Abstract
Ancient and diverse antibiotic resistance genes (ARGs) have previously been identified from soil1,2,3, including genes identical to those in human pathogens4. Despite the apparent overlap between soil and clinical resistomes4,5,6, factors influencing ARG composition in soil and their movement between genomes and habitats remain largely unknown3. General metagenome functions often correlate with the underlying structure of bacterial communities7,8,9,10,11,12. However, ARGs are proposed to be highly mobile4,5,13, prompting speculation that resistomes may not correlate with phylogenetic signatures or ecological divisions13,14. To investigate these relationships, we performed functional metagenomic selections for resistance to 18 antibiotics from 18 agricultural and grassland soils. The 2,895 ARGs we discovered were mostly new, and represent all major resistance mechanisms15. We demonstrate that distinct soil types harbour distinct resistomes, and that the addition of nitrogen fertilizer strongly influenced soil ARG content. Resistome composition also correlated with microbial phylogenetic and taxonomic structure, both across and within soil types. Consistent with this strong correlation, mobility elements (genes responsible for horizontal gene transfer between bacteria such as transposases and integrases) syntenic with ARGs were rare in soil by comparison with sequenced pathogens, suggesting that ARGs may not transfer between soil bacteria as readily as is observed between human pathogens. Together, our results indicate that bacterial community composition is the primary determinant of soil ARG content, challenging previous hypotheses that horizontal gene transfer effectively decouples resistomes from phylogeny13,14.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
D'Costa, V. M. et al. Antibiotic resistance is ancient. Nature 477, 457–461 (2011)
Allen, H. K., Moe, L. A., Rodbumrer, J., Gaarder, A. & Handelsman, J. Functional metagenomics reveals diverse beta-lactamases in a remote Alaskan soil. ISME J. 3, 243–251 (2009)
Allen, H. K. et al. Call of the wild: antibiotic resistance genes in natural environments. Nature Rev. Microbiol. 8, 251–259 (2010)
Forsberg, K. J. et al. The shared antibiotic resistome of soil bacteria and human pathogens. Science 337, 1107–1111 (2012)
Wright, G. D. Antibiotic resistance in the environment: a link to the clinic? Curr. Opin. Microbiol. 13, 589–594 (2010)
Benveniste, R. & Davies, J. Aminoglycoside antibiotic-inactivating enzymes in actinomycetes similar to those present in clinical isolates of antibiotic-resistant bacteria. Proc. Natl Acad. Sci. USA 70, 2276–2280 (1973)
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nature Biotechnol. 31, 814–821 (2013)
Fierer, N. et al. Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J. 6, 1007–1017 (2012)
Fierer, N. et al. Cross-biome metagenomic analyses of soil microbial communities and their functional attributes. Proc. Natl Acad. Sci. USA 109, 21390–21395 (2012)
Fierer, N. et al. Reconstructing the microbial diversity and function of pre-agricultural tallgrass prairie soils in the United States. Science 342, 621–624 (2013)
Muegge, B. D. et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332, 970–974 (2011)
Zaneveld, J. R., Lozupone, C., Gordon, J. I. & Knight, R. Ribosomal RNA diversity predicts genome diversity in gut bacteria and their relatives. Nucleic Acids Res. 38, 3869–3879 (2010)
Smillie, C. S. et al. Ecology drives a global network of gene exchange connecting the human microbiome. Nature 480, 241–244 (2011)
Stokes, H. W. & Gillings, M. R. Gene flow, mobile genetic elements and the recruitment of antibiotic resistance genes into Gram-negative pathogens. FEMS Microbiol. Rev. 35, 790–819 (2011)
Walsh, C. Molecular mechanisms that confer antibacterial drug resistance. Nature 406, 775–781 (2000)
Pehrsson, E. C., Forsberg, K. J., Gibson, M. K., Ahmadi, S. & Dantas, G. Novel resistance functions uncovered using functional metagenomic investigations of resistance reservoirs. Front. Microbiol. 4, 145 (2013)
Delmont, T. O. et al. Structure, fluctuation and magnitude of a natural grassland soil metagenome. ISME J. 6, 1677–1687 (2012)
Jacoby, G. A. & Munoz-Price, L. S. The new beta-lactamases. N. Engl. J. Med. 352, 380–391 (2005)
Nalbantoglu, O. U., Way, S. F., Hinrichs, S. H. & Sayood, K. RAIphy: phylogenetic classification of metagenomics samples using iterative refinement of relative abundance index profiles. BMC Bioinform. 12, 41 (2011)
Ramirez, K. S., Lauber, C. L., Knight, R., Bradford, M. A. & Fierer, N. Consistent effects of nitrogen fertilization on soil bacterial communities in contrasting systems. Ecology 91, 3463–3470; discussion 3503–3414. (2010)
Ventura, M. et al. Genomics of Actinobacteria: tracing the evolutionary history of an ancient phylum. Microbiol. Mol. Biol. Rev. 71, 495–548 (2007)
Aminov, R. I. & Mackie, R. I. Evolution and ecology of antibiotic resistance genes. FEMS Microbiol. Lett. 271, 147–161 (2007)
Davies, J. & Davies, D. Origins and evolution of antibiotic resistance. Microbiol. Mol. Biol. Rev. 74, 417–433 (2010)
Medeiros, A. A. Evolution and dissemination of beta-lactamases accelerated by generations of beta-lactam antibiotics. Clin. Inf. Diseases 24, (Suppl. 1)19–45 (1997)
Dantas, G., Sommer, M. O., Oluwasegun, R. D. & Church, G. M. Bacteria subsisting on antibiotics. Science 320, 100–103 (2008)
Boucher, H. W. et al. Bad bugs, no drugs: no ESKAPE!. Clin. Inf. Diseases 48, 1–12 (2009)
Knapp, C. W., Dolfing, J., Ehlert, P. A. & Graham, D. W. Evidence of increasing antibiotic resistance gene abundances in archived soils since 1940. Environ. Sci. Technol. 44, 580–587 (2010)
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1–I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997)
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008)
de la Bastide, M. & McCombie, W. R. Assembling Genomic DNA sequences with PHRAP 2008/04/23 edn, Vol. 11 (John Wiley, 2007)
Moore, A. M. et al. Pediatric fecal microbiota harbor diverse and novel antibiotic resistance genes. PLoS ONE 8, e78822 (2013)
Tatusov, R. L., Galperin, M. Y., Natale, D. A. & Koonin, E. V. The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 28, 33–36 (2000)
Zhu, W., Lomsadze, A. & Borodovsky, M. Ab initio gene identification in metagenomic sequences. Nucleic Acids Res. 38, e132 (2010)
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–37 (2011)
Haft, D. H. et al. TIGRFAMs: a protein family resource for the functional identification of proteins. Nucleic Acids Res. 29, 41–43 (2001)
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Res. 28, 263–266 (2000)
McArthur, A. G. et al. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 57, 3348–3357 (2013)
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006)
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature Methods 7, 335–336 (2010)
Lozupone, C., Hamady, M. & Knight, R. UniFrac–an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinform. 7, 371 (2006)
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. ISME J. 5, 169–172 (2011)
Clarke, K. R. & Gorley, R. N. PRIMER v6: User Manual/Tutorial 6th edn, Ch. 13 (PRIMER-E, 2006)
Acknowledgements
We thank M. Pesesky for access to and assistance with the data set used to benchmark RAIphy’s performance, B. Wang for suggested improvements to Illumina library preparation, the Genome Technology Access Center at Washington University in St Louis for generating Illumina sequence data, M. Sherman for discussions on modelling pathogen HGT potential, and members of the Dantas laboratory for discussions on the results and analyses presented here. This work was supported by awards to G.D. through the Children’s Discovery Institute (MD-II-2011-117), the International Center for Advanced Renewable Energy and Sustainability at Washington University, the National Academies Keck Futures Initiatives (Synthetic Biology, SB2), and the NIH Director’s New Innovator Award (DP2-DK-098089). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. M.K.G. is supported by a Mr and Mrs Spencer T. Olin Fellowship for Women in Graduate Study at Washington University. K.J.F. received support from the NIGMS Cell and Molecular Biology Training Grant (GM 007067) and from the NHGRI Genome Analysis Training Program (T32 HG000045). K.J.F. and M.K.G. are NSF graduate research fellows (award number DGE-1143954).
Author information
Authors and Affiliations
Contributions
N.F., C.L.L. and R.K. provided soils and 16S rRNA gene sequencing data; G.D. conceived the functional selections; S.P. created metagenomic libraries, performed functional selections, and prepared sequencing libraries; K.J.F. assembled sequence data from functional selections and annotated ARGs with assistance from M.K.G.; M.K.G. built the custom ARG profile HMM database; K.J.F. performed genomic and ecological analyses and wrote the manuscript with contributions from M.K.G., R.K., N.F. and G.D.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Functional selections of 18 soil metagenomes for resistance against 18 antibiotics.
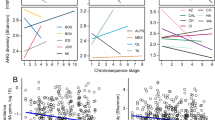
a, Phenotypic results of selections. A dark grey cell means that a resistance phenotype was observed whereas white cells indicate the absence of any drug-tolerant transformants. Grassland soils from CC are labelled in red and agricultural soils from KBS are labelled in blue. b, c, Alpha diversity representations. On the left is depicted the number of distinct ARG annotations observed as increasing numbers of ARGs are sampled from each soil. On the right, Shannon diversity scores (an ecological metric that quantifies within-sample diversity) are shown at each rarefaction step.
Extended Data Figure 2 Three prominent ARG classes are present in nearly all bacterial genomes and can provide antibiotic resistance when overexpressed.
a, Generalized as red circles are dihydrofolate reductases, D-alanine—D-alanine (D-ala D-ala) ligases, which are the molecular targets of the drugs trimethoprim (TR) and D-cycloserine (CY) respectively (black stars), and thymidylate synthases, which can provide trimethoprim resistance by circumventing the need for an active dihydrofolate reductase. When overexpressed in functional selections, these genes can provide antibiotic resistance. We found substantial diversity in these genes (average pairwise amino acid identity 39.3 ± 12.2%), suggesting that variants were captured from many bacterial lineages. b, Relative to other ARG mechanisms, large numbers of dihydrofolate reductases, thymidylate synthases, and D-ala D-ala ligases were found in all soils, with these ARGs representing 92.5% of resistance genes identified from selections containing trimethoprim or D-cycloserine antibiotics. Therefore, these selections encompass large genetic diversity, but constrained functional diversity, with a broad range of genes encoding limited functional traits. c, When considered in isolation, these functions were not different between the KBS and CC soils (P > 0.05, ANOSIM), indicating that trimethoprim and D-cycloserine resistance function is similarly distributed across the surveyed soil types.
Extended Data Figure 3 Total counts of β-lactamases recovered from antibiotic selections.
All soils (black), CC soils (red), and KBS soils (blue).
Extended Data Figure 4 Total counts of ARGs categorized by their predicted phylogenetic origin.
The number of ARGs is indicated on the y axis and the ARG types are colour-coded in the key.
Extended Data Figure 5 PCoA analysis plots of Bray–Curtis distances between soil resistomes.
The PCoA was calculated using all ORFs captured from functional selections without trimethoprim and D-cycloserine, and shows significant separation between CC (red) and KBS (blue) resistomes (P < 10−5, ANOSIM).
Extended Data Figure 6 PCoA across CC (red, grassland) and KBS (blue, agricultural) soils.
a–c, PCoA generated from all 16S data available from ref. 8, using Bray–Curtis (a), weighted Unifrac (b) and unweighted Unifrac (c) dissimilarity metrics. Samples cluster by soil location and N level, as previously demonstrated. d–f, The same PCoA plots generated using only samples with sufficient 16S and resistome data (that is, those used in Procrustes and Mantel analyses). Excluding the two high-N KBS soils with insufficient resistome data eliminates the clustering pattern observed for KBS soils in a–c. The asterisk denotes the high-N KBS soil common to both sets of analyses.
Extended Data Figure 7 Phylum level relative abundance of combined CC and KBS data sets for major soil bacteria.
a, 16s rRNA data are depicted in black. Phylogenetic inferences based on the sequence composition of the assembled, resistance-conferring DNA fragments are depicted in red. The relative abundances of Actinobacteria and Acidobacteria represent the largest discrepancies between data sets. b, Actinobacteria are most dramatically enriched in resistance-conferring DNA fragments, in accord with their role in producing antibiotics, but despite their high GC-content and predicted transcriptional incompatibilities with E. coli. Levels of Proteobacteria, the phylum to which E. coli belongs, are largely unchanged following functional selection, suggesting that any potential bias introduced to the selections by heterologous expression in E. coli is minimal compared to the effect of ARG-content of the source organisms.
Extended Data Figure 8 Procrustes analysis demonstrates that when soils cluster by bacterial composition, resistomes aggregate with phylogenetic groupings.
a–c, Procrustes analysis of the ARG content (Bray–Curtis) of CC (red) and KBS (blue) soils compared to community composition calculated by Bray–Curtis (a), weighted Unifrac (b) and unweighted Unifrac (c) dissimilarity metrics. d–f, The same Procrustes transformations for CC soils only. For a given soil, black lines connect to functional resistome data while the green lines connect to points generated from 16S gene sequence data. The M2 fit reported is from a Procrustes transformation over the first two principal coordinates while the P-value is calculated from a distribution of empirically determined M2 values over 10,000 Monte Carlo label permutations. For M2/P values calculated using all principal coordinates, refer to Supplementary Table 8.
Extended Data Figure 9 Procrustes analysis demonstrates that when soils do not form distinct phylogenetic clusters, we are unable to detect significant correlation between ARG content and phylogenetic architecture.
See Extended Data Fig. 6 for the phylogenetic relationships between these soils. a–c, Procrustes analysis of the ARG content (Bray–Curtis) of KBS (agricultural, blue) soils compared to 16S rRNA gene sequence using unweighted Unifrac (a), weighted Unifrac (b) and Bray–Curtis (c) similarity metrics. d–f, The same Procrustes transformations for the CC soils (grassland, red) without high-N amendment, showing that soil groupings must be distinguishable by bacterial composition to detect correlations with resistome content, regardless of soil type. For a given soil, black lines connect to functional resistome data while the green lines connect to points generated from 16S rRNA gene sequence data. The M2 fit reported is from a Procrustes transformation over the first two principal coordinates while the P value is calculated from a distribution of empirically determined M2 values over 10,000 Monte Carlo label permutations.
Extended Data Figure 10 Histogram of nucleotide per cent identity from pairwise alignments of all predicted mobility elements, suggesting that assembly does not inappropriately condense mobile DNA elements into too few sequences.
The blue trace depicts a normal distribution with the same mean and standard deviation empirically observed across all pairwise comparisons (n = 666).
Supplementary information
Supplementary information
This file contains Supplementary Tables 1-9. (PDF 225 kb)
Supplementary Data 1
This file contains all recovered contigs, their ORFs, annotations, and information regarding the samples from which each contig was derived. (XLSX 364 kb)
Supplementary Data 2
This file outlines all 433 human pathogen genomes and 153 non-pathogenic soil genomes used in comparisons of resistance gene mobility in these different environments. (XLSX 33 kb)
Rights and permissions
About this article
Cite this article
Forsberg, K., Patel, S., Gibson, M. et al. Bacterial phylogeny structures soil resistomes across habitats. Nature 509, 612–616 (2014). https://doi.org/10.1038/nature13377
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13377
This article is cited by
-
Longitudinal dynamics of farmer and livestock nasal and faecal microbiomes and resistomes
Nature Microbiology (2024)
-
Antibiotic- and metal-resistant endophytes inhabit Armeria maritima hyperaccumulator
Plant and Soil (2024)
-
Impact of wastewater treatment plant effluent discharge on the antibiotic resistome in downstream aquatic environments: a mini review
Frontiers of Environmental Science & Engineering (2024)
-
The priority of yeast to select among various DNA options to repair genome breaks by homologous recombination
Molecular Biology Reports (2024)
-
Deciphering the Role of WWTPs in Cold Environments as Hotspots for the Dissemination of Antibiotic Resistance Genes
Microbial Ecology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.