Abstract
Alveoli are gas-exchange sacs lined by squamous alveolar type (AT) 1 cells and cuboidal, surfactant-secreting AT2 cells. Classical studies suggested that AT1 arise from AT2 cells, but recent studies propose other sources. Here we use molecular markers, lineage tracing and clonal analysis to map alveolar progenitors throughout the mouse lifespan. We show that, during development, AT1 and AT2 cells arise directly from a bipotent progenitor, whereas after birth new AT1 cells derive from rare, self-renewing, long-lived, mature AT2 cells that produce slowly expanding clonal foci of alveolar renewal. This stem-cell function is broadly activated by AT1 injury, and AT2 self-renewal is selectively induced by EGFR (epidermal growth factor receptor) ligands in vitro and oncogenic Kras(G12D) in vivo, efficiently generating multifocal, clonal adenomas. Thus, there is a switch after birth, when AT2 cells function as stem cells that contribute to alveolar renewal, repair and cancer. We propose that local signals regulate AT2 stem-cell activity: a signal transduced by EGFR-KRAS controls self-renewal and is hijacked during oncogenesis, whereas another signal controls reprogramming to AT1 fate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Accessions
Gene Expression Omnibus
Data deposits
Microarray datasets were deposited at Gene Expression Omnibus (accession code GSE49346) and GEXC (https://gexc.stanford.edu/population/detail/998 and https://gexc.stanford.edu/population/detail/999).
References
Bertalanffy, F. D. & Leblond, C. P. Structure of respiratory tissue. Lancet 266, 1365–1368 (1955)
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2013. CA Cancer J. Clin. 63, 11–30 (2013)
Rock, J. R. & Hogan, B. L. Epithelial progenitor cells in lung development, maintenance, repair, and disease. Annu. Rev. Cell Dev. Biol. 27, 493–512 (2011)
Chapman, H. A. et al. Integrin α6β4 identifies an adult distal lung epithelial population with regenerative potential in mice. J. Clin. Invest. 121, 2855–2862 (2011)
Adamson, I. Y. & Bowden, D. H. Derivation of type 1 epithelium from type 2 cells in the developing rat lung. Lab. Invest. 32, 736–745 (1975)
Spencer, H. & Shorter, R. G. Cell turnover in pulmonary tissues. Nature 194, 880 (1962)
Evans, M. J. & Bils, R. F. Identification of cells labeled with tritiated thymidine in the pulmonary alveolar walls of the mouse. Am. Rev. Respir. Dis. 100, 372–378 (1969)
Rock, J. R. et al. Multiple stromal populations contribute to pulmonary fibrosis without evidence for epithelial to mesenchymal transition. Proc. Natl Acad. Sci. USA 108, E1475–E1483 (2011)
Barkauskas, C. E. et al. Type 2 alveolar cells are stem cells in adult lung. J. Clin. Invest. 123, 3025–3036 (2013)
Evans, M. J., Cabral, L. J., Stephens, R. J. & Freeman, G. Renewal of alveolar epithelium in the rat following exposure to NO2 . Am. J. Pathol. 70, 175–198 (1973)
Adamson, I. Y. & Bowden, D. H. The type 2 cell as progenitor of alveolar epithelial regeneration. A cytodynamic study in mice after exposure to oxygen. Lab. Invest. 30, 35–42 (1974)
Kim, C. F. et al. Identification of bronchioalveolar stem cells in normal lung and lung cancer. Cell 121, 823–835 (2005)
McQualter, J. L., Yuen, K., Williams, B. & Bertoncello, I. Evidence of an epithelial stem/progenitor cell hierarchy in the adult mouse lung. Proc. Natl Acad. Sci. USA 107, 1414–1419 (2010)
Kumar, P. A. et al. Distal airway stem cells yield alveoli in vitro and during lung regeneration following H1N1 influenza infection. Cell 147, 525–538 (2011)
Kajstura, J. et al. Evidence for human lung stem cells. N. Engl. J. Med. 364, 1795–1806 (2011)
Rawlins, E. L. et al. The role of Scgb1a1+ Clara cells in the long-term maintenance and repair of lung airway, but not alveolar, epithelium. Cell Stem Cell 4, 525–534 (2009)
Van Keymeulen, A. et al. Distinct stem cells contribute to mammary gland development and maintenance. Nature 479, 189–193 (2011)
Burri, P. H. & Moschopulos, M. Structural analysis of fetal rat lung development. Anat. Rec. 234, 399–418 (1992)
Buckingham, S., McNary, W. F., Jr & Sommers, S. C. Pulmonary alveolar cell inclusions: their development in the rat. Science 145, 1192–1193 (1964)
Williams, M. C. & Dobbs, L. G. Expression of cell-specific markers for alveolar epithelium in fetal rat lung. Am. J. Respir. Cell Mol. Biol. 2, 533–542 (1990)
Hughes, G. M. Ultrastructure of the lung of Neoceratodus and Lepidosiren in relation to the lung of other vertebrates. Folia Morphol. (Praha) 21, 155–161 (1973)
Miller, L. A., Wert, S. E. & Whitsett, J. A. Immunolocalization of sonic hedgehog (Shh) in developing mouse lung. J. Histochem. Cytochem. 49, 1593–1603 (2001)
Messier, B. & Leblond, C. P. Cell proliferation and migration as revealed by radioautography after injection of thymidine-H3 into male rats and mice. Am. J. Anat. 106, 247–285 (1960)
Bowden, D. H., Adamson, I. Y. & Wyatt, J. P. Reaction of the lung cells to a high concentration of oxygen. Arch. Pathol. 86, 671–675 (1968)
Herbst, R. S., Heymach, J. V. & Lippman, S. M. Lung cancer. N. Engl. J. Med. 359, 1367–1380 (2008)
Sutherland, K. D. & Berns, A. Cell of origin of lung cancer. Mol. Oncol. 4, 397–403 (2010)
Xu, X. et al. Evidence for type II cells as cells of origin of K-Ras-induced distal lung adenocarcinoma. Proc. Natl Acad. Sci. USA 109, 4910–4915 (2012)
Lin, C. et al. Alveolar type II cells possess the capability of initiating lung tumor development. PLoS ONE 7, e53817 (2012)
Nikitin, A. Y. et al. Classification of proliferative pulmonary lesions of the mouse: recommendations of the mouse models of human cancers consortium. Cancer Res. 64, 2307–2316 (2004)
Takanami, I., Takeuchi, K. & Kodaira, S. Tumor-associated macrophage infiltration in pulmonary adenocarcinoma: association with angiogenesis and poor prognosis. Oncology 57, 138–142 (1999)
Brody, J. S., Burki, R. & Kaplan, N. Deoxyribonucleic acid synthesis in lung cells during compensatory lung growth after pneumonectomy. Am. Rev. Respir. Dis. 117, 307–316 (1978)
Ding, B. S. et al. Endothelial-derived angiocrine signals induce and sustain regenerative lung alveolarization. Cell 147, 539–553 (2011)
Kretzschmar, K. & Watt, F. M. Lineage tracing. Cell 148, 33–45 (2012)
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001)
Harfe, B. D. et al. Evidence for an expansion-based temporal Shh gradient in specifying vertebrate digit identities. Cell 118, 517–528 (2004)
Clausen, B. E., Burkhardt, C., Reith, W., Renkawitz, R. & Forster, I. Conditional gene targeting in macrophages and granulocytes using LysMcre mice. Transgenic Res. 8, 265–277 (1999)
Ventura, A. et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007)
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001)
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007)
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010)
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nature Neurosci. 13, 133–140 (2010)
Rinkevich, Y., Lindau, P., Ueno, H., Longaker, M. T. & Weissman, I. L. Germ-layer and lineage-restricted stem/progenitors regenerate the mouse digit tip. Nature 476, 409–413 (2011)
Faust, N., Varas, F., Kelly, L. M., Heck, S. & Graf, T. Insertion of enhanced green fluorescent protein into the lysozyme gene creates mice with green fluorescent granulocytes and macrophages. Blood 96, 719–726 (2000)
Metzger, R. J., Klein, O. D., Martin, G. R. & Krasnow, M. A. The branching programme of mouse lung development. Nature 453, 745–750 (2008)
Messier, E. M., Mason, R. J. & Kosmider, B. Efficient and rapid isolation and purification of mouse alveolar type II epithelial cells. Exp. Lung Res. 38, 363–373 (2012)
Fujino, N. et al. A novel method for isolating individual cellular components from the adult human distal lung. Am. J. Respir. Cell Mol. Biol. 46, 422–430 (2012)
Dobbs, L. G., Williams, M. C. & Brandt, A. E. Changes in biochemical characteristics and pattern of lectin binding of alveolar type II cells with time in culture. Biochim. Biophys. Acta 846, 155–166 (1985)
Sugahara, K., Mason, R. J. & Shannon, J. M. Effects of soluble factors and extracellular matrix on DNA synthesis and surfactant gene expression in primary cultures of rat alveolar type II cells. Cell Tissue Res. 291, 295–303 (1998)
Seita, J. et al. Gene Expression Commons: an open platform for absolute gene expression profiling. PLoS ONE 7, e40321 (2012)
Giangreco, A. et al. Stem cells are dispensable for lung homeostasis but restore airways after injury. Proc. Natl Acad. Sci. USA 106, 9286–9291 (2009)
Lynch, P. J. & Jaffe, C. C. Bronchial anatomy detail of alveoli and lung circulation. http://commons.wikimedia.org/wiki/File%3ABronchial_anatomy.jpg (2006)
Burri, P. H. in Comprehensive Physiology 1–46 http://dx.doi.org/10.1002/cphy.cp030101 (Wiley, 2011)
Acknowledgements
We thank A. Andalon for technical assistance; H. Chapman (SftpC–Cre–ER-rtTA), B. Hogan (CCSP–Cre–ER), H. Ueno and I. Weissman (Rainbow), H. Clevers (Confetti), L. Luo (mTmG), and J. Sage (KrasLSL-G12D) for strains; B. Stripp for goat anti-CCSP antibody; F. H. Espinoza for annotated gene lists; R. Metzger, H. Chapman, and members of the Krasnow laboratory for discussions and comments on the manuscript; and Maria Petersen for help preparing figures and the manuscript.
Author information
Authors and Affiliations
Contributions
T.J.D. conducted the experiments except the gene expression profiling and AT2 cell cultures, which were done by D.G.B.; T.J.D., D.G.B. and M.A.K. conceived the experiments, analysed the data and wrote the manuscript. This work was supported by a Parker B. Francis Foundation Fellowship and NIH 5KO8HL084095 Award (T.J.D.), NIH T32HD007249 (D.G.B.), and an NHLBI U01HL099995 Progenitor Cell Biology Consortium grant (M.A.K.). M.A.K. is an investigator of the Howard Hughes Medical Institute.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Mature and developing structure of the lung and alveoli.
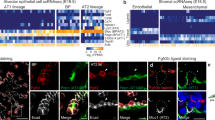
a, Mature lung showing close up (inset) of transition from bronchial tree to alveoli (from http://www.mayoclinic.com/, with permission). b, Alveoli are surrounded by a dense capillary network in which de-oxygenated blood (blue) from pulmonary arteries is oxygenated (red) and then returned to the heart through pulmonary veins (modified from ref. 51). c, E16.5 Shh–Cre > mTmG mouse lung in which GFP is expressed throughout the epithelium. Lobes are labelled (RAcc, right accessory; RCr, right cranial; RMid, right middle; RCa, right caudal; L, left) and boxed region shows the tip of the accessory lobe that was used for developmental analyses. By E16.5, the bronchial tree has formed and 1 day later sacculation begins as flat (squamous) AT1 and cuboidal AT2 cells mature to generate a functional gas exchange interface. Saccules subsequently undergo subdivision (‘secondary septation’) into mature alveoli that provide an extensive gas-exchange surface. Scale bar, 1 mm; d, Schematic of cell morphogenesis during sacculation (sac) (from ref. 52). Progenitors form flat AT1 cells adjacent to capillaries and AT2 cells specialized to secrete surfactant. e, f, e′, f′, Schematics (e, f) and images (e′, f′) of E-cadherin-stained accessory lobe tips at E18.5 and postnatal day 2 (PN2) showing longitudinal (e) and frontal (f) views of maturing AT1 (1) and AT2 (2) cells. P, proximal; D, distal; red, cell junctions (jxn); yellow, apical surfaces. Note lack of AT1 cells distally in sacculating airway. Scale bar, 10 μm (e′, f′). g, Quantification of ultrastructural classification of cell types in sacculating airways in E18.3 lungs (see Fig. 1r–u). Values shown are the numbers of each progenitor and cell type observed with the indicated ultrastructural features. No cells (n = 36) had features of an AT2> AT1 intermediate (AT1/2) or mature AT2 cell. E, embryonic day; PN, postnatal day.
Extended Data Figure 2 Clonal analysis of alveolar progenitor cells and lineage marking and tracing alveolar type 2 (AT2) cells with LysM-Cre.
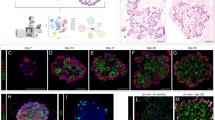
a, b, Shh–Cre–ER > mTmG embryos were induced in utero with a limiting dose (2 mg) of tamoxifen (tamox) at E15 to pulse-label epithelial cells at the distal lung tips (alveolar progenitors) with GFP (0.2 labelled cells per embryonic lung lobe) shortly before the onset of differentiation then examined 34 days later at PN30. a, An isolated clone (dashed circle) expressing the GFP lineage tag (green) in a PN30 lung. b, Close up of boxed region showing several flat AT1 cells (open arrows) and a cuboidal AT2 cell (filled arrows) within the alveolar clone, indicating that the tagged progenitor was bipotent. Tagged cells are interspersed with unrecombined cells (tdTomato, red). E, embryonic day; PN, postnatal day; Scale bar, 50 μm. c–f, Close-ups of alveoli of 1 (c, e) and 2 month old (d, f) LysM-Cre > mTmG lungs stained for the AT2 lineage tag (GFP, green) and the AT2 (c, d) or AT1 (e, f) markers indicated. Note that at 1 month lineage marked cells (green) express the AT2 (c) but not the AT1 marker (e). At 2 months (d, f), the lineage mark is also observed in some flat AT1 cells. Filled arrows, AT2 cells; open arrows, AT1 cells; E, embryonic day; PN, postnatal day. Scale bar, 20 μm (c–f).
Extended Data Figure 3 Proliferation analysis of bipotent progenitors and alveolar epithelial cells.
a–d, Late gestational (E17.5, a) and early postnatal (PN7, 14, 21; b–d) lungs stained for Nkx2.1 (green) for epithelial and Ki67 (red) for actively cycling cells. Note essentially exclusive labelling, indicating minimal proliferation of bipotent progenitors (a) or AT1 and AT2 cells (b–d). Arrow, a rare proliferating AT2 cell. Dashes outline distal epithelial tips; dotted line indicates mesothelium. E, embryonic day; PN, postnatal day; Scale bar, 35 μm.
Extended Data Figure 4 Quantification of cell type labelling and long-term lineage contribution of LysM-Cre and SftpC–Cre–ER marked cells.
a, Alveolar region of a PN 3 month (mo) Shh–Cre> R26EYFP mouse lung co-stained for GFP (green, epithelial cytoplasm) and LysM (red). Inset shows close-up of boxed region. LysM is detected in cytoplasm of many AT2 cells (filled arrow) but not AT1 cells (open arrow). b, Bronchoalveolar lung region of a PN 2 mo LysM reporter mouse expressing GFP from the endogenous locus stained for E-cadherin (red) to mark airway epithelium and GFP (green) to mark LysM-expressing cells. Note AT2 (filled arrows) but not bronchiolar cells (dashes mark bronchoalveolar junction (Badj)) express the LysM reporter. c, PN 17 mo LysM-Cre > mTmG lung stained for E-cadherin (red) and the AT2 lineage marker (green). Note many marked AT2 cells (filled arrows) but absence of lineage-marked cells in the terminal bronchiole (dashes, Badj). d, e, Lungs from LysM-EGFP (d) and LysM-Cre > mTmG (e) mice of the indicated ages stained for ciliated (acetylated tubulin, acTub, red) and neuroendocrine (NE) cell (CGRP, blue) markers and GFP (green) show no co-expressing cells, indicating ciliated (filled arrowhead) and NE (open arrowhead) cell types do not express LysM and do not derive from AT2 cells. Br, bronchus. f–h, Lungs from LysM-Cre > mTmG mice of the indicated ages stained for the AT2 lineage tag (green), the AT2 cell marker SftpC (red), and the Clara cell marker CCSP (blue) show CCSP+/SftpC+ (double-positive) cells (*) at the Badj, some of which are tagged. Marked double-positive cells are solitary (g) or in doublets (h). i, Lung from PN 1 mo SftpC–Cre–ER > mTmG mouse (administered 1 mg tamoxifen at PN19) analysed 13 days after pulse-labelling and stained as in d. Note no co-expressing cells, indicating that ciliated (filled arrowhead) and NE (open arrowhead) cell types are not tagged. j–l, Lungs from mice labelled as in panel i analysed 13 days (j) and 192 days (k, l) later, stained as in f. Note double-positive CCSP+/SftpC+ cells (*) at the Badj, some of which are pulse-labelled (j). After 192 day chase, marked double-positive cells are solitary (k) or in doublets (l). m, Quantification of lung cell types marked under different labelling and lineage trace conditions. The number of marked and total cells of each type scored (and Badj analysed for CCSP+/SftpC+ cells) is shown for each genotype and age analysed. For the SftpC–Cre–ER line, the dosage of tamoxifen (tamox) and interval time until analysis is also indicated. Because bronchial maintenance involves proliferation without significant cell dispersion50 (as we find for alveolar maintenance), the presence of marked CCSP+/SftpC+ cells primarily in isolation (>90%) at advanced ages suggests they did not contribute significantly to physiological bronchiolar renewal. d, days; PN, postnatal. Scale bar, 50 μm (a–l), 10 μm (insets in a).
Extended Data Figure 5 Lineage tracing alveolar type 2 (AT2) cells using SftpC–Cre–ER.
SftpC–Cre–ER > mTmG mice were administered 1 mg tamoxifen (Tamox) at PN18 then analysed later by staining for the lineage label (GFP, green) and AT2 cells (SftpC, red). a, b, 13 days later only AT2 cells are marked (a), whereas after 212 days flat AT1 cells also express the AT2 lineage tag (b). A peripheral renewal focus (mesothelium indicated by dotted line) involving multiple alveoli (asterisks) is shown, similar to results using LysM–Cre. Quantification revealed that 94% of AT2 cells were marked 13 days after tamoxifen induction and 97% after 192 days (see Extended Data Fig. 5m), indicating that the SftpC+ population is maintained by self-duplication, and that new AT2 cells do not derive from another cell population during physiological ageing. PN, postnatal day; bar, 100 μm.
Extended Data Figure 6 Alveolar type 2 (AT2) founder cell functional marker expression and self-duplication in vivo, and reprogramming into AT1 cells in vitro.
a–c, 16 month LysM-Cre > Confetti mouse lung stained for mCFP lineage tag (green), AT2 cell marker (Nkx2.1, red), and DAPI (blue) to identify clonal renewal foci. a, AT2 founder cell (filled arrow) associated with a daughter AT1 cell (open arrow) shown to express the A2L marker LAMP-1 (white), a protein associated with surfactant-containing lysosomes in mature, secretory AT2 cells. b, c, A clonal focus (b, boxed region) in which an AT2 cell has generated two additional AT2 cells (filled arrows) but no AT1 cells (c, close up of boxed region), demonstrating isolated self-duplication without AT1 cell reprogramming in vivo. Scale bars, 25 μm (a–c). d, Freshly isolated AT2 cells (Fig. 3f) cultured 4 days on glass with 10% serum adopt flat AT1 morphology (E-cadherin, green) and initiate AT1 marker expression (Aqp5, not shown). Scale bar, 10 μm.
Extended Data Figure 7 Clonogenic response to activated Kras using SftpC–Cre–ER and CCSP–Cre–ER.
a–c, Lungs of SftpC–Cre–ER (a) and CCSP–Cre–ER (b) mice carrying oncogenic KrasLSL-G12D/+ (Kras*) and Rainbow (Rbw), injected with 3 mg (a) or 1 mg (b) of tamoxifen (tamox) at indicated ages to induce Kras* expression and clonal lineage marking of StfpC- and CCSP-positive cells, respectively. Lungs were analysed after 17 days at PN43 (a) or after 39 days at PN81 (b). Note that multifocal tumours result when SftpC–Cre–ER mice are induced at 25 (a) days of age, whereas induction of Kras* in CCSP–Cre–ER mice at 42 days (b) shows many unresponsive cells and doublets throughout the bronchi (Br) as well as small clonal tumours located at bronchoalveolar duct junctions (Badj). c, An Sftpc–Cre–ER mouse carrying Kras* and mTmG alleles induced at PN116 by injection of 2 mg tamox and analysed after 53 days also demonstrates adenomas. Dotted line indicates mesothelium; PN, postnatal day; scale bar, 100 μm.
Extended Data Figure 8 Marker expression analysis of alveolar type 2 (AT2) cell derived adenomas.
LysM–Cre > mTmG, KrasLSL-G12D/+ (abbreviated Kras*) lungs stained for the AT2-lineage marker (GFP, green), the nuclear stain DAPI (blue), and the indicated AT2, AT1, and Clara cell markers (red). a–e, Note tumour cells (green) maintain expression of AT2 markers (a, b) and do not turn on Clara (e) or AT1 markers (c, d), except possibly rare cells (*). c′ and d′ show control (non-tumour) regions with normal AT1 staining (arrows). Scale bars, 10 μm.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-3. Supplementary Table 1 contains validated, robust alveolar epithelial cell markers. Supplementary Table 2 shows percentiles of receptor genes expressed by bipotential progenitors and LysM-lineage alveolar type 2 (AT2) cells; and expression levels of EGF receptor family members with histograms of probe-specific levels. Supplementary Table 3 contains genes highly selectively expressed by bipotential progenitors and LysM-lineage alveolar type 2 (AT2) cells at the 90th percentile or higher; and annotation enrichment analysis for these gene profiles. (PDF 475 kb)
Rights and permissions
About this article
Cite this article
Desai, T., Brownfield, D. & Krasnow, M. Alveolar progenitor and stem cells in lung development, renewal and cancer. Nature 507, 190–194 (2014). https://doi.org/10.1038/nature12930
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12930
This article is cited by
-
Bufei Yishen formula protects the airway epithelial barrier and ameliorates COPD by enhancing autophagy through the Sirt1/AMPK/Foxo3 signaling pathway
Chinese Medicine (2024)
-
USP13 drives lung squamous cell carcinoma by switching lung club cell lineage plasticity
Molecular Cancer (2023)
-
Machine learning-identified stemness features and constructed stemness-related subtype with prognosis, chemotherapy, and immunotherapy responses for non-small cell lung cancer patients
Stem Cell Research & Therapy (2023)
-
CTGF promotes the repair and regeneration of alveoli after acute lung injury by promoting the proliferation of subpopulation of AEC2s
Respiratory Research (2023)
-
E26 transformation-specific transcription variant 5 in development and cancer: modification, regulation and function
Journal of Biomedical Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.