Abstract
In bacterial cells, processing of double-stranded DNA breaks for repair by homologous recombination is dependent upon the recombination hotspot sequence χ (Chi)1,2 and is catalysed by either an AddAB- or RecBCD-type helicase–nuclease (reviewed in refs 3, 4). These enzyme complexes unwind and digest the DNA duplex from the broken end until they encounter a χ sequence5, whereupon they produce a 3′ single-stranded DNA tail onto which they initiate loading of the RecA protein6. Consequently, regulation of the AddAB/RecBCD complex by χ is a key control point in DNA repair and other processes involving genetic recombination. Here we report crystal structures of Bacillus subtilis AddAB in complex with different χ-containing DNA substrates either with or without a non-hydrolysable ATP analogue. Comparison of these structures suggests a mechanism for DNA translocation and unwinding, suggests how the enzyme binds specifically to χ sequences, and explains how χ recognition leads to the arrest of AddAB (and RecBCD) translocation that is observed in single-molecule experiments7,8,9.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lam, S. T., Stahl, M. M., McMilin, K. D. & Stahl, F. W. Rec-mediated recombinational hot spot activity in bacteriophage lambda. II. A mutation which causes hot spot activity. Genetics 77, 425–433 (1974)
Chédin, F., Noirot, P., Biaudet, V. & Ehrlich, S. D. A five-nucleotide sequence protects DNA from exonucleolytic degradation by AddAB, the RecBCD analogue of Bacillus subtilis. Mol. Microbiol. 29, 1369–1377 (1998)
Dillingham, M. S. & Kowalczykowski, S. C. RecBCD enzyme and the repair of double-stranded DNA breaks. Microbiol. Mol. Biol. Rev. 72, 642–671 (2008)
Wigley, D. B. Bacterial DNA repair: recent insights into the mechanism of RecBCD, AddAB and AdnAB. Nature Rev. Microbiol. 11, 9–13 (2013)
Ponticelli, A. S., Schultz, D. W., Taylor, A. F. & Smith, G. R. Chi-dependent DNA strand cleavage by RecBC enzyme. Cell 41, 145–151 (1985)
Anderson, D. G. & Kowalczykowski, S. C. The translocating RecBCD enzyme stimulates recombination by directing RecA protein onto ssDNA in a chi-regulated manner. Cell 90, 77–86 (1997)
Spies, M. et al. A molecular throttle: the recombination hotspot chi controls DNA translocation by the RecBCD helicase. Cell 114, 647–654 (2003)
Spies, M., Amitani, I., Baskin, R. J. & Kowalczykowski, S. C. RecBCD enzyme switches lead motor subunits in response to chi recognition. Cell 131, 694–705 (2007)
Carrasco, C., Gilhooly, N. S., Dillingham, M. S. & Moreno-Herrero, F. On the mechanism of recombination hotspot scanning during double-stranded DNA break resection. Proc. Natl Acad. Sci. USA 110, E2562–E2571 (2013)
Saikrishnan, K. et al. Insights into Chi recognition from the structure of an AddAB-type helicase-nuclease complex. EMBO J. 31, 1568–1578 (2012)
Singleton, M. R., Dillingham, M. S., Gaudier, M., Kowalczykowski, S. C. & Wigley, D. B. Crystal structure of RecBCD enzyme reveals a machine for processing DNA breaks. Nature 432, 187–193 (2004)
Yeeles, J. T., Gwynn, E. J., Webb, M. R. & Dillingham, M. S. The AddAB helicase-nuclease catalyses rapid and processive DNA unwinding using a single Superfamily 1A motor domain. Nucleic Acids Res. 39, 2271–2285 (2011)
Singleton, M. R., Dillingham, M. S. & Wigley, D. B. Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 76, 23–50 (2007)
Unciuleac, M. C. & Shuman, S. Characterization of the mycobacterial AdnAB DNA motor provides insights into the evolution of bacterial motor-nuclease machines. J. Biol. Chem. 285, 2632–2641 (2010)
Velankar, S. S., Soultanas, P., Dillingham, M. S., Subramanya, H. S. & Wigley, D. B. Crystal structures of complexes of PcrA DNA helicase with a DNA substrate indicate an inchworm mechanism. Cell 97, 75–84 (1999)
Wu, C. G., Bradford, C. & Lohman, T. M. Escherichia coli RecBC helicase has two translocase activities controlled by a single ATPase motor. Nature Struct. Mol. Biol. 17, 1210–1217 (2010)
Handa, N. et al. Molecular determinants responsible for recognition of the single-stranded DNA regulatory sequence, χ, by RecBCD enzyme. Proc. Natl Acad. Sci. USA 109, 8901–8906 (2012)
Yang, L. et al. Mutation of χ-recognition by RecBCD reveals a regulated molecular latch and suggests a channel-bypass mechanism for biological control. Proc. Natl Acad. Sci. USA 109, 8907–8912 (2012)
Kovall, R. & Matthews, B. W. Toroidal structure of λ-exonuclease. Science 277, 1824–1827 (1997)
Zhang, J., McCabe, K. A. & Bell, C. E. Crystal structures of λ exonuclease in complex with DNA suggest an electrostatic ratchet mechanism for processivity. Proc. Natl Acad. Sci. USA 108, 11872–11877 (2011)
Taylor, A. F. & Smith, G. R. RecBCD enzyme is a DNA helicase with fast and slow motors of opposite polarity. Nature 423, 889–893 (2003)
Thaler, D. S. et al. Recombination of bacteriophage λ in recD mutants of Escherichia coli. Genome 31, 53–67 (1989)
Dohoney, K. M. & Gelles, J. χ-sequence recognition and DNA translocation by single RecBCD helicase/nuclease molecules. Nature 409, 370–374 (2001)
Handa, N., Bianco, P. R., Baskin, R. J. & Kowalczykowski, S. C. Direct visualization of RecBCD movement reveals cotranslocation of the RecD motor after χ recognition. Mol. Cell 17, 745–750 (2005)
Wong, C. J., Rice, R. L., Baker, N. A., Ju, T. & Lohman, T. M. Probing 3′-ssDNA loop formation in E. coli RecBCD/RecBC-DNA complexes using non-natural DNA: a model for “Chi” recognition complexes. J. Mol. Biol. 362, 26–43 (2006)
Anderson, D. G., Churchill, J. J. & Kowalczykowski, S. C. A single mutation, RecBD1080A eliminates RecA protein loading but not Chi recognition by RecBCD enzyme. J. Biol. Chem. 274, 27139–27144 (1999)
Yeeles, J. T., Cammack, R. & Dillingham, M. S. An iron-sulfur cluster is essential for the binding of broken DNA by AddAB-type helicase-nucleases. J. Biol. Chem. 284, 7746–7755 (2009)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Acknowledgements
We thank the European Synchrotron Radiation Facility and Diamond synchrotrons for access to beamlines. The work was funded by the Royal Society, the Wellcome Trust and the European Research Council (M.S.D.), Cancer Research UK (D.B.W.) and EMBO (W.W.K.).
Author information
Authors and Affiliations
Contributions
W.W.K. and D.B.W. designed the experiments. W.W.K., X.F., M.W. and N.B.C. performed the experiments. W.W.K., M.W., M.S.D. and D.B.W. analysed the data and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 DNA substrates used in the structure determinations.
We have previously determined the crystal structure of an initiation complex of AddAB using the DNA substrate shown at the top10. The substrates used in the current study differ by the additions shown in blue below. Five T residues were added to make a 5′ tail for the fork. The 3′ tail was extended by the addition of an AddAB χ sequence (AGCGG), preceded by a ‘spacer’ sequence of between 1 and 7 T residues to extend the distance between the fork junction and the χ sequence along the 3′ tail to mimic the products of sequential unwinding of a DNA fork. Oligo 023 used in this study contained a six-base T spacer. Oligo 027 contained a seven-T spacer plus an additional four T residues beyond the χ sequence as shown above. AddAB failed to recognize χ in substrates with six or less residues in the spacer but was able to bind to χ with seven T residues in the spacer region.
Extended Data Figure 2 Cartoon representation of the DNA-binding site in the two complexes.
Residues from AddA in orange, AddB in cyan, hydrogen bonds and charged interactions as arrows, hydrophobic and stacked interactions as solid lines.
Extended Data Figure 3 Nucleotide-binding sites.
Difference electron density (Fo – Fc, contoured at 2.5σ) for bound nucleotide at the ATP-binding sites in the AddA and AddB subunits in the ADPNP and χ complexes.
Extended Data Figure 4 Conformational variability of domain 2A of AddA in different χ-bound structures.
The two extreme variants are shown for illustration. Although domain 1A (green and light green) of the AddA protein (orange) remains in an essentially constant position, the position of the 2A domain (red and light red) varies from one that is almost superimposable with the ‘apo’ complex that lacks ADPNP, to one that is midway between the apo and ADPNP-bound conformations plus a variety of conformations between these two extreme cases. The extreme cases differ by a rotation of 5° compared to 7° for the difference between the ADPNP and apo structures. By contrast, the conformation of the 2A domain of AddA adopts an almost invariable conformation in different structures of the AddAB–DNA complex grown with ADPNP with a variety of DNA substrates in which χ is not bound.
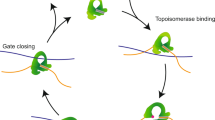
Extended Data Figure 5 Cartoon summarizing our current knowledge of the molecular structures of different functional states along the AddAB reaction pathway.
We now have crystal structures for the first three functional states as shown in the figure. These show how the protein complex interacts with DNA (initiation complex), what happens when ATP binds and how this leads to DNA unwinding (translocation complex) and what happens when this complex encounters a χ sequence (χ recognition complex). At least two further states have yet to be visualized: (1) what happens after the pause at χ (χ reactivation complex) and (2) how the protein interacts with and loads RecA (RecA-loading complex).
Supplementary information
DNA translocation mechanism
This video depicts a morph between the binary and ternary complexes. The details are described in the main text. Note that this is likely a simplification for the mechanism which assumes just two states and additional states on the unwinding pathway may well exist. The video shows two cycles of ATP binding and hydrolysis. (MOV 3591 kb)
Translocation mechanism emphasising the role of the motor domains of AddA
Similar to Supplementary video 1, although this video emphasises the role of the AddA domains in the unwinding mechanism. The 1A domain is shown in green, 2A in red and 1B in blue. The remainder of the protein is shown translucent orange (AddA) or cyan (AddB) for context. (MOV 3554 kb)
Rights and permissions
About this article
Cite this article
Krajewski, W., Fu, X., Wilkinson, M. et al. Structural basis for translocation by AddAB helicase–nuclease and its arrest at χ sites. Nature 508, 416–419 (2014). https://doi.org/10.1038/nature13037
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13037
This article is cited by
-
Modeling DNA Unwinding by AddAB Helicase–Nuclease and Modulation by Chi Sequences: Comparison with AdnAB and RecBCD
Cellular and Molecular Bioengineering (2019)
-
Phylogenomics of Cas4 family nucleases
BMC Evolutionary Biology (2017)
-
Bacteriophage T5 gene D10 encodes a branch-migration protein
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.