Abstract
Mitochondrial function is challenged by toxic by-products of metabolism as well as by pathogen attack1,2. Caenorhabditis elegans normally responds to mitochondrial dysfunction with activation of mitochondrial-repair, drug-detoxification and pathogen-response pathways1,2,3,4,5,6,7. Here, from a genome-wide RNA interference (RNAi) screen, we identified 45 C. elegans genes that are required to upregulate detoxification, pathogen-response and mitochondrial-repair pathways after inhibition of mitochondrial function by drug-induced or genetic disruption. Animals defective in ceramide biosynthesis are deficient in mitochondrial surveillance, and addition of particular ceramides can rescue the surveillance defects. Ceramide can also rescue the mitochondrial surveillance defects of other gene inactivations, mapping these gene activities upstream of ceramide. Inhibition of the mevalonate pathway, either by RNAi or statin drugs, also disrupts mitochondrial surveillance. Growth of C. elegans with a significant fraction of bacterial species from their natural habitat causes mitochondrial dysfunction. Other bacterial species inhibit C. elegans defence responses to a mitochondrial toxin, revealing bacterial countermeasures to animal defence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Melo, J. A. & Ruvkun, G. Inactivation of conserved C. elegans genes engages pathogen- and xenobiotic-associated defenses. Cell 149, 452–466 (2012)
Runkel, E. D., Liu, S., Baumeister, R. & Schulze, E. Surveillance-activated defenses block the ROS-induced mitochondrial unfolded protein response. PLoS Genet. 9, e1003346 (2013)
Benedetti, C., Haynes, C. M., Yang, Y., Harding, H. P. & Ron, D. Ubiquitin-like protein 5 positively regulates chaperone gene expression in the mitochondrial unfolded protein response. Genetics 174, 229–239 (2006)
Haynes, C. M., Petrova, K., Benedetti, C., Yang, Y. & Ron, D. ClpP mediates activation of a mitochondrial unfolded protein response in C. elegans. Dev. Cell 13, 467–480 (2007)
Haynes, C. M., Yang, Y., Blais, S. P., Neubert, T. A. & Ron, D. The matrix peptide exporter HAF-1 signals a mitochondrial UPR by activating the transcription factor ZC376.7 in C. elegans.. Mol. Cell 37, 529–540 (2010)
Nargund, A. M., Pellegrino, M. W., Fiorese, C. J., Baker, B. M. & Haynes, C. M. Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 337, 587–590 (2012)
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011)
Pellegrino, M. W., Nargund, A. M. & Haynes, C. M. Signaling the mitochondrial unfolded protein response. Biochim. Biophys. Acta 1833, 410–416 (2013)
Estes, K. A., Dunbar, T. L., Powell, J. R., Ausubel, F. M. & Troemel, E. R. bZIP transcription factor zip-2 mediates an early response to Pseudomonas aeruginosa infection in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 107, 2153–2158 (2010)
Félix, M. A. & Duveau, F. Population dynamics and habitat sharing of natural populations of Caenorhabditis elegans and C. briggsae. BMC Biol. 10, 59 (2012)
Lee, S. S. et al. A systematic RNAi screen identifies a critical role for mitochondria in C. elegans longevity. Nature Genet. 33, 40–48 (2003)
Menuz, V. et al. Protection of C. elegans from anoxia by HYL-2 ceramide synthase. Science 324, 381–384 (2009)
Deng, X. et al. Ceramide biogenesis is required for radiation-induced apoptosis in the germ line of C. elegans. Science 322, 110–115 (2008)
Sentelle, R. D. et al. Ceramide targets autophagosomes to mitochondria and induces lethal mitophagy. Nature Chem. Biol. 8, 831–838 (2012)
Colombini, M. Ceramide channels and their role in mitochondria-mediated apoptosis. Biochim. Biophys. Acta 1797, 1239–1244 (2010)
Wenner Moyer, M. The search beyond statins. Nature Med. 16, 150–153 (2010)
Graham, D. J. et al. Incidence of hospitalized rhabdomyolysis in patients treated with lipid-lowering drugs. J. Am. Med. Assoc. 292, 2585–2590 (2004)
Rauthan, M., Ranji, P., Aguilera Pradenas, N., Pitot, C. & Pilon, M. The mitochondrial unfolded protein response activator ATFS-1 protects cells from inhibition of the mevalonate pathway. Proc. Natl Acad. Sci. USA 110, 5981–5986 (2013)
Hage-Sleiman, R., Esmerian, M. O., Kobeissy, H. & Dbaibo, G. p53 and ceramide as collaborators in the stress response. Int. J. Mol. Sci. 14, 4982–5012 (2013)
Senkal, C. E. et al. Alteration of ceramide synthase 6/C16-ceramide induces activating transcription factor 6-mediated endoplasmic reticulum (ER) stress and apoptosis via perturbation of cellular Ca2+ and ER/Golgi membrane network. J. Biol. Chem. 286, 42446–42458 (2011)
Breslow, D. K. & Weissman, J. S. Membranes in balance: mechanisms of sphingolipid homeostasis. Mol. Cell 40, 267–279 (2010)
Bookout, A. L. & Mangelsdorf, D. J. Quantitative real-time PCR protocol for analysis of nuclear receptor signaling pathways. Nucl. Recept. Signal. 1, e012 (2003)
Phillips, C. M., McDonald, K. L. & Dernburg, A. F. Cytological analysis of meiosis in Caenorhabditis elegans. Methods Mol. Biol. 558, 171–195 (2009)
Wagner, B. K. et al. A small-molecule screening strategy to identify suppressors of statin myopathy. ACS Chem. Biol. 6, 900–904 (2011)
Acknowledgements
We thank C. Haynes (Memorial Sloan Kettering Cancer Center), F. Ausubel (Massachusetts General Hospital) and the Caenorhabditis Genetics Center for providing strains; and the V. Mootha laboratory (Massachusetts General Hospital) for providing C2C12 cells. We thank J. Larkins-Ford, C. Phillips, S. Garcia and M. Pellegrino for technical support, as well as M. A. Félix for bacterial strains and for guidance in the collection of microbial strains from C. elegans natural habitats. Y.L. is supported by the Helen Hay Whitney research fellowship; B.S.S. is supported by the Charles King Trust postdoctoral fellowship. The work is supported by a grant from the National Institutes of Health, awarded to G.R. (NIH AG043184-16).
Author information
Authors and Affiliations
Contributions
Y.L. and G.R. designed experiments; Y.L., B.S.S. and P.C.B. carried out experiments. Y.L., B.S.S. and G.R. wrote the paper. G.R. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Mitochondrial dysfunction activates homeostatic, detoxification and pathogen responses.
a, Drug-induced food-avoidance phenotypes on live or dead bacteria. Dead bacteria were obtained by heating bacteria at 90 °C for 30 min. Avoidance behaviour was also observed when antimycin was added to dead bacteria, showing that the drug acts directly on C. elegans and is not transformed by the bacteria. b, Quantification of the food-avoidance phenotypes of Extended Data Fig. 1a (n = 4). Error bars represent s.d. c, A dose response of hsp-6p::gfp induction with the addition of antimycin.
Extended Data Figure 2 Diagram of the genome-wide RNAi screen workflow.
For the detailed experimental procedure, see Methods.
Extended Data Figure 4 sptl-1 is required for mitochondrial surveillance.
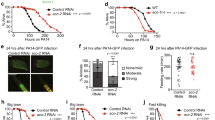
a, A graph showing the fold change in C. elegans sptl-1 transcript level compared to control (n = 3). Error bars represent s.d. b, hsp-6p::gfp worms raised on control or sptl-1(RNAi) (n = 40). c, Body wall muscle animals expressing a mitochondrially localized GFP reporter. The animals were subjected to sptl-1(RNAi) for 36 h and transferred onto the second RNAi (control, fzo-1, fis-1 or drp-1). The hyperfusion of mitochondria observed under sptl-1(RNAi) is dependent on the fusion machinery as disruption of mitochondrial fusion by inactivating fzo-1 or fis-1 partially restored the tubular structure. By contrast, inhibition of the gene drp-1 that governs fission led to mitochondrial hyperfusion. The images were taken 36 h after they were placed on the second RNAi. d, A graph showing the percentage of worms which avoid the spg-7(RNAi) bacteria lawn 48 h after they were initially placed on the plates (n = 4). Error bars represent s.d. e, A graph showing the percentage of worms that avoid the bacteria lawn 8 h after the addition of antimycin (n = 4). Error bars represent s.d. f, Wild-type N2 or sptl-1(ok1693) mutant animals raised in the presence or absence of 0.8 μM antimycin. Photos were taken 4 days after the synchronized L1 worms were placed on the plates. g, hsp-6p::gfp animals were raised on sptl-1(RNAi) for 36 h and transferred onto a subset of wild microbes. The images were taken after 2 days.
Extended Data Figure 5 Ceramide biosynthesis is required for mitochondrial surveillance.
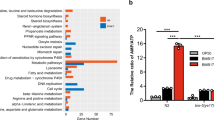
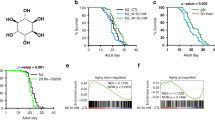
a, Genotyping of the sphingolipid metabolism pathway mutant alleles. b, c, Fold difference in hsp-6 transcript levels in wild type or sphingolipid metabolism pathway mutants (b) and hyl-1 or lagr-1 mutants (c) (n = 3). Error bars represent s.d. d, hsp-6p::gfp worms raised on sptl-1(RNAi) in the presence of increasing amounts of ceramide. e, hsp-6p::gfp in the presence or absence of ceramide.
Extended Data Figure 6 Ceramide biogenesis is required for mitochondrial surveillance.
a, The percentage of worms that avoid the bacterial lawn 8 h after the addition of antimycin. Animals were pre-treated with control RNAi, control RNAi with ceramide, sptl-1(RNAi) or sptl-1(RNAi) with ceramide (n = 4). Error bars represent s.d. b, Time course experiment for the induction of hsp-6p::gfp with antimycin. c, Dissected young adults after 4 h antimycin treatment were stained with anti-COX-IV antibody (green) and anti-ceramide antibody (red). d, Nomarski (top) and fluorescent (bottom) images of intestinal cells in atfs-1;hsp-16p::atfs-1Δ1-32.myc::gfp transgenic animals. e, hsp-6p::gfp worms raised on indicated RNAi in the presence or absence of ceramide.
Extended Data Figure 7 Inhibition of the mevalonate pathway disrupts mitochondrial surveillance.
a, hsp-6p::gfp animals raised on control or hmgs-1(RNAi) in the presence of antimycin. b, Immunoblotting of GFP expressed by hsp-6p::gfp animals, with or without antimycin. c, Antimycin-induced food avoidance in control or hmgs-1(RNAi) animals. d, Quantification of food avoidance (n = 4). Error bars represent s.d. e, Body wall muscle of control or hmgs-1(RNAi) animals expressing a mitochondrially localized GFP reporter. f, hsp-6p::gfp animals raised on hmgs-1(RNAi), or hmgs-1(RNAi) with addition of mevalonate exposed to antimycin. g, hsp-6p::gfp animals treated with antimycin after pre-treatment with simvastatin or mevastatin. h, Mitochondrial immunostaining in HEK293T cells. i, ATP levels in C2C12 myotubes after treating with simvastatin or mevastatin (n = 3). Error bars represent s.d.
Extended Data Figure 8 Statin treatment impairs mitochondrial surveillance.
a, Diagram of the mevalonate pathway for the biosynthesis of cholesterol, ubiquinone and haem A, and protein N-glycosylation and prenylation. b, hsp-6p::gfp animals treated with antimycin after pre-treatment with increasing concentration of simvastatin. c, hsp-6p::gfp animals with mock, 80 μg ml−1 simvastatin or 80 μg ml−1 mevastatin treatment. d, hsp-6p::gfp animals raised on control RNAi, hmgs-1(RNAi), or hmgs-1(RNAi) with the addition of geranylgeranyl pyrophosphate. The animals were then treated with antimycin to induce mitochondrial damage. Statin toxicity has been proposed to be caused by the inhibition of Rab prenylation24. Geranylgeranyl pyrophosphate (GGPP), a precursor of protein prenylation rescued the statin side effect in cell culture24. GGPP also partially rescued the deficiency of mitochondrial surveillance and activated hsp-6p::gfp in antimycin-treated hmgs-1(RNAi) animals.
Extended Data Figure 9 Mitochondrial immunostaining in HEK293T cells.
The cells were treated with DMSO, 10 μM simvastatin or 10 μM mevastatin for 2 days.
Supplementary information
Movement abilities of animals treated with control RNAi
Animals were filmed after 48 hours of growth on control RNAi. (MOV 1725 kb)
Movement abilities of animals treated with sptl-1(RNAi)
Animals were filmed after 48 hours of growth on sptl-1(RNAi) (MOV 2107 kb)
Rights and permissions
About this article
Cite this article
Liu, Y., Samuel, B., Breen, P. et al. Caenorhabditis elegans pathways that surveil and defend mitochondria. Nature 508, 406–410 (2014). https://doi.org/10.1038/nature13204
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13204
This article is cited by
-
Mitochondrial aconitase suppresses immunity by modulating oxaloacetate and the mitochondrial unfolded protein response
Nature Communications (2023)
-
A novel N6-Deoxyadenine methyltransferase METL-9 modulates C. elegans immunity via dichotomous mechanisms
Cell Research (2023)
-
C. elegans monitor energy status via the AMPK pathway to trigger innate immune responses against bacterial pathogens
Communications Biology (2022)
-
Transcriptional response of Meloidogyne incognita to non-fumigant nematicides
Scientific Reports (2022)
-
Neuroligin-mediated neurodevelopmental defects are induced by mitochondrial dysfunction and prevented by lutein in C. elegans
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.