Abstract
The capacity of numerous bacterial species to tolerate antibiotics and other toxic compounds arises in part from the activity of energy-dependent transporters. In Gram-negative bacteria, many of these transporters form multicomponent ‘pumps’ that span both inner and outer membranes and are driven energetically by a primary or secondary transporter component1,2,3,4,5,6,7. A model system for such a pump is the acridine resistance complex of Escherichia coli1. This pump assembly comprises the outer-membrane channel TolC, the secondary transporter AcrB located in the inner membrane, and the periplasmic AcrA, which bridges these two integral membrane proteins. The AcrAB–TolC efflux pump is able to transport vectorially a diverse array of compounds with little chemical similarity, thus conferring resistance to a broad spectrum of antibiotics. Homologous complexes are found in many Gram-negative species, including in animal and plant pathogens. Crystal structures are available for the individual components of the pump2,3,4,5,6,7 and have provided insights into substrate recognition, energy coupling and the transduction of conformational changes associated with the transport process. However, how the subunits are organized in the pump, their stoichiometry and the details of their interactions are not known. Here we present the pseudo-atomic structure of a complete multidrug efflux pump in complex with a modulatory protein partner8 from E. coli. The model defines the quaternary organization of the pump, identifies key domain interactions, and suggests a cooperative process for channel assembly and opening. These findings illuminate the basis for drug resistance in numerous pathogenic bacterial species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Atomic coordinates and structure factors for the reported crystal structures have been deposited in the PDB under accessions 4C48 (AcrB/AcrZ/DARPin) and 4CDI (AcrB/AcrZ). The cryo-EM map has been deposited in the Electron Microscopy Data Bank under accession EMD-5915.
References
Du, D., Venter, H., Pos, K. M. & Luisi, B. F. in Microbial Efflux Pumps: Current Research Ch. 3 (Caister Academic, 2013)
Koronakis, V., Sharff, A., Koronakis, E., Luisi, B. F. & Hughes, C. Crystal structure of the bacterial membrane protein TolC central to multidrug efflux and protein export. Nature 405, 914–919 (2000)
Murakami, S., Nakashima, R. & Yamashita, E. Cry & Yamaguchi, A. Crystal structure of bacterial multidrug efflux transporter AcrB. Nature 419, 587–593 (2002)
Mikolosko, J., Bobyk, K., Zgurskaya, H. I. & Ghosh, P. Conformational flexibility in the multidrug efflux system protein AcrA. Structure 14, 577–587 (2006)
Seeger, M. A. et al. Structural asymmetry of AcrB trimer suggests a peristaltic pump mechanism. Science 313, 1295–1298 (2006)
Murakami, S., Nakashima, R., Yamashita, E., Matsumoto, T. & Yamaguchi, A. Crystal structures of a multidrug transporter reveal a functionally rotating mechanism. Nature 443, 173–179 (2006)
Eicher, T. et al. Transport of drugs by the multidrug transporter AcrB involves an access and a deep binding pocket that are separated by a switch-loop. Proc. Natl Acad. Sci. USA 109, 5687–5692 (2012)
Hobbs, E. C., Yin, X., Paul, B. J., Astarita, J. L. & Storz, G. Conserved small protein associates with the multidrug efflux pump AcrB and differentially affects antibiotic resistance. Proc. Natl Acad. Sci. USA 109, 16696–16701 (2012)
Sennhauser, G., Amstutz, P., Briand, C., Storchenegger, O. & Grütter, M. G. Drug export pathway of multidrug exporter AcrB revealed by DARPin inhibitors. PLoS Biol. 5, e7 (2007)
Törnroth-Horsefield, S. et al. Crystal structure of AcrB in complex with a single transmembrane subunit reveals another twist. Structure 15, 1663–1673 (2007)
Janganan, T. K. et al. Evidence for the assembly of a bacterial tripartite multidrug pump with a stoichiometry of 3:6:3. J. Biol. Chem. 286, 26900–26912 (2011)
Stegmeier, J. F., Polleichtner, G., Brandes, N., Hotz, C. & Andersen, C. Importance of the adaptor (membrane fusion) protein hairpin domain for the functionality of multidrug efflux pumps. Biochemistry 45, 10303–10312 (2006)
Narita, S.-i., Eda, S., Yoshihara, E. & Nakae, T. Linkage of the efflux-pump expression level with substrate extrusion rate in the MexAB–OprM efflux pump of Pseudomonas aeruginosa. Biochem. Biophys. Res. Commun. 308, 922–926 (2003)
Mima, T., Joshi, S., Gomez-Escalada, M. & Schweizer, H. P. Identification and characterization of TriABC-OpmH, a triclosan efflux pump of Pseudomonas aeruginosa requiring two membrane fusion proteins. J. Bacteriol. 189, 7600–7609 (2007)
Su, C. C. et al. Crystal structure of the CusBA heavy metal efflux complex of Escherichia coli. Nature 470, 558–562 (2011)
Yum, S. et al. Crystal structure of the periplasmic component of a tripartite macrolide-specific efflux pump. J. Mol. Biol. 387, 1286–1297 (2009)
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nature Methods 5, 53–55 (2008)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Henderson, R. et al. Tilt-pair analysis of images from a range of different specimens in single-particle electron cryomicroscopy. J. Mol. Biol. 413, 1028–1046 (2011)
Berthaud, A., Manzi, J., Pérez, J. & Mangenot, S. Modeling detergent organization around aquaporin-O using small angle X-ray scattering. J. Am. Chem. Soc. 134, 10080–10088 (2012)
Symmons, M. F., Bokma, E., Koronakis, E., Hughes, C. & Koronakis, V. The assembled structure of a complete tripartite bacterial multidrug efflux pump. Proc. Natl Acad. Sci. USA 106, 7173–7178 (2009)
Touzé, T. et al. Interactions underlying assembly of the Escherichia coli AcrAB–TolC multidrug efflux system. Mol. Microbiol. 53, 697–706 (2004)
Xu, Y. et al. Assembly and channel opening of outer membrane protein in tripartite drug efflux pumps of Gram-negative bacteria. J. Biol. Chem. 287, 11740–11750 (2012)
Trépout, S. et al. Structure of reconstituted bacterial membrane efflux pump by cryo-electron tomography. Biochim. Biophys. Acta 1798, 1953–1960 (2010)
Tikhonova, E. B., Yamada, Y. & Zgurskaya, H. I. Sequential mechanism of assembly of multidrug efflux pump AcrAB–TolC. Chem. Biol. 18, 454–463 (2011)
Ge, Q., Yamada, Y. & Zgurskaya, H. The C-terminal domain of AcrA is essential for the assembly and function of the multidrug efflux pump AcrAB–TolC. J. Bacteriol. 191, 4365–4371 (2009)
Janganan, T. K., Bavro, V. N., Zhang, L., Borges-Walmsley, M. I. & Walmsley, A. R. Tripartite efflux pumps: energy is required for dissociation, but not assembly or opening of the outer membrane channel of the pump. Mol. Microbiol. 88, 590–602 (2013)
Bavro, V. N. et al. Assembly and channel opening in a bacterial drug efflux machine. Mol. Cell 30, 114–121 (2008)
Tikhonova, E. B. & Zgurskaya, H. I. AcrA, AcrB, and TolC of Escherichia coli form a stable intermembrane multidrug efflux complex. J. Biol. Chem. 279, 32116–32124 (2004)
Miroux, B. & Walker, J. E. Over-production of proteins in Escherichia coli: mutant hosts that allow synthesis of some membrane proteins and globular proteins at high levels. J. Mol. Biol. 260, 289–298 (1996)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D 62, 72–82 (2006)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Brunger, A. T. Version 1.2 of the Crystallography and NMR system. Nature Protocols 2, 2728–2733 (2007)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Murshudov, G. N., Vagin, A. A., Lebedev, A., Wilson, K. S. & Dodson, E. J. Efficient anisotropic refinement of macromolecular structures using FFT. Acta Crystallogr. D 55, 247–255 (1999)
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011)
Emsley, P., Lohkamp, B., Scott, W. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Welch, A., Awah, C. U., Jing, S., van Veen, H. W. & Venter, H. Promiscuous partnering and independent activity of MexB, the multidrug transporter protein from Pseudomonas aeruginosa. Biochem. J. 430, 355–364 (2010)
Lobedanz, S. et al. A periplasmic coiled-coil interface underlying TolC recruitment and the assembly of bacterial drug efflux pumps. Proc. Natl Acad. Sci. USA 104, 4612–4617 (2007)
Stark, H. GraFix: stabilization of fragile macromolecular complexes for single particle cryo-EM. Methods Enzymol. 481, 109–126 (2010)
Li, X. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nature Methods 10, 584–590 (2013)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Murray, S. C. et al. Validation of cryo-EM structure of IP3R1 channel. Structure 21, 900–909 (2013)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Pei, X. Y. et al. Structures of sequential open states in a symmetrical opening transition of the TolC exit duct. Proc. Natl Acad. Sci. USA 108, 2112–2117 (2011)
Tsukazaki, T. et al. Structure and function of a membrane component SecDF that enhances protein export. Nature 474, 235–238 (2011)
Hsieh, Y. H. et al. Reconstitution of functionally efficient SecA-dependent protein-conducting channels: transformation of low-affinity SecA-liposome channels to high-affinity SecA-SecYEG-SecDF·YajC channels. Biochem. Biophys. Res. Commun. 431, 388–392 (2013)
du Plessis, D. J., Nouwen, N. & Driessen, A. J. The Sec translocase. Biochim. Biophys. Acta 1808, 851–865 (2011)
Ashkenazy, H., Erez, E., Martz, E., Pupko, T. & Ben-Tal, N. ConSurf 2010: calculating evolutionary conservation in sequence and structure of proteins and nucleic acids. Nucleic Acids Res. 38, W529–W533 (2010)
Jiang, W. et al. Bridging the information gap: computational tools for intermediate resolution structure interpretation. J. Mol. Biol. 308, 1033–1044 (2001)
Acknowledgements
This work was supported by the Wellcome Trust and Human Frontier Science Program (B.L.) and partly by a National Institutes of Health grant (P41GM103832 to W.C.). J.E.V. is supported by a Herschel Smith scholarship. T.O.-A. is the recipient of a Cambridge Trust scholarship, an Adam Glinsman award and a Faculty for the Future Fellowship from the Schlumberger Foundation. We thank M. Pos for kindly providing pBAD-AcrAB-TolC. We thank L. Packman for mass-spectrometric analyses and S. J. Ludtke, M. F. Schmid, M. Pos and R. van Veen for helpful advice and discussions. We are grateful to the staff at the Diamond Light Source for access to facilities and invaluable help.
Author information
Authors and Affiliations
Contributions
D.D., W.C. and B.F.L. designed the experiments; D.D. and N.R.J. purified and crystallized AcrBZ complexes. D.D. and B.F.L. solved the crystal structure of AcrBZ complexes. D.D., N.R.J. and E.K. purified AcrABZ–TolC complexes. Z.W., J.E.V. and W.C. obtained and analysed the single-particle cryo-EM data. D.D. and B.F.L. devised a model of AcrABZ–TolC based on the cryo-EM map. T.O.-A. and H.V. conducted MIC and efflux assays on the AcrABZ–TolC pump. D.D., J.E.V., W.C. and B.F.L. analysed results. All authors contributed to writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Co-purification of the AcrBZ and AcrBZ–DARPin complexes.
a, Gel filtration profile of AcrBZ complex and SDS–polyacrylamide gel electrophoresis (SDS–PAGE) analysis of the peak fractions. b, Gel filtration profile of the AcrBZ–DARPin complex and SDS–PAGE analysis. The proteins were enriched by immobilized metal affinity chromatography (IMAC; results not shown) before the size-exclusion chromatography step. AcrZ has a heptahistidine tag on the C terminus as an IMAC affinity tag, and the C-terminal histidines of AcrB have been removed to prevent its binding to the matrix. Standard proteins thyrogloblin (669 kDa) and aldolase (158 kDa) (Bio-Rad) were eluted from the same column at volumes of 11.4 and 14.7 ml, respectively. See also Fig. 1.
Extended Data Figure 2 Validation of the AcrBZ crystal structure.
An AcrB protomer taken from the refined structure of the AcrBZ–DARPin complex at a resolution of 3.3 Å was used in molecular replacement of the 3.7 Å data of the AcrBZ complex (Extended Data Table 1). The model of AcrB without AcrZ was refined with REFMAC38 jelly-body and reference structure restraints, and a difference map calculated in which AcrZ does not contribute to the phases. Shown are the positive features of the difference map in the vicinity of AcrZ (green carbon atoms) taken by superposing the AcrB protomers from the AcrBZ–DARPin and AcrBZ (red) structures. The unbiased map shows the presence of AcrZ and indicates that the presence of DARPin does not disrupt the interactions with AcrB. The model of the AcrBZ–DARPin complex at a resolution of 3.3 Å was refined without AcrZ to generate a simulated annealing omit map, which confirmed the location of AcrZ (data not shown). See also Fig. 1.
Extended Data Figure 3 The AcrBZ complex resembles AcrB–YajC, and AcrBZ interactions involve conserved residues.
a, Superposition of AcrB–YajC (orange and yellow; PDB accession 2RDD) onto AcrBZ (red and green). See also Fig. 1. We were not able to identify an interaction between AcrB and C-terminal histidine-tagged YajC from E. coli, suggesting that the interaction is less avid than the AcrB–AcrZ pairing (data not shown). It is interesting to note that the contacting surface is also conserved in another RND protein, SecDF, which is involved in protein translocation and membrane insertion48. Therefore, it seems likely that SecDF might be modulated by a similar helical peptide. Indeed, YajC forms a complex with SecDF49,50 and could act as an allosteric modulator in protein biogenesis. b, Sequence conservation of AcrZ. The secondary structure is annotated at the top and asterisks indicate residues that directly contact AcrB. c, Sequence variation of the surface of AcrB homologues, showing conservation of the surface that contacts AcrZ. Right-hand side includes the bound AcrZ in stick representation (green). The colour spectrum ranges from purple (most conserved) to blue (least conserved). This figure was made using ConSurf51.
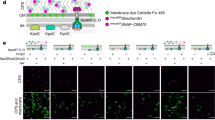
Extended Data Figure 4 Schematic representations and purification of the fusion proteins used to assemble the efflux pump.
a, The AcrA–AcrB fusion, with two flexible poly(Gly-Ser) linkers. The two C-terminal histidines of AcrB have been removed to prevent binding to the NTA matrix during co-purification. b, The AcrA–AcrZ–His5 fusion and TolC co-expression construct. The numbers above the bars correspond to the residues of the protein, and, owing to the restriction site used for cloning, a single glycine residue was inserted after the start codon in both AcrA and AcrZ. The flexible poly(Gly-Ser) linker permits the protomers to manoeuvre. c, Co-purification of TolC with the AcrABZ complexes. SDS–PAGE of the eluate from gel filtration following nickel affinity chromatography purification. See also Fig. 2.
Extended Data Figure 5 The fusion assemblies can drive efflux in vivo.
a, b, Drug-transport assays of the AcrAB and AcrAZ fusion proteins in the MCΔtolCΔacrAB strain show that the engineered pump retains partial activity for efflux of ethidium bromide (a) and trimethylammonium-diphenylhexatriene (b). The results show representative traces from six biological replicates. For ethidium bromide, initial influx rates of 10.3 ± 0.6, 6.4 ± 0.6 and 3.6 ± 0.4 arbitrary units (AU) per min were obtained for the non-expressing control cells, the cells expressing both fusion proteins plus TolC and the wild-type cells expressing AcrAB–TolC, respectively. For trimethylammonium-diphenylhexatriene, initial influx rates of 187 ± 6, 151 ± 3 and 54 ± 2 AU min−1 were obtained for the non-expressing control cells, the cells expressing both fusion proteins plus TolC and the wild-type cells expressing AcrAB–TolC, respectively. c, MIC data on antimicrobial susceptibility of E. coli MCΔtolCΔacrAB cells expressing wild-type AcrA–AcrB–TolC or the fusion proteins with TolC. These data indicate that the fusion of AcrA–AcrZ and the insertion of AcrA into AcrB both diminish the capacity for drug resistance. The plasmids pET21a, pET21a-AcrB or pET21a-acrB328-polyGS-acrA-polyGlySer-acrB329 (AcrAB), together with plasmid pRSFDuet-1-acrA-polyGlySer-acrZHis5-tolC (AcrAZ + TolC), were transformed into MCΔtolCΔacrAB. Cells were tested for their ability to resist increasing concentrations of cytotoxic drugs. As a positive control, cells were transformed with plasmid pBAD_AcrA+AcrB+TolC, which encodes for the native AcrA–AcrB–TolC efflux pump.
Extended Data Figure 6 Electron microscopy images and class averages of the AcrABZ–TolC complex.
a, Negative-stain electron microscopy images of the purified AcrABZ–TolC complex after GraFix treatment. White circles indicate particles with long axis almost normal to the viewing plane; black circles show particles with the long axis parallel to the viewing plane. b, c, Class averages from cryo-EM data and reconstructions with C1 symmetry and with C3 symmetry, respectively. The galleries in the side panels show representative two-dimensional class averages of the purified pump. The top shows views perpendicular to the long axis of the drumstick shape, the middle shows inclined views, and the bottom shows views along the long axis. The reconstructed images and cross-sections indicate the presence of a six-fold symmetry, which is consistent with six AcrA protomers. The pseudo-atomic model has three protomers each of AcrB, AcrZ and TolC, and as TolC and AcrB each have an internal structural repeat, they have pseudo six-fold symmetry at low resolution (cut view). The maps are consistent with each other. See also Fig. 2.
Extended Data Figure 7 Resolution estimation and validation of the cryo-EM map.
a, Fourier shell correlation (FSC) of two independently determined cryo-EM maps of the efflux pump assembly after their alignment by Foldhunter52. b, Tilt-pair validation of the efflux pump assembly. Each point represents a pair of particles with experimentally known relative tilt. The radius indicates the computationally determined amount of tilt, and the azimuth indicates tilt direction. The red circle denotes particle pairs that cluster around the experimental tilt axis geometry, thus validating the map in Fig. 2.
Extended Data Figure 8 Sections through the electron microscopy map.
a, Slices 1–4: view of planes normal to the three-fold symmetry axis of the pump. b, Slice 5 of the model assembly showing the opening of the TolC periplasmic domain. The same transverse plane through the cryo-EM map is shown with TolC in the closed state, as seen in the crystal structure of the isolated protein (left; PDB accession 1EK9), the partially open structure, made by mutations (middle; PDB accession 2XMN) and the modelled fully opened structure (right).
Rights and permissions
About this article
Cite this article
Du, D., Wang, Z., James, N. et al. Structure of the AcrAB–TolC multidrug efflux pump. Nature 509, 512–515 (2014). https://doi.org/10.1038/nature13205
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13205
This article is cited by
-
Escherichia coli resistance mechanism AcrAB-TolC efflux pump interactions with commonly used antibiotics: a molecular dynamics study
Scientific Reports (2024)
-
Antimicrobial, antiproliferative activities and molecular docking of metabolites from Alternaria alternata
AMB Express (2023)
-
Molecular mechanisms of antibiotic resistance revisited
Nature Reviews Microbiology (2023)
-
Di-berberine conjugates as chemical probes of Pseudomonas aeruginosa MexXY-OprM efflux function and inhibition
npj Antimicrobials and Resistance (2023)
-
Scaffold size-dependent effect on the enhanced uptake of antibiotics and other compounds by Escherichia coli and Pseudomonas aeruginosa
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.