Abstract
Bacterial type IV secretion systems translocate virulence factors into eukaryotic cells1,2, distribute genetic material between bacteria and have shown potential as a tool for the genetic modification of human cells3. Given the complex choreography of the substrate through the secretion apparatus4, the molecular mechanism of the type IV secretion system has proved difficult to dissect in the absence of structural data for the entire machinery. Here we use electron microscopy to reconstruct the type IV secretion system encoded by the Escherichia coli R388 conjugative plasmid. We show that eight proteins assemble in an intricate stoichiometric relationship to form an approximately 3 megadalton nanomachine that spans the entire cell envelope. The structure comprises an outer membrane-associated core complex1 connected by a central stalk to a substantial inner membrane complex that is dominated by a battery of 12 VirB4 ATPase subunits organized as side-by-side hexameric barrels. Our results show a secretion system with markedly different architecture, and consequently mechanism, to other known bacterial secretion systems1,4,5,6.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fronzes, R., Christie, P. J. & Waksman, G. The structural biology of type IV secretion systems. Nature Rev. Microbiol. 7, 703–714 (2009)
Schröder, G. & Lanka, E. The mating pair formation system of conjugative plasmids—a versatile secretion machinery for transfer of proteins and DNA. Plasmid 54, 1–25 (2005)
Schröder, G., Schuelein, R., Quebatte, M. & Dehio, C. Conjugative DNA transfer into human cells by the VirB/VirD4 type IV secretion system of the bacterial pathogen Bartonella henselae. Proc. Natl Acad. Sci. USA 108, 14643–14648 (2011)
Cascales, E. & Christie, P. J. Definition of a bacterial type IV secretion pathway for a DNA substrate. Science 304, 1170–1173 (2004)
Johnson, T. L., Abendroth, J., Hol, W. G. & Sandkvist, M. Type II secretion: from structure to function. FEMS Microbiol. Lett. 255, 175–186 (2006)
Marlovits, T. C. et al. Structural insights into the assembly of the type III secretion needle complex. Science 306, 1040–1042 (2004)
Fronzes, R. et al. Structure of a type IV secretion system core complex. Science 323, 266–268 (2009)
Kerr, J. E. & Christie, P. J. Evidence for VirB4-mediated dislocation of membrane-integrated VirB2 pilin during biogenesis of the Agrobacterium VirB/VirD4 type IV secretion system. J. Bacteriol. 192, 4923–4934 (2010)
Rivera-Calzada, A. et al. Structure of a bacterial type IV secretion core complex at subnanometre resolution. EMBO J. 32, 1195–1204 (2013)
Wallden, K. et al. Structure of the VirB4 ATPase, alone and bound to the core complex of a type IV secretion system. Proc. Natl Acad. Sci. USA 109, 11348–11353 (2012)
Passmore, L. A. et al. Structural analysis of the anaphase-promoting complex reveals multiple active sites and insights into polyubiquitylation. Mol. Cell 20, 855–866 (2005)
Chandran, V. et al. Structure of the outer membrane complex of a type IV secretion system. Nature 462, 1011–1015 (2009)
Peña, A. et al. The hexameric structure of a conjugative VirB4 protein ATPase provides new insights for a functional and phylogenetic relationship with DNA translocases. J. Biol. Chem. 287, 29925–29932 (2012)
Durand, E., Oomen, C. & Waksman, G. Biochemical dissection of the ATPase TraB, the VirB4 homologue of the Escherichia coli pKM101 conjugation machinery. J. Bacteriol. 192, 2315–2323 (2010)
Locht, C., Coutte, L. & Mielcarek, N. The ins and outs of pertussis toxin. FEBS J. 278, 4668–4682 (2011)
Pantoja, M., Chen, L. S., Chen, Y. C. & Nester, E. W. Agrobacterium type IV secretion is a two-step process in which export substrates associate with the virulence protein VirJ in the periplasm. Mol. Microbiol. 45, 1325–1335 (2002)
Ripoll-Rozada, J., Zunzunegui, S., de la Cruz, F., Arechaga, I. & Cabezon, E. Functional interactions of VirB11 traffic ATPases with VirB4 and VirD4 molecular motors in type IV secretion systems. J. Bacteriol. 195, 4195–4201 (2013)
Schraidt, O. & Marlovits, T. C. Three-dimensional model of Salmonella’s needle complex at subnanometer resolution. Science 331, 1192–1195 (2011)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: Semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Heymann, J. B. Bsoft: Image and molecular processing in electron microscopy. J. Struct. Biol. 133, 156–169 (2001)
van Heel, M. et al. Single-particle electron cryo-microscopy: towards atomic resolution. Q. Rev. Biophys. 33, 307–369 (2000)
vanHeel, M., Harauz, G., Orlova, E. V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996)
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nature Methods 9, 671–675 (2012)
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
De Carlo, S., Boisset, N. & Hoenger, A. High-resolution single-particle 3D analysis on GroEL prepared by cryo-negative staining. Micron 39, 934–943 (2008)
Scheres, S. H. W., Nunez-Ramirez, R., Sorzano, C. O. S., Carazo, J. M. & Marabini, R. Image processing for electron microscopy single-particle analysis using XMIPP. Nature Protocols 3, 977–990 (2008)
Orlov, S. S. Theory of three-dimensional reconstruction. 1. Conditions of a complete set of projections. Sov. Phys. Crystallogr. 20, 312–314 (1975)
Kudryashev, M. et al. In situ structural analysis of the Yersinia enterocolitica injectisome. eLife 2, e00792 (2013)
Acknowledgements
This work was financed by grant 098302 from the Wellcome Trust to G.W. and by equipment grant 079605 from the Wellcome Trust. R.F. and A.D. were financed by Institut Pasteur and the CNRS. F.G. was the recipient of ‘Bourse Roux’ from Institut Pasteur. We thank G. Péhau-Arnaudet for support with the electron microscopes at Institut Pasteur.
Author information
Authors and Affiliations
Contributions
H.H.L., F.G., R.F., E.V.O. and G.W. designed the experiments. H.H.L., F.G., S.C. and F.L. generated the clones. H.H.L. first purified the T4SS3–10 complex, and collected and processed the negative-stain electron microscope data for that complex. F.G., assisted by A.D., purified the T4SS3–10 complex and performed the stoichiometry, gold labelling and membrane wash experiments for that complex. A.R.-C. purified, collected electron microscope data and processed the TrwH/TrwF/TrwE complex. N.B., assisted by A.R., collected T4SS3–10 complex cryo-negative-stain data. H.H.L. and A.R.-C. purified, collected electron microscope data and processed TrwK and MBP–TrwK. A.R.-C. and F.L. purified the TrwM/TrwK complex and determined its stoichiometry. H.H.L. drafted the paper, and H.H.L., F.G., R.F., E.V.O. and GW finalized it.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Conserved gene architecture between E. coli R388 plasmid and A. tumefaciens T4S systems. Confirmation of T4SS3–10 complex constituents and the anomalous migration of VirB6/TrwI by SDS–PAGE.
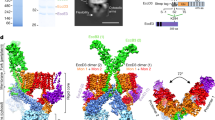
a, In this study, R388 virB1/trwN to virB10/trwE were overexpressed for purification of the T4SS3–10 complex using a StrepII tag located at the VirB10/TrwE C terminus. The figure was adapted from ref. 2. b, Left, western blot of specific individually cloned His6- or Flag-tagged gene variants in the T4SS1–10 clone. Anti-His6 or anti-Flag antibody was used as required. Note that western blot against Flag-tagged VirB2/TrwL in the purified T4SS3–10 complex gel gave no signal. Also, extensive LC–MS of trypsin-digested T4SS3–10 complex where VirB2 was untagged yielded no trace of VirB2/TrwL peptides. Thus, VirB2/TrwL could not be detected. VirB2/TrwL undergoes processing from a propilin to an assembly-competent pilin by a poorly understood mechanism that is reliant on signal peptidases and other poorly defined host encoded enzymes. Overexpression of such processing enzymes may therefore be required for overexpression of VirB2/TrwL and its incorporation into the T4SS3–10 complex. However, another possibility for the failure to observe VirB2 is that VirB2 might not tolerate insertion of tags in its sequence. Note that a fully assembled T4SS3–10 complex was observed by negative-stain electron microscopy for all His6- or Flag-tagged variants investigated. Right, VirB6/TrwI purified in isolation as an N-terminal MBP fusion protein. Cleavage of MBP confirms that VirB6/TrwI migrates anomalously by SDS–PAGE with a mass of approximately 27 kDa rather than at 35.1 kDa as calculated from its sequence. This seems not to be a proteolysis product as western blot of C-terminal Flag-tagged VirB6/TrwI shows a similar migration pattern (left panel).
Extended Data Figure 2 VirB10/TrwE, VirB9/TrwF and VirB7/TrwH form a pKM101-like core complex with C14 symmetry.
a, VirB10/TrwE, VirB9/TrwF and VirB7/TrwH (the R388 core complex) co-elute as a single peak during gel filtration (1, column exclusion limit; 2, 665 kDa; 3, 75 kDa). b, SDS–PAGE comparison of pKM101 and R388 core complexes. VirB9/TrwF and VirB10/TrwE were confirmed by N-terminal sequencing. c, Typical negative-stain electron microscopy micrograph of R388 core complexes. d, Comparison of the R388 and pKM101 core complexes. Top and side view class averages for the two complexes (left panel). The R388 core complex shows clear C14 symmetry as shown by the rotational auto-correlation function plot (right panel).
Extended Data Figure 3 The T4SS3–10 complex shows significant flexibility between the core complex and IMC.
a, A gallery of pairs of typical T4SS3–10 complex class averages chosen to show the IMC in 15° rotational increments around the particle long axis. Note the different tilts of the core complex in relation to the IMC. b, Variance analysis image to highlight observed flexibility between the core complex and IMC. The image shows the variance between 15 ‘front view’ T4SS3–10 complex class averages aligned using only their IMC region (see Methods for further detail). c, Angular distribution plot for the position of the core complex relative to the IMC. The plot consists of the same 15 ‘front view’ T4SS3–10 complex class averages as introduced in b (see Methods for further detail). d, Plots showing the assigned angular distribution of constituent classes around the Euler sphere for both the core complex/stalk and IMC reconstructions. Note that angular reconstitution was used for Euler angle assignment of the core complex/stalk, whereas projection matching was used for the IMC. The equatorial band distribution of angles reflects the fact that elongated molecules such as the T4SS3–10 are mostly represented by side views. Such an even equatorial band distribution provides the basis for a reliable reconstruction as demonstrated in ref. 29.
Extended Data Figure 4 Fourier shell correlation and depiction of separate core complex and IMC reconstructions.
a, Fourier shell correlation curves of the final models corresponding to the core complex (left) and IMC (right). Resolution according to the Fourier shell correlation = 0.5 criterion corresponds to approximately 18 and 23 Å, respectively. b, From top row to bottom row: representative class averages, respective variance images, corresponding re-projections and corresponding surface views of the final map for the core complex reconstruction. c, Side and top views of the core complex reconstruction. d, As described in b, but for the IMC reconstruction. e, Side and top views of the IMC reconstruction.
Extended Data Figure 5 Schematic representation of components in selected regions of the T4SS3–10 complex.
a, Top view representation of the stalk and arches. In the asymmetric T4SS3–10 complex reconstruction, four linkers are observed in the stalk. However, given the flexibility in this region, other unresolved linkers may exist. The arches are attached to the IMC barrels with up to six thin interconnecting flexible linkers (orange numerals, 1–6). The observed number of linkers varied during processing although a dominant triad was always observed in positions 1, 3 and 5. b, Surface representation and schematic (middle) of a single IMC barrel segment. Given the trimer of dimer arrangement of densities within the middle tier of each IMC barrel, each barrel segment (Fig. 3e) must consist of two subunits. The barrel densities comprise VirB4/TrwK as described in Fig. 4.
Extended Data Figure 6 Verification of the T4SS3–10 structure by cryo-negative staining.
a, Three image tilt series of cryo-negatively stained T4SS3–10 complexes. Scale bar, 100 nm. b, Tracking of specific individual particles (red or blue circle in a) at different tilt angles. Inset schematic diagrams aid interpretation of either side views (single barrel) or front views (two barrels). Note that the blue encircled particle has been rotated by about 180° in the image plane to show the particle with the core complex at the top. c, Representative characteristic class averages of cryo-negatively stained T4SS3–10 complexes (from left to right: side, oblique and front views). d, Equivalent of c, but using negative stain.
Extended Data Figure 7 Class averages derived from a test data set of the T4SS3–10/His6-B6 complex; biochemical and electron microscopy studies of VirB4/TrwK, and maltose binding protein (MBP) fused with VirB4/TrwK.
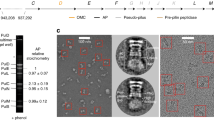
a, Comparison of class averages of the T4SS3–10/His6-B6 complex against those of the T4SS3–10 complex show them to be similar. Views presented here are similar to those shown in Extended Data Fig. 3a at 0, 45 and 90°. b, Gel filtration profile showing the co-elution of a VirB4/TrwK and VirB3/TrwM complex (1, column exclusion limit; 2, 665 kDa; 3, 75 kDa). The SDS–PAGE of this complex is shown in Fig. 4B. c, Left, SDS–PAGE of VirB4/TrwK purified from the membrane fraction. Right, in the presence of LDAO detergent, VirB4/TrwK exists as a monomer as shown by size exclusion chromatography. d, Negative-stain electron microscopy image of VirB4/TrwK as purified in c. Class averages (inset) show monomeric VirB4/TrwK to form an elongated protein with general dimensions of approximately 105 Å × 40 Å. Inset scale bar, 5 nm. e, Left, SDS–PAGE of N-terminal MBP–VirB4/TrwK fusion purified in the absence of detergents from the cytoplasmic fraction. Right, the negative-stain electron microscopy image of MBP–VirB4/TrwK fusion protein is similar to that of monomeric VirB4/TrwK as shown in d. However, an extra MBP density is clearly observable in class averages (inset) and indicates that the VirB4/TrwK N terminus is located in the centre of its long axis. The dimensions of the VirB4/TrwK monomer are compatible with the fitting of six VirB4/TrwK monomers within each IMC barrel. f, Western blot against the membrane fraction of a modified T4SS3–10/B4-His6 complex purification using anti-His6 antibody. In the absence of detergent, the membrane fraction was washed iteratively twice with 6 M urea, then once with 0.1 M sodium carbonate pH 11, and finally once with 2 M KCl. Substantial VirB4/TrwK signal remained associated with the membrane fraction after all four stringent washes.
Extended Data Figure 8 Gold labelling of TrwI/VirB6 and TrwK/VirB4.
T4SS3–10, T4SS3–10/B4-His6 and T4SS3–10/His6-B6 complexes were purified and applied to carbon-coated grids. His6 tags were specifically labelled by Ni-NTA nanogold beads (5 nm diameter) directly on the grid before negative staining with 2% uranyl acetate. a, Overview and gallery of 12 representative gold-labelled particles of T4SS3–10/B4-His6. b, As a, but for T4SS3–10/His6-B6. c, Overview of the Ni-NTA nanogold bead labelling control experiment using the purified T4SS3–10 complex. No gold-labelling was observed. The scale bar in all overview panels is 50 nm.
Extended Data Figure 9 The structure of the T4SS3–10 complex is compatible with a general two-step mechanism for T4S systems.
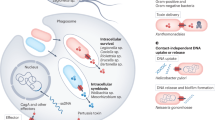
a, In earlier studies in which the pKM101 core complex (VirB7/TraN, VirB9/TraO and VirB10/TraF) was purified in vitro in isolation7,10, the flexible N terminus of VirB10/TraF is in an alternate ‘compacted’ conformation (left) that is positioned immediately under the I-layer to form the core complex ‘base’. When the core complex is either in situ and is constrained by the inner and outer membranes, or is constrained by other components of the secretion system as observed in vitro in the T4SS3–10 complex, the VirB10/TrwE N terminus is in an ‘extended’ conformation (right). The absolute arrangement of the VirB10/TrwE N terminus within the core complex when extended is yet to be determined although the 14 strands may form a rotationally discontinuous punctuated collar that contributes to, or surrounds, the stalk. It should also be noted that we cannot exclude the possibility that, in the membrane environment, the core and inner membrane complexes might be in closer proximity, possibly resulting in the stalk fitting up into the core complex chamber and in VirB10 adopting a less stretched conformation. See b for key to colour scheme. b, Schematic of the T4SS3–10 complex showing three possible entry routes for substrate across the inner membrane and into the core complex inner chamber. In R388 conjugal transfer, the coupling protein VirD4/TrwB probably pumps DNA substrate into the periplasm (route 2), although passage by VirB4/TrwK may currently not be discounted (route 1). Other T4S systems such as that from Bordetella use the Sec pathway to transfer effectors across the inner membrane (route 3). For clarity, only VirB nomenclature is used in the colour scheme.
Extended Data Figure 10 Comparison of the type III and type IV secretion systems.
In contrast to the T4S system, the type III secretion (T3S) system (Electron Microscopy Data Bank accession number EMD-1875) has an obvious continuous secretion pore that spans the entire cell envelope, including inner and outer membranes. In addition, the distance between putatively assigned inner and outer membrane locations is slightly extended for the T3S system relative to the T4S system, although this could simply be due to the different systems being derived from different species: Salmonella and Escherichia, respectively. In situ, the T3S system is known to vary in length within the intermembrane space, from 30 to 36 nm (ref. 30).
Rights and permissions
About this article
Cite this article
Low, H., Gubellini, F., Rivera-Calzada, A. et al. Structure of a type IV secretion system. Nature 508, 550–553 (2014). https://doi.org/10.1038/nature13081
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13081
This article is cited by
-
Structural and functional diversity of type IV secretion systems
Nature Reviews Microbiology (2023)
-
Comparative genome analysis unravels pathogenicity of Xanthomonas albilineans causing sugarcane leaf scald disease
BMC Genomics (2022)
-
Structure of a type IV secretion system core complex encoded by multi-drug resistance F plasmids
Nature Communications (2022)
-
Possible drugs for the treatment of bacterial infections in the future: anti-virulence drugs
The Journal of Antibiotics (2021)
-
Architecture of the outer-membrane core complex from a conjugative type IV secretion system
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.