, D. I., Gilbert, R. J. & Brierley, I. A mechanical explanation of RNA pseudoknot function in programmed ribosomal frameshifting. Nature 441, 244–247 (2006)
Weiss, R. B., Dunn, D. M., Shuh, M., Atkins, J. F. & Gesteland, R. F. E. coli ribosomes re-phase on retroviral frameshift signals at rates ranging from 2 to 50 percent. New Biol. 1, 159–169 (1989)
Léger, M., Dulude, D., Steinberg, S. V. & Brakier-Gingras, L. The three transfer RNAs occupying the A, P and E sites on the ribosome are involved in viral programmed −1 ribosomal frameshift. Nucleic Acids Res. 35, 5581–5592 (2007)
Larsen, B., Gesteland, R. F. & Atkins, J. F. Structural probing and mutagenic analysis of the stem-loop required for Escherichia coli dnaX ribosomal frameshifting: programmed efficiency of 50%. J. Mol. Biol. 271, 47–60 (1997)
Larsen, B., Wills, N. M., Gesteland, R. F. & Atkins, J. F. rRNA–mRNA base pairing stimulates a programmed −1 ribosomal frameshift. J. Bacteriol. 176, 6842–6851 (1994)
Chen, J. et al. High-throughput platform for real-time monitoring of biological processes by multicolor single-molecule fluorescence. Proc. Natl Acad. Sci. USA 111, 664–669 (2014)
Kim, H. K. et al. A frameshifting stimulatory stem loop destabilizes the hybrid state and impedes ribosomal translocation. Proc. Natl Acad. Sci. USA 111, 5538–5543 (2014)
Chen, J., Petrov, A., Tsai, A., O’Leary, S. E. & Puglisi, J. D. Coordinated conformational and compositional dynamics drive ribosome translocation. Nature Struct. Mol. Biol. 20, 718–727 (2013)
Chen, J., Tsai, A., Petrov, A. & Puglisi, J. D. Nonfluorescent quenchers to correlate single-molecule conformational and compositional dynamics. J. Am. Chem. Soc. 134, 5734–5737 (2012)
Aitken, C. E. & Puglisi, J. D. Following the intersubunit conformation of the ribosome during translation in real time. Nature Struct. Mol. Biol. 17, 793–800 (2010)
Tsuchihashi, Z. & Brown, P. O. Sequence requirements for efficient translational frameshifting in the Escherichia coli dnaX gene and the role of an unstable interaction between tRNALys and an AAG lysine codon. Genes Dev. 6, 511–519 (1992)
Bertrand, C., Prere, M. F., Gesteland, R. F., Atkins, J. F. & Fayet, O. Influence of the stacking potential of the base 3′ of tandem shift codons on −1 ribosomal frameshifting used for gene expression. RNA 8, 16–28 (2002)
Johansson, M., Zhang, J. & Ehrenberg, M. Genetic code translation displays a linear trade-off between efficiency and accuracy of tRNA selection. Proc. Natl Acad. Sci. USA 109, 131–136 (2012)
Tourigny, D. S., Fernandez, I. S., Kelley, A. C. & Ramakrishnan, V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science 340, 1235490 (2013)
Hughes, D., Atkins, J. F. & Thompson, S. Mutants of elongation factor Tu promote ribosomal frameshifting and nonsense readthrough. EMBO J. 6, 4235–4239 (1987)
Qu, X. et al. The ribosome uses two active mechanisms to unwind messenger RNA during translation. Nature 475, 118–121 (2011)
Liu, C. Y., Qureshi, M. T. & Lee, T. H. Interaction strengths between the ribosome and tRNA at various steps of translocation. Biophys. J. 100, 2201–2208 (2011)
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003)
Qin, P., Yu, D., Zuo, X. & Cornish, P. V. Structured mRNA induces the ribosome into a hyper-rotated state. EMBO Rep. 15, 185–190 (2014)
Jacks, T. et al. Characterization of ribosomal frameshifting in HIV-1 gag-pol expression. Nature 331, 280–283 (1988)
Liao, P. Y., Choi, Y. S., Dinman, J. D. & Lee, K. H. The many paths to frameshifting: kinetic modelling and analysis of the effects of different elongation steps on programmed -1 ribosomal frameshifting. Nucleic Acids Res. 39, 300–312 (2011)
Uemura, S. et al. Real-time tRNA transit on single translating ribosomes at codon resolution. Nature 464, 1012–1017 (2010)
Marshall, R. A., Dorywalska, M. & Puglisi, J. D. Irreversible chemical steps control intersubunit dynamics during translation. Proc. Natl Acad. Sci. USA 105, 15364–15369 (2008)
Dorywalska, M. et al. Site-specific labeling of the ribosome for single-molecule spectroscopy. Nucleic Acids Res. 33, 182–189 (2005)
Blanchard, S. C., Kim, H. D., Gonzalez, R. L., Jr, Puglisi, J. D. & Chu, S. tRNA dynamics on the ribosome during translation. Proc. Natl Acad. Sci. USA 101, 12893–12898 (2004)
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008)
Frank, J. & Agrawal, R. K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000)
Agirrezabala, X. et al. Visualization of the hybrid state of tRNA binding promoted by spontaneous ratcheting of the ribosome. Mol. Cell 32, 190–197 (2008)
Munro, J. B., Altman, R. B., O’Connor, N. & Blanchard, S. C. Identification of two distinct hybrid state intermediates on the ribosome. Mol. Cell 25, 505–517 (2007)
Blanchard, S. C., Gonzalez, R. L., Kim, H. D., Chu, S. & Puglisi, J. D. tRNA selection and kinetic proofreading in translation. Nature Struct. Mol. Biol. 11, 1008–1014 (2004)
Moazed, D. & Noller, H. F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989)
Cornish, P. V., Ermolenko, D. N., Noller, H. F. & Ha, T. Spontaneous intersubunit rotation in single ribosomes. Mol. Cell 30, 578–588 (2008)
Marshall, R. A., Aitken, C. E. & Puglisi, J. D. GTP hydrolysis by IF2 guides progression of the ribosome into elongation. Mol. Cell 35, 37–47 (2009)
Wen, J. D. et al. Following translation by single ribosomes one codon at a time. Nature 452, 598–603 (2008)
Li, G. W., Oh, E. & Weissman, J. S. The anti-Shine–Dalgarno sequence drives translational pausing and codon choice in bacteria. Nature 484, 538–541 (2012)
Acknowledgements
This work was supported by US National Institutes of Health (NIH) grant GM51266 to J.C., A.T. and J.D.P.; by NIH grant GM099687 to A.P., S.E.O’L. and J.D.P.; Wenner-Gren Foundations (Stockholm) to M.J.; and by a Stanford Interdisciplinary Graduate Fellowship to J.C. The authors thank D. Hsu and R. Dalal (Pacific Biosciences Inc.) for their assistance on the ZMW instrumentation; and S. Yan, H. K. Kim and I. Tinoco Jr (University of California at Berkeley) for helpful discussions. J.C. would like to thank I Lin for support.
Author information
Authors and Affiliations
Contributions
J.C. performed all the experiments and the data analysis. J.C., A.P., A.T. and J.D.P. designed the project and wrote the manuscript. M.J. and S.E.O’L. assisted with reagent preparation. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Explanation and schematics of the experimental signals.
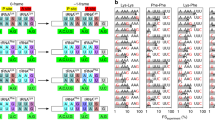
a, Different stages of the elongation cycle during translation. Frameshifting has been proposed to occur at each of the steps: (1) during accommodation of the A-site tRNA4, (2) subsequent to accommodation, but before peptide bond formation8, (3) during tRNA hybrid-state intermediates10, (4) during translocation9, and (5) at the start of the next found of elongation11. b, The ribosome starts each round of elongation in the non-rotated state. In this ‘locked’ state, the P-site tRNA is stably bound in the classical state, preserving the reading frame of the mRNA18. Upon A-site tRNA selection and peptide bond formation, the 30S subunit rotates 3–10° anti-clockwise with respect to the 50S subunit to the rotated state (pre-translocation)26,35,36. This ‘unlocked’ state permits tRNA motions and the tRNA can fluctuate freely between the classical state and hybrid state37, thus facilitating translocation of tRNA and movement of ribosome by one codon over mRNA38,39. Peptide-bond formation also triggers spontaneous fluctuations of the L1 stalk between open and closed conformations as well as spontaneous rotations in ribosome conformations36,40. EF-G then catalyses translocation, and the ribosome returns to the non-rotated state (post-translocation). To monitor the rotational state of the ribosome in real time, we employed FRET between the small (30S) and large (50S) subunits. The 30S subunit was site-specifically labelled with Cy3B on helix 44, and a non-fluorescent quencher, BHQ-2, was placed on helix 101 of 50S subunit18,31,32. Reagent delivery of BHQ-50S, tRNA ternary complex, and EF-G to surface immobilized Cy3B-30S pre-initiation complexes in ZMWs results in IF2-guided 70S assembly during initiation and establishment of FRET between the two ribosomal subunits: upon subunit joining41, the green (Cy3B) intensity drops, which is followed by alternating low-high-low intensities. Each alternating cycle corresponds to the ribosome translating a single codon, with the two intensity states consistent with the two rotational states of the ribosome: the low intensity state (high FRET) defining the non-rotated (locked) ribosome conformation and the high intensity state (low FRET) the rotated (unlocked) conformation18. The rotated- and non-rotated-state lifetimes at each codon can be statistically analysed. c, During each cycle of elongation, the ribosome selects the aminoacyl-tRNA in a ternary complex with EF-Tu–GTP, and positions the tRNA in the A site. Upon A-site tRNA accommodation, the ribosome rapidly catalyses peptide bond formation with the P-site tRNA33,38. Translocation moves A- and P-site tRNA–mRNA complexes to the E and P site respectively, catalysed by the EF-G. The compositional dynamics of tRNA and EF-G on the ribosome, here defined as the relative timing of their arrival and departure during elongation, can be observed by labelling the tRNA or EF-G with Cy3 or Cy5. Cy5/Cy3–tRNA arrival to the surface immobilized ribosomes is marked by a red/green fluorescent pulse. Translation can be monitored by the arrival and departure of dye-labelled tRNA. Each productive tRNA binding event results in a fluorescence pulse that lasts as a ribosome translates 2 codons—beginning with arrival of tRNA in the A site, continuing through tRNA translocation to the P site, arrival of A-site tRNA to the next codon, a second round of translocation, and ending with spontaneous dissociation of tRNA from the E site30. To track tRNA and EF-G dynamics on translating ribosomes at near-physiological concentrations of fluorescent factors (0.1–1 µM), we used ZMWs to detect hundreds of individual ribosomes30. d, Substituting the traditional FRET acceptor, Cy5, with BHQ-2 allowed the use of Cy5 to label other translation components for correlation studies. The Cy3B intensity reports on the conformational state of the ribosome, whereas Cy5 pulses indicate arrival, occupancy and departure of ribosomal ligands.
Extended Data Figure 2 Phe codon in the −1 frame confirms the characteristic long pause during frameshifting.
a, Histogram of the fraction of ribosomes translating to a particular codon for the dnaX wild-type mRNA, with a schematic. Many of the ribosomes translate up to 12 codons to a 0 frame stop codon, though a large percentage of ribosomes translate up to 9 codons to a −1 frame stop codon. There are also ribosomes that stall at codon 7, limited by Cy3B photobleaching or end of movie (8 min) from the long rotated state pause. Interestingly, there is also a number of ribosomes that stall at codon 8 (see Extended Data Fig. 10 for discussion). By parsing the number of ribosomes that translate beyond codon 9 and up to codon 9, the frameshifting percentage can be calculated (75%). However, as non-frameshifted ribosomes may terminate early, this would lead to a slight over-estimate of the frameshifting percentage (∼3–10%). The frameshifting efficiency has been independently confirmed using a Cy5–tRNAPhe score, as described below. Number of molecules analysed, n = 256. b, A UUC(Phe) codon is introduced in the −1 frame downstream of the slippery site. Frameshifting can be scored by an appearance of a Cy5 (red) pulse with Cy5–tRNAPhe in addition to the Cy3B/BHQ ribosome FRET signal. This allows us to independently score for frameshifting. c, Using the Cy5–tRNAPhe as a score to confirm frameshifting, we get the same dynamics and lifetimes: the non-rotated state lifetimes remain constant at each codon, and the rotated state lifetime increases tenfold at the seventh FRET cycle (codon Lys7 and Lys8 at the slippery sequence due to uncoupled translocation). This confirms and justifies our results in Fig. 1. Number of molecules analysed, n = 474. Error bars, s.e. d, By using the Cy5–tRNAPhe as a score, we can parse the rotated state lifetimes into ribosomes that frameshifted and ribosomes that did not frameshift. We also get the same results as Fig. 1: non-frameshifted ribosomes translate through the frameshift sequence seemingly unaffected; frameshifted ribosomes exhibit the characteristic long-rotated state pause at the seventh FRET cycle (codon Lys7 and Lys8 due to uncoupled translocation). Number of molecules analysed, n = 474. Error bars, s.e.
Extended Data Figure 3 Hairpin and the internal Shine–Dalgarno sequence are important for frameshifting.
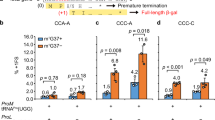
a, mRNA sequence of the no hairpin (no HP) mutant. The mRNA consists of the same sequence as the wild-type dnaX frameshift sequence, but with the sequence after the UGA stop codon in the 0 frame in the hairpin deleted. b, Non-rotated and rotated state lifetimes in the presence of 80 nM EF-G and 1 µM tRNAtot. The non-rotated state lifetimes are constant at each codon. There is an increase in rotated state lifetime at codon Lys7. Number of molecules analysed, n = 124. Error bars, s.e. c, mRNA sequence of the no Shine–Dalgarno (no SD) mutant. The mRNA consists of the same sequence as the wild-type dnaX frameshift sequence, but with the original internal Shine–Dalgarno sequence GGGAGC mutated to AGGCGC, decreasing the rRNA–mRNA interaction energy from −4.70 kcal mol−1 to −0.00 kcal mol−1. d, Non-rotated and rotated state lifetimes in the presence of 80 nM EF-G and 1 µM tRNAtot. The non-rotated state lifetime is constant. There is an increase in rotated state lifetime at codon Lys7. Number of molecules analysed, n = 225. Error bars, s.e. e, Frameshifting percentages of the no Shine–Dalgano and no hairpin mutant. Without the Shine–Dalgarno sequence, frameshifting percentage drops by half. Without the hairpin, frameshifting percentage drops to a quarter of the wild-type sequence. This indicates that both the internal Shine–Dalgarno sequence and the hairpin are required for efficient frameshifting, confirming that the hairpin and internal Shine–Dalgarno are stimulatory elements for frameshifting. These stimulatory elements may present a barrier and tension to translocation that is a prerequisite for efficient frameshifting.
Extended Data Figure 4 Hairpin and internal Shine–Dalgarno sequences increases the energy barrier to translocation.
a, Translation of a short linear mRNA, 6(FK), in the presence of 80 nM EF-G and 1 µM tRNAtot, with a sample trace. b, Histogram of fraction of ribosomes translating to a particular codon. Most of the ribosomes translate up to 12 codons. Ribosomes translate <12 codons are due to photobleaching of the Cy3B dye, or non-processive ribosomes. This gives us a background level of ∼3–10% for our frameshifting efficiency analysis. The small number of ribosomes that translate beyond codon 12 are probably errors in our statistical analysis or read-through of the stop codon. Number of molecules analysed, n = 462. c, Rotated and non-rotated state lifetimes are fairly constant. Number of molecules analysed, n = 462. Error bars, s.e. d, Translation of a Phe-Lys sequence preceded by an internal Shine–Dalgarno sequence (same Shine–Dalgarno sequence used in the dnaX frameshift mRNA of this study) in the presence of 80 nM EF-G and 1 µM tRNAtot, with an example trace. e, Histogram of fraction of ribosomes translating to a particular codon. Number of molecules analysed, n = 60. f, There is an increase in rotated state lifetime at codon 5–7. There is an increase in the rotated state lifetimes ∼3–4-fold over 3–5 codons downstream of the Shine–Delgano-like sequence, whereas the non-rotated state lifetime remains unaffected. The internal Shine–Delgano-like sequences may base pair with the 3′ end of the 16S rRNA and slow down ribosomes in the pre-translocation state, echoing several work done previously by tracking ribosome movement and ribosome profiling42,43. Number of molecules analysed, n = 60. Error bars, s.e. g, Translation of a Phe-Lys sequence followed by a hairpin (same hairpin used in the dnaX frameshift mRNA of this study) in the presence of 80 nM EF-G and 1 µM tRNAtot, with an example trace. h, Histogram of fraction of ribosomes translating to a particular codon. Number of molecules analysed, n = 332. i, Non-rotated state lifetimes are fairly constant. There is an increase in rotated state lifetime at codon 5, exactly 3 codons before the start of the hairpin, placing the ribosome directly at the first encounter of the hairpin. The relative position also matches where we see the long-rotated state pause during frameshift. This echoes the work which showed the ribosome is capable of translating through the secondary structure through two mechanisms: ribosome translocating when encountering an open-state junction, occurring naturally or induced by the ribosome, or mechanically unwinding by the ribosome when encountering a closed-state junction24. When the ribosome encounters an open-state junction, translation proceeds at a constant rate; however, when a closed-state junction is encountered, the ribosome actively unwinds the secondary structure, resulting in a slight waiting time for translocation, after which the hairpin is biased by the ribosome into an open-state and translation occurs normally. The shunt to either pausing in the rotated state (which leads to uncoupled translocation) or normal translation during frameshifting is probably due to this mechanism. Number of molecules analysed, n = 332. Error bars, s.e.
Extended Data Figure 5 Dynamics of frameshifting at different factor concentrations.
a, Example trace and schematic of a ribosome translating the dnaX frameshift mRNA at much higher factor concentrations (6 µM tRNAtot and 480 nM EF-G). b, Frameshifting efficiency does not depend on EF-G and tRNAtot concentrations. c, Increasing the tRNAtot and EF-G concentrations twofold (from 1 µM tRNAtot and 80 nM EF-G to 2 µM tRNAtot and 160 nM EF-G) decreases both the rotated state lifetime and non-rotated state lifetimes. This confirms that our ribosome FRET signal depends correctly on factor concentration. Number of molecules analysed, n = 256 (1 µM tRNAtot and 80 nM EF-G), n = 234 (2 µM tRNAtot and 160 nM EF-G). Error bars, s.e. d, Increasing the tRNAtot concentration (from 1 µM to 3 µM) while keeping EF-G concentrations constant decrease the non-rotated state lifetimes threefold as expected. The rotated state lifetimes remain the same except for codon Lys7; this is expected because the rotated state lifetime depends only on concentration of EF-G. Unexpectedly, the rotated state lifetime at codon Lys7 (Lys8 after uncoupled translocation) is also slightly decreased, suggesting a linkage between tRNA and EF-G dynamics at that long rotated-state stall. This echoes our results in Fig. 2 that tRNA (tRNALys in this case) samples the uncoupled rotated-state after translocation, and the tRNA sampling and accommodation and EF-G sampling may help to re-establish the ribosome’s reading frame and reverse-rotate subsequently. Thus, increasing tRNA concentrations (especially tRNALys in this case) will decrease the long rotated-state lifetime. Number of molecules analysed, n = 526. Error bars, s.e. e, Increasing the EF-G concentration (from 80 nM to 240 nM) while keeping tRNAtot concentration constant decreases the rotated state lifetime threefold. The non-rotated state lifetime, which depends on the tRNA concentration, remains the same. However, the decrease in the rotated lifetime at codon Lys7 is only around twofold, rather than threefold as expected. This echoes our result in Fig. 2 as well as in b above, suggesting that tRNA sampling also plays a role at this codon. Number of molecules analysed, n = 314. Error bars, s.e. f, Increasing the EF-G and tRNAtot concentrations further to 6 µM tRNAtot and 480 nM EF-G further decreases the rotated and non-rotated state lifetimes. However, the rotated state lifetime at codon Lys7 remains the same when compared with 2 µM tRNAtot and 160 nM EF-G. The long tRNALys sampling events observed in Fig. 2 may be contributing to the long-rotated state lifetime at codon Lys7.
Extended Data Figure 6 Slippery sequence mutation (A21G–A24G) decreases frameshifting percentage.
a, Sample trace of a ribosome translating the dnaX A21G–A24G mutant mRNA in the presence of 80 nM EF-G and 1 µM tRNAtot. There seems to be a slightly longer pause at codon Lys7. b, Histogram of the fraction of ribosomes translating to a particular codon for the dnaX A21G–A24G mutant mRNA. Most of the ribosomes translate up to 12 codons to a 0 frame stop codon. The buildup of ribosomes stalled at codon 9 present during frameshifting disappears. By parsing the number of ribosomes that translate beyond codon 9 and up to codon 9, the frameshifting percentage can be calculated (12%). c, The rotated-state lifetime. The long stall at Lys7 is decreased with the slippery site mutant, suggesting that the extra-long pause is indeed a result of frameshifting. The slight increase in lifetime at Lys7 is due to the effects of the hairpin and internal Shine–Dalgarno sequence. Number of molecules analysed, n = 230. Error bars, s.e. d, A UUC(Phe) is introduced in the −1 frame downstream of the slippery site of the A21G–A24G mutant, similar to above. The A21G–A24G mutation is known to decrease frameshifting efficiency down to background levels19. e, The non-rotated state lifetime and rotated-state lifetime match with our results using codon counting (see above). In the absence of frameshifting, there is still an increase in rotated state lifetime at codon Lys7, due to the increased energy barrier to translocation by the hairpin and internal Shine–Dalgarno sequence, though this increased lifetime is still much less than the Lys7 rotated state lifetime during frameshifting. Number of molecules analysed, n = 538. Error bars, s.e. f, Using Cy5–tRNAPhe as a score for frameshifting, frameshifting percentage matches with our previous results. The slippery sequence A21G–A24G mutant decreases frameshifting percentage down to background levels. Number of molecules analysed, from left to right, n = 474, n = 538.
Extended Data Figure 7 tRNA dynamics during frameshifting with the dnaX GCA(Ala) to GUA(Val) mutant mRNA.
a, The 3 nucleotides upstream of the slippery sequence (GCA(Ala)) are mutated to GUA(Val) (named the C20U mutant) so that E-site tRNA dynamics can be observed during frameshifting since tRNAVal can be labelled with Cy3-maleimide (see Fig. 2). This allows us to estimate the time to translocation during the long rotated-state pause at codon Lys7, as translocation of the Cy3–tRNAVal from the P-site to the E-site leads to rapid departure of the tRNAVal and disappearance of the Cy3 signal. We want to make sure that the C20U mutation does not affect frameshifting dynamics. The non-rotated state and rotated state lifetimes, as well as frameshifting percentages, are consistent with what we have observed before for the wild-type sequence. Number of molecules analysed, n = 266. Error bars, s.e. b, Post-synchronization density plot of tRNA–tRNA FRET between the Cy3–tRNAVal in the P site and the incoming Cy5–tRNALys at Lys7 in the A site at the slippery sequence. The tRNAs are in a hybrid state upon encountering of hairpin and engagement with the internal Shine–Dalgarno sequence. Thus, uncoupled translocation occurs with normal tRNA hybrid state formation. After translocation, the Cy3–tRNAVal departs from the ribosome, resulting in a disappearance of FRET. After translocation and uncoupling with ribosome reverse-rotation, the now P-site tRNALys is probably in a ‘distorted’ conformation, according to the structure by Namy et al.9. Number of molecules analysed, left n = 227, right n = 337.
Extended Data Figure 8 tRNALys transit and sampling dynamics.
a, Example trace of Cy5–tRNALys transit during translation of the dnaX wild-type mRNA, indicating the definition of pulse lifetime and time between pulse. b, The time between tRNALys pulse and lifetime of each pulse for the first three Lys codons are consistent with what is expected, and decrease expectedly with the increase of EF-G concentration and tRNAtotal(ΔLys) concentration ([Cy5–tRNALys] = 200 nM). The time between pulse for the first Lys pulse corresponds to the decoding of the first codon Lys1 of the mRNA after 50S subunit joining to the immobilized 30S, which is short as expected and does not depend on factor concentration. The second Lys pulse has a longer time between pulse, corresponding to the ribosome translating four codons from Lys1 to Lys5. The third pulse has a slightly shorter time between pulse, corresponding to the ribosome translating from codon Lys5 to Lys7. The lifetimes for the first two pulses are short, as the ribosomes decode and translocate the corresponding codons normally. The lifetime for the third pulse at codon Lys7 is long, corresponding to the ribosome at the long rotated-state pause during frameshifting. Number of molecules analysed, n = 179 (1 µM tRNAtot and 80 nM EF-G) and n = 212 (2 µM tRNAtot and 160 nM EF-G). Error bars, s.e. c, Mean number of additional tRNALys sampling pulses to the long rotated-state pause at codon Lys7, sampling lifetimes, and sampling arrival times, at various concentrations of EF-G and tRNAtot ([Cy5–tRNALys] = 200 nM) for ribosomes translating the dnaX wild-type frameshifting sequence. There is a mean number of 2.3 sampling tRNALys pulses, which remains constant at the various factor concentrations. There is not a clear dependency of the sampling arrival times and lifetimes on concentration of EF-G and the other tRNAs, probably because the factors are all competing for the ribosomal A site. Number of molecules analysed, from left to right, n = 179, n = 212, n = 180 and n = 162. Error bars, s.e. d, The time between tRNALys pulse and lifetime of each pulse for the first three Lys codons are the same for the dnaX wild-type AAG sequence and the AAG(AAA) mutant sequence. Number of molecules analysed, n = 212 (AAG) and n = 454 (AAG(AAA)). Error bars, s.e. e, By translating the AAG(AAA) mutant in the presence of Cy5–tRNALys, we see similar dynamics as the dnaX wild-type sequence. Cy5–tRNALys samples the A site at codon Lys8 after uncoupled translocation at the frameshift site. The fraction of ribosomes exhibiting >4 tRNALys pulses are the same for the wild-type sequence and the AAG(AAA) mutant. The mean number of tRNALys sampling pulses, the mean arrival time, and the mean lifetimes of the sampling pulses to the long rotated-state stall are the same for the AAG(AAA) mutant and the dnaX wild-type sequence. Number of molecules analysed, n = 212 (left, wild-type AAG), n = 454 (right, AAG(AAA)). Error bars, s.e.
Extended Data Figure 9 tRNA sampling dynamics and slippage during frameshifting.
a, Sample traces of Cy5–tRNAPhe (red) sampling to the A-site Phe8 codon during the long rotated-state pause correlated with Cy3B/BHQ ribosome FRET signal (green) for the AAG(UUU) mutant. b, By translating the AAG(UUU) mutant in the presence of Cy5–tRNAPhe (red) and correlating with the Cy3B/BHQ ribosome FRET signal (green), we can observe the fraction of ribosomes exhibiting only 1 Cy5–tRNAPhe pulse or > 1 Cy5–tRNAPhe pulse sampling to the long rotated state pause after codon Lys7. There is a significant number of ribosomes exhibiting > 1 Cy5–tRNAPhe pulse even when there is only one Phe codon, suggesting that even without frameshifting, many of the ribosomes still pause in an uncoupled rotated state after Lys7, where tRNAPhe samples the exposed UUU codon in the A site. Number of molecules analysed, n = 106. c, The arrival time of the first tRNA sampling to the long stalled codon for wild-type mRNA (with Cy5–tRNALys) and AAG(UUU) with Cy5–tRNAPhe are the same. Although frameshifting in principle could occur through an incomplete +2 translocation9,10 with weakened codon–anticodon–ribosome interactions and the final reading-frame determined through the Lys8 tRNALys sampling in the −1 frame, our data support +3 translocation that weakens codon–anticodon–ribosome interactions followed by tRNA sampling and accommodation that defines the reading frame and shifts −1 through effects of the hairpin and internal-Shine–Dalgarno interaction. For +2 translocation, we would expect to see tRNA sampling to both −1 frame and the 0 frame of A-site codon. For the AAG(UUU) mutant, tRNAPhe will sample the 0 frame U25U26U27, whereas tRNAIle will sample the −1 frame A24U25U26. As Cy5–tRNAPhe arrival times and lifetimes for sampling to the AAG(UUU) mutant match Cy5–tRNALys arrival times and lifetimes for the wild-type sequence, there is probably no competition between tRNAPhe and tRNAIle, suggesting that the AUU codon is not initially exposed for tRNAIle sampling. Furthermore, for the +2 model, we would not expect a AAG(UUU) mutation to lead to a decrease in frameshifting efficiency. Thus, our results favour a +3 translocation followed by a −1 shift driven by sampling, accommodation and base-pairing stability. Unfortunately, our single-molecule assay is blind to the actual movement of the ribosome on the mRNA, so the details of this mechanism will require further exploration. See Extended Data Fig. 10 for possible implications for heterogeneous frameshift products observed previously28. Error bars, s.e. Number of molecules analysed, from left to right, n = 212, n = 106. d, Mean sampling lifetime and mean sampling arrival time for Cy5–tRNAPhe to the Phe8 codon for the AAG(UUU) mutant. The arrival time and lifetime are the same as Cy5–tRNALys sampling to the Lys8 codon for the dnaX wild-type sequence. Number of molecules analysed n = 106. e, Example trace for Cy5–tRNALys transit through the dnaX AAG(UUU) mutant mRNA. For the AAG(UUU) mutant, we see only three Lys pulses, as expected, as the fourth Lys codon (Lys8) is mutated to a Phe codon. Most of the ribosomes (∼80%) exhibit only three Cy5–tRNALys pulses, indicating that the additional Cy5–tRNALys sampling pulses we saw characteristic of frameshifting are indeed sampling to the Lys8 codon. Sampling now is by tRNAPhe, which are dark and invisible to our observations. The time between pulses are consistent with both the wild-type sequence and AAG(AAA) mutant. The lifetime of the third pulse (at Lys7) is long, consistent with the long pause at that codon. Number of molecules analysed, n = 318. Error bars, s.e. f, Sample trace for Cy5–tRNALys transit through the dnaX AAG(AAC) mutant mRNA. For the AAG(AAC) mutant, we see mostly only three Lys pulses (∼75%) since the fourth Lys codon (Lys8) is mutated to a Asn codon. Number of molecules analysed n = 406. This further argues against the +2 translocation model (see c). For +2 translocation, we would expect to see long tRNALys sampling to the −1 frame AAA codon, which is not observed significantly. The additional Cy5–tRNALys pulses have a shorter lifetime and longer arrival time when compared with the translation of the wild-type mRNA, suggesting that these pulses are non-cognate sampling to the AAC codon in the 0 frame or sampling unstably to the AAA codon in the −1 frame. Even though our data support a +3 translocation followed by a −1 slippage, multiple frameshifting pathways probably occur. The details of this mechanism will require further exploration.
Extended Data Figure 10 Hetereogeneous frameshift products.
a, Two different protein products are possible after −1 frameshifting, dependent on whether peptide bond formation occurs during sampling in the −1 or 0 frame. For the first scenario, tRNA sampling to the last three nucleotides of the slippery sequence (YYZ) redefines the ribosome in the −1 frame (YYY), after which the tRNA dissociates to leave an empty A-site codon. After the long rotated state is reverse-rotated by EF-G, tRNAYYY decodes that codon normally, creating a frameshift product denoted by XXY-YYY. For the second scenario, peptide bond formation occurs after slippage of tRNAYYZ into the −1 frame; peptide bond formation occurs slowly, as the P- and A-site tRNAs would probably not be positioned correctly in the rotated ribosomal conformation. EF-G would then normally and rapidly resolve the newly-created A/P hybrid state and the ribosome reverse-rotates. In this case, the frameshift product will be denoted by XXY-YYZ. b, Histogram of the fraction of ribosomes translating to a particular codon for the dnaX AAG(UUU) mRNA, with a schematic. As the frameshifting percentage for the AAG(UUU) sequence is low, we see that most of the ribosomes translate up to 12 codons to a 0 frame stop codon. However, there is a significant number of ribosomes that translate to 11 codons (∼25%), compared to ∼5% for our previous experiments. There are two possible scenarios for tRNAPhe sampling to the long rotated-state pause to codon Phe8. In the first case, the tRNAPhe defines the reading frame and falls off, after which the ribosome resolves itself through the action of EF-G, followed by the normal decoding of Phe codon at codon 8. In this case, we get 12 cycles of low-high-low FRET intensity, and hence 12 codons translated by our signal. In the second case, the tRNAPhe defines the reading frame, followed by slow peptide bond formation. After peptide bond formation, the ribosome returns to the canonical hybrid and rotated state, for which EF-G then catalyses reverse rotation. In this case, one low-high-low FRET cycle is missed, so we count 11 codons translated by our signal. This explains the heterogeneity in frameshift products observed in many frameshift systems28. Number of molecules analysed, n = 353. c, Sample traces of Cy5–tRNAPhe (red) sampling to the long rotated-state pause at codon Lys7 correlated with Cy3B/BHQ ribosome FRET signal (green), showing the two possible scenarios for tRNA sampling. Case 1 (as described in a) leads to correlation of tRNA arrival and ribosome rotation after the long rotated state pause whereas case 2 leads to overlap of a tRNAPhe pulse with the reverse-rotation of the long pause. Both scenarios occur when translating the AAG(UUU) mutant, with ∼58% of ribosomes for case 1 and 42% for case 2. For the second case, the time between the last Cy5–tRNAPhe arrival and the ribosome reverse-rotation is 27.2 s, much longer than the 7.7 s during normal decoding and translocation, suggesting a slow peptidyltransfer reaction. Our results provide a possible explanation for why heterogeneous frameshifting products are observed in many frameshifting systems28. However, the details of this mechanism will require further exploration. Number of molecules analysed, n = 55.
Rights and permissions
About this article
Cite this article
Chen, J., Petrov, A., Johansson, M. et al. Dynamic pathways of −1 translational frameshifting. Nature 512, 328–332 (2014). https://doi.org/10.1038/nature13428
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13428
This article is cited by
-
Frameshift and wild-type proteins are often highly similar because the genetic code and genomes were optimized for frameshift tolerance
BMC Genomics (2022)
-
Specific length and structure rather than high thermodynamic stability enable regulatory mRNA stem-loops to pause translation
Nature Communications (2022)
-
Oligodendrocyte differentiation alters tRNA modifications and codon optimality-mediated mRNA decay
Nature Communications (2022)
-
Length-dependent motions of SARS-CoV-2 frameshifting RNA pseudoknot and alternative conformations suggest avenues for frameshifting suppression
Nature Communications (2022)
-
Altered tRNA dynamics during translocation on slippery mRNA as determinant of spontaneous ribosome frameshifting
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.