Abstract
Drug resistance is a major hurdle in oncology. Responses of acute myeloid leukaemia (AML) patients to cytarabine (Ara-C)-based therapies are often short lived with a median overall survival of months1,2,3,4. Therapies are under development to improve outcomes and include targeting the eukaryotic translation initiation factor (eIF4E) with its inhibitor ribavirin5,6,7. In a Phase II clinical trial in poor prognosis AML5, ribavirin monotherapy yielded promising responses including remissions; however, all patients relapsed. Here we identify a novel form of drug resistance to ribavirin and Ara-C. We observe that the sonic hedgehog transcription factor glioma-associated protein 1 (GLI1) and the UDP glucuronosyltransferase (UGT1A) family of enzymes are elevated in resistant cells. UGT1As add glucuronic acid to many drugs, modifying their activity in diverse tissues8. GLI1 alone is sufficient to drive UGT1A-dependent glucuronidation of ribavirin and Ara-C, and thus drug resistance. Resistance is overcome by genetic or pharmacological inhibition of GLI1, revealing a potential strategy to overcome drug resistance in some patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Deschler, B., de Witte, T., Mertelsmann, R. & Lubbert, M. Treatment decision-making for older patients with high-risk myelodysplastic syndrome or acute myeloid leukemia: problems and approaches. Haematologica 91, 1513–1522 (2006)
Deschler, B. & Lubbert, M. Acute myeloid leukemia: epidemiology and etiology. Cancer 107, 2099–2107 (2006)
Pulsoni, A. et al. Survival of elderly patients with acute myeloid leukemia. Haematologica 89, 296–302 (2004)
Hiddemann, W. et al. Management of acute myeloid leukemia in elderly patients. J. Clin. Oncol. 17, 3569–3576 (1999)
Assouline, S. et al. Molecular targeting of the oncogene eIF4E in acute myeloid leukemia (AML): a proof-of-principle clinical trial with ribavirin. Blood 114, 257–260 (2009)
Kentsis, A., Topisirovic, I., Culjkovic, B., Shao, L. & Borden, K. L. Ribavirin suppresses eIF4E-mediated oncogenic transformation by physical mimicry of the 7-methyl guanosine mRNA cap. Proc. Natl Acad. Sci. USA 101, 18105–18110 (2004)
Kraljacic, B. C., Arguello, M., Amri, A., Cormack, G. & Borden, K. Inhibition of eIF4E with ribavirin cooperates with common chemotherapies in primary acute myeloid leukemia specimens. Leukemia 25, 1197–1200 (2011)
Tukey, R. H. & Strassburg, C. P. Human UDP-glucuronosyltransferases: metabolism, expression, and disease. Annu. Rev. Pharmacol. Toxicol. 40, 581–616 (2000)
Jarvis, S. M., Thorn, J. A. & Glue, P. Ribavirin uptake by human erythrocytes and the involvement of nitrobenzylthioinosine-sensitive (es)-nucleoside transporters. Br. J. Pharmacol. 123, 1587–1592 (1998)
Willis, R. C., Carson, D. A. & Seegmiller, J. E. Adenosine kinase initiates the major route of ribavirin activation in a cultured human cell line. Proc. Natl Acad. Sci. USA 75, 3042–3044 (1978)
Cancer Genome Atlas Research. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013)
Mas, C. & Ruiz i Altaba, A. Small molecule modulation of HH-GLI signaling: current leads, trials and tribulations. Biochem. Pharmacol. 80, 712–723 (2010)
LoRusso, P. M. et al. Pharmacokinetic dose-scheduling study of hedgehog pathway inhibitor vismodegib (GDC-0449) in patients with locally advanced or metastatic solid tumors. Clin. Cancer Res. 17, 5774–5782 (2011)
Lauth, M., Bergstrom, A., Shimokawa, T. & Toftgard, R. Inhibition of GLI-mediated transcription and tumor cell growth by small-molecule antagonists. Proc. Natl Acad. Sci. USA 104, 8455–8460 (2007)
Dutton, G. Glucuronidation of Drugs and Other Compounds Ch. 1 (CRC, 1980)
Olah, E. et al. Molecular mechanisms in the antiproliferative action of taxol and tiazofurin. Anticancer Res. 16, 2469–2477 (1996)
Fisher, M. B., Campanale, K., Ackermann, B. L., VandenBranden, M. & Wrighton, S. A. In vitro glucuronidation using human liver microsomes and the pore-forming peptide alamethicin. Drug Metab. Dispos. 28, 560–566 (2000)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Culjkovic-Kraljacic, B., Baguet, A., Volpon, L., Amri, A. & Borden, K. L. B. The oncogene eIF4E reprograms the nuclear pore complext to promote mRNA export and oncogenic transformation. Cell Reports 2, 207–215 (2012)
Engtrakul, J. J., Foti, R. S., Strelevitz, T. J. & Fisher, M. B. Altered AZT (3′-azido-3′-deoxythymidine) glucuronidation kinetics in liver microsomes as an explanation for underprediction of in vivo clearance: comparison to hepatocytes and effect of incubation environment. Drug Metab. Dispos. 33, 1621–1627 (2005)
Barbier, O. et al. 3′-azido-3′-deoxythimidine (AZT) is glucuronidated by human UDP-glucuronosyltransferase 2B7 (UGT2B7). Drug Metab. Dispos. 28, 497–502 (2000)
Cheson, B. D. et al. Revised recommendations of the International Working Group for Diagnosis, Standardization of Response Criteria, Treatment Outcomes, and Reporting Standards for Therapeutic Trials in Acute Myeloid Leukemia. J. Clin. Oncol. 21, 4642–4649 (2003)
Horny, H. P., Wehrmann, M., Steinke, B. & Kaiserling, E. Assessment of the value of immunohistochemistry in the subtyping of acute leukemia on routinely processed bone marrow biopsy specimens with particular reference to macrophage-associated antibodies. Hum. Pathol. 25, 810–814 (1994)
Acknowledgements
We are grateful for reading of the manuscript by A. Kentsis, M. Osborne, F. Pettersson, S. del Rincon and C. Perreault. We are grateful for the use of Pharmascience facilities thanks to M. Goodman and D. Goodman, for specimens from the Banques de Cellules Leucémiques Du Québec (BCLQ), and for technical discussions with M. Ouellet. We thank nursing and support staff at all the clinical sites, and A. Ramteke for technical support. K.L.B.B. is supported by funds from the National Institutes of Health (RO1 80728 and 98571), IRICoR and Translational Research Program grants from the Leukemia and Lymphoma Society USA. K.L.B.B. holds a Canada Research Chair and H.A.Z. holds a Cole Foundation Fellowship and a CNRS Lebanon Fellowship.
Author information
Authors and Affiliations
Contributions
H.A.Z. and B.C.-K. designed and performed experiments, analysed data and wrote the manuscript; L.C., A.A. and G.C. performed experiments; P.G., S.P. and C.T.J. analysed data; A.A.R. performed experiments and analysed data; J.B.J. and S.J.M. designed experiments; S.A. was medical monitor for the trial, treated patients and analysed clinical data; E.C. coordinated clinical trials and analysed clinical data; B.L., J.B. and M.W.B. treated patients, provided specimens and analysed clinical data; W.H.M. designed experiments, analysed data and edited the manuscript; K.L.B.B. designed experiments, analysed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.J.M., J.B.J. and A.A.R. are/were employees of Pharmascience.
Extended data figures and tables
Extended Data Figure 1 Ribavirin resistance in some ribavirin monotherapy clinical trial patients.
Bone marrow biopsies for responding (patient 8 achieved a PR; patient 11, a CR, is shown in Fig. 1a) and non-responding patients (patient 9 was a PD and patient 13 a SD). Note abundance of blasts before treatment and at EOT and reduction in blasts and restoration of haematopoiesis during response. For Wright–Giemsa-stained samples magnification was ×50 with oil (patients 8 and 13), and for lysozyme staining ×20 (patient 9).
Extended Data Figure 2 Characterization of FaDu- and THP-1-derived resistant cell lines.
a, Detailed growth curves for FRI and FRII resistant cells. All cell lines have indistinguishable doubling times in the absence of ribavirin (far right panel). b, THP-1 resistant (TR) cell lines are not sensitive to treatment with ribavirin at the doses and times used. Ribavirin no longer targets eIF4E activity (that is, Mcl-1) in resistant cells (far right panel). There were no changes in eIF4E levels between resistant and parental cell lines (and Fig. 1c). Actin provides a loading control. c, Resistance is retained after 6 months of growth in the absence of ribavirin. d, Incubation of live cells with 3H-ribavirin indicates that THP-1 resistant cells have impaired uptake of ribavirin similar to FRI cells. e, eIF4E cap binding and eIF4G binding activity are retained in FRII cells. f, FRII cells are sensitive to eIF4E knockdown measured by cell growth. g, Effects of RNAi-mediated knockdown of GLI1 or eIF4E on UGT1A levels. Western blots were probed as indicated. RNAi-mediated knockdown of GLI1 led to reduced levels of UGT1A whereas knockdown of eIF4E did not. For UGT1A, a pan-UGT1A antibody was used. Antibody controls for UGT1A and GLI1 are shown in Extended Data Fig. 8c. Results are representative of at least three independent experiments. Average values are reported and error bars indicate ±standard deviations. Experiments were carried out in triplicate, three independent times. Western blots are representative of at least three independent experiments.
Extended Data Figure 3 Pro-drug metabolism is impaired in type I resistance.
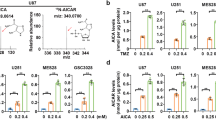
a, b, Western analysis reveals that ADK levels were reduced in FRI cells (a) and THP1 resistant cells (b). Treatments were 48 h at 20 µM ribavirin. c, d, Knockdown of ADK leads to ribavirin resistance as shown by cell growth. Western blot confirms knockdown of ADK. Hsp90 provides a loading control. e, f, ENT1 and ADK mRNA levels for patients’ specimens. Patient 11 (CR) was responding clinically at (and before) 84 and 112 days and relapsed around day 252, when both ADK and ENT1 mRNA levels decreased. Analysis of RNA samples isolated before and at the end of the first 28-day cycle for patient 9 (who did not respond to ribavirin) compared to a healthy volunteer. Averaged values for ADK and ENT1 RNAs were normalized to glucose 6-phosphate dehydrogenase (G6PDH). Error bars indicate ± s.d.; centre values are averages. All experiments were performed in triplicate at least three independent times. **P < 0.01, ***P < 0.001 (two-tailed Student’s t-test). Results are representative of at least three independent experiments.
Extended Data Figure 4 Confocal micrographs of leukaemic blasts isolated from bone marrows of responding and non-responding patients before treatment, at response or at EOT.
Immunostaining for GLI1 and UGT1A are shown. DAPI is in blue. Note nuclear accumulation of GLI1 in non-responding patients, indicating elevated GLI1 activity. High levels of GLI1 and UGT1A suggest primary resistance. All confocal settings were identical between specimens and thus lower signal is indicative of less protein. A ×100 objective with no digital zoom was used for patients 9, 11, 13, 17 and 19. The same objective but a digital zoom of ×2 was used for patients 8, A, B and C. Note patient C was still in remission at EOT (see main text). For each patient, staining was carried out three independent experiments. Controls for specificity of GLI1 and UGT1A antibodies are provided in Extended Data Fig. 8c.
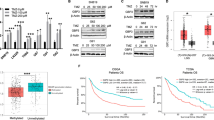
Extended Data Figure 5 Higher GLI1 expression is found in poor cytogenetic risk group and predicts a trend of worse survival outcome in AML.
To study the prognostic value of GLI1 gene expression in AML, we mined the publicly available AML data set published by The Cancer Genome Atlas Research Network11. a, GLI1 gene expression in 176 de novo AML patients grouped by cytogenetic risks. The expression level is represented by RPKM value (reads per kilobase of transcript per million mapped reads in RNA-seq). Each patient is represented by a symbol. Error bars represent median ± IQR (interquartile range) of each group. Nonparametric Mann–Whitney U-test was used to analyse the differences between groups. A total of 176 de novo AML patients with complete mRNA-seq and cytogenetic risk classification data are included in this analysis. b, c, Kaplan–Meier plots of events-free survival (EFS) and overall survival (OS) of 168 de novo AML patients segregated by median GLI1 expression (RPKM = 0.8596) (b) or high GLI1 expression (RPKM greater than or equal to 2) (c). Each tick on the survival curve represents a censored event because the patient is still alive at the end of the TCGA study. A total of 168 de novo AML patients with complete mRNA-seq, and reliable EFS and OS data are included in this analysis (patient information details are described in the Supplementary Table 1 of the NEJM study11. Mantel–Cox test was performed to calculate log-rank P values. We also observed that abnormally low levels of GLI1 were also correlated with poor outcome (data not shown), suggesting that GLI1 levels must be in a ‘Goldilocks’ zone.
Extended Data Figure 6 Effects of modulation of GLI1 levels on UGT1A.
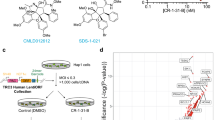
a, Effects of the direct GLI1 inhibitor GANT61 on restoring ribavirin sensitivity (20 μM) in FRII cells. Effects are dependent on GANT61 dose. b–d, Controls for eIF4E–ribavirin immunoprecipitations (IP) shown in Figs 1e and 3b. Inputs, supernatants (Sn) and IP controls for 3H ribavirin anti-eIF4E IPs are shown for GDC-0449-treated cells (b), FRII and FRI cells (c), and RNAi-mediated knockdown of GLI1 (d). e–g, qPCR analysis of GLI1 (e) and UGT1A (f) using a pan-UGT1A primer or primers for specific UGT1As (g). mRNA levels were normalized to RNA polymerase IIa. These findings are consistent with Extended Data Table 1, which indicates lower levels of UGT1A mRNA levels. Further, UGT1A3 and UGT1A8 decreased similarly (data not shown). Experiments were carried out in triplicate, at least three independent times. Average values are reported and error bars indicate standard deviations. These findings, that GLI1 elevation leads to reduced mRNA levels but increased protein levels, are counterintuitive. We propose that GLI1 elevation increases protein stability of UGT1As (see below) and this leads to some sort of feedback mechanism leading to reduced UGT1As. Other scenarios are possible but the main point that GLI1 elevation leads to increased UGT1A protein production is clear. h, GLI1 increases UGT1A protein stability as shown by studies with the proteasomal inhibitor MG132 (MG) and a pan-UGT1A antibody. Here, MG132 addition stabilizes levels of UGT1A in parental cells, but in FRII cells where levels are already increased, there is no further increase with MG132. This indicates that UGT1A proteins are already stabilized in the FRII cells. All results are representative of three independent experiments. i, Western blot analysis with a pan-UGT2B antibody indicates that UGT2B levels are unchanged in FRII relative to FaDu cells,suggesting the glucuronidation effects are mediated mostly through the UGT1A family. 293T cells are shown for comparison. Tubulin provides a loading control.
Extended Data Figure 7 MS analysis of ribavirin and Ara-C glucuronidation.
a, MS/MS collision-induced fragmentation analysis indicates that a breakdown product of the ribavirin glucuronide missing the ribose ring (exact mass 288.07) was further fragmented into a fragment of this glucuronide (exact mass 244.08, red asterisk) and to the triazole ring, the key moiety of ribavirin (exact mass 112.04). No ribose-glucuronide or ribose fragment was detected in our experiments, suggesting that this is not a major glucuronidation site in these cells. However, we cannot rule out that this exists and could not be detected. b, Microsomes expressing UGT1A1, UGT1A4, UGT1A6 and UGT1A9 were treated with RTP, underwent hydrophilic interaction liquid chromatography (HILIC) and the resulting extracted ion chromatogram (EIC) is shown. The Rib-Glu peak is clearly present and fragmentation analysis as in a confirms that this is glucuronidated ribavirin. We note that microsomes only expressing UGT1A1 do not glucuronidate RTP; and that RTP, but not ribavirin, is glucuronidated in microsomes. These studies suggest that UGT1A4, UGT1A6 and/or UGT1A9 are required for glucuronidation, as is some phosphorylation event before glucuronidation. c, Using HILIC chromatography, we isolated the fraction containing the Rib-Glu peak in b. A portion of this was re-assessed by MS/MS to be sure that the correct peak was isolated. This material was used in the 3H-ribavirin competition assay in Fig. 4m. Material was quantified using a standard curve of ribavirin (see Methods). d, Western blot demonstrating equal loading of eIF4E–GST in the 3H-ribavirin pulldown assay shown in Fig. 4m. All results are representative of at least three independent experiments. e, AraC is glucuronidated (AraC-Glu) in FRII cells but not parental FaDu cells where AraC-TP (triphosphate) is observed. AraC-TP is also observed in FRII cells, but at much lower levels than AraC-Glu. Treatment of FRII cells with GDC-0449 results in the loss of the AraC-Glu peak and causes no alteration to the parental FaDu cells. Fragmentation strongly suggests that the cytosine is the major site of glucuronidation (data not shown). We did not observe masses consistent with an arabinose breakdown product or an arabinose-glucuronide but cannot rule out that they are present at low levels or that our isolation procedure precluded their detection. f, Structures of AraC and AraC-TP are shown. The red arrow indicating the most likely glucuronidation site, as per our mass spectrometry data. Note that no glucuronides were observed when reactions were incubated in the absence of UDP-glucuronic acid (data not shown).
Extended Data Figure 8 Investigations into why GLI1 levels are elevated in FRII cells.
a, Analysis of expression of a subset of hedgehog signalling pathway proteins. Western blots are probed as indicated and are representative of three independent experiments. Tubulin and Hsp90 provided loading controls. Patched 1 (PTCH1) was the most significant change. PTCH1 is 210 kDa, with an often observed degradation product at 170 kDa. IHH, Indian hedgehog; SHH, sonic hedgehog, Smo, smoothened. b, GLI1 DNA methylation. CpG methylation was interrogated on bisulphite-converted DNA from GLI1 promoter region and first exon. The amplicon covered 29 CpGs, 25 of them located within a CpG island. DNA CpG methylation is shown as per cent methylation for FaDu (top) and FRII (bottom) cells. There was no difference observed between the cell lines. c, Antibody controls. Analysis of GLI1 and pan-UGT1A antibodies as a function of RNAi-mediated knockdown of these proteins as indicated. Note that UGT1A family members have approximately the same molecular weight. Results are representative of at least three independent experiments.
Rights and permissions
About this article
Cite this article
Zahreddine, H., Culjkovic-Kraljacic, B., Assouline, S. et al. The sonic hedgehog factor GLI1 imparts drug resistance through inducible glucuronidation. Nature 511, 90–93 (2014). https://doi.org/10.1038/nature13283
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13283
This article is cited by
-
1H, 13C, 15N Backbone and sidechain chemical shift assignments of the C-terminal domain of human UDP-glucuronosyltransferase 2B17 (UGT2B17-C)
Biomolecular NMR Assignments (2023)
-
Chemotherapy-related hyperbilirubinemia in pediatric acute lymphoblastic leukemia: a genome-wide association study from the AIEOP-BFM ALL study group
Journal of Experimental & Clinical Cancer Research (2023)
-
The NSAID glafenine rescues class 2 CFTR mutants via cyclooxygenase 2 inhibition of the arachidonic acid pathway
Scientific Reports (2022)
-
1H, 13C and 15N chemical shift assignments of the C-terminal domain of human UDP-Glucuronosyltransferase 2B7 (UGT2B7-C)
Biomolecular NMR Assignments (2021)
-
Up-regulation of GLI1 in vincristine-resistant rhabdomyosarcoma and Ewing sarcoma
BMC Cancer (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.