Abstract
Heterotaxy is a disorder of left–right body patterning, or laterality, that is associated with major congenital heart disease1. The aetiology and mechanisms underlying most cases of human heterotaxy are poorly understood. In vertebrates, laterality is initiated at the embryonic left–right organizer, where motile cilia generate leftward flow that is detected by immotile sensory cilia, which transduce flow into downstream asymmetric signals2,3,4,5,6. The mechanism that specifies these two cilia types remains unknown. Here we show that the N-acetylgalactosamine-type O-glycosylation enzyme GALNT11 is crucial to such determination. We previously identified GALNT11 as a candidate disease gene in a patient with heterotaxy7, and now demonstrate, in Xenopus tropicalis, that galnt11 activates Notch signalling. GALNT11 O-glycosylates human NOTCH1 peptides in vitro, thereby supporting a mechanism of Notch activation either by increasing ADAM17-mediated ectodomain shedding of the Notch receptor or by modification of specific EGF repeats. We further developed a quantitative live imaging technique for Xenopus left–right organizer cilia and show that Galnt11-mediated Notch1 signalling modulates the spatial distribution and ratio of motile and immotile cilia at the left–right organizer. galnt11 or notch1 depletion increases the ratio of motile cilia at the expense of immotile cilia and produces a laterality defect reminiscent of loss of the ciliary sensor Pkd2. By contrast, Notch overexpression decreases this ratio, mimicking the ciliopathy primary ciliary dyskinesia. Together our data demonstrate that Galnt11 modifies Notch, establishing an essential balance between motile and immotile cilia at the left–right organizer to determine laterality, and reveal a novel mechanism for human heterotaxy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Sutherland, M. J. & Ware, S. M. Disorders of left–right asymmetry: heterotaxy and situs inversus. Am. J. Med. Genet. C. Semin. Med. Genet. 151C, 307–317 (2009)
Nonaka, S. et al. Randomization of left–right asymmetry due to loss of nodal cilia generating leftward flow of extraembryonic fluid in mice lacking KIF3B motor protein. Cell 95, 829–837 (1998)
McGrath, J., Somlo, S., Makova, S., Tian, X. & Brueckner, M. Two populations of node monocilia initiate left–right asymmetry in the mouse. Cell 114, 61–73 (2003)
Yoshiba, S. et al. Cilia at the node of mouse embryos sense fluid flow for left-right determination via Pkd2. Science 338, 226–231 (2012)
Schweickert, A. et al. Cilia-driven leftward flow determines laterality in Xenopus . Curr. Biol. 17, 60–66 (2007)
Kramer-Zucker, A. G. et al. Cilia-driven fluid flow in the zebrafish pronephros, brain and Kupffer’s vesicle is required for normal organogenesis. Development 132, 1907–1921 (2005)
Fakhro, K. A. et al. Rare copy number variations in congenital heart disease patients identify unique genes in left-right patterning. Proc. Natl Acad. Sci. USA 108, 2915–2920 (2011)
Greenway, S. C. et al. De novo copy number variants identify new genes and loci in isolated sporadic tetralogy of Fallot. Nature Genet. 41, 931–935 (2009)
Zaidi, S. et al. De novo mutations in histone-modifying genes in congenital heart disease. Nature 498, 220–223 (2013)
Bennett, E. P. et al. Control of mucin-type O-glycosylation: a classification of the polypeptide GalNAc-transferase gene family. Glycobiology 22, 736–756 (2012)
Schweickert, A. et al. The nodal inhibitor Coco is a critical target of leftward flow in Xenopus . Curr. Biol. 20, 738–743 (2010)
Deblandre, G. A., Wettstein, D. A., Koyano-Nakagawa, N. & Kintner, C. A two-step mechanism generates the spacing pattern of the ciliated cells in the skin of Xenopus embryos. Development 126, 4715–4728 (1999)
Artavanis-Tsakonas, S. & Muskavitch, M. A. Notch: the past, the present, and the future. Curr. Top. Dev. Biol. 92, 1–29 (2010)
Wettstein, D. A., Turner, D. L. & Kintner, C. The Xenopus homolog of Drosophila Suppressor of Hairless mediates Notch signaling during primary neurogenesis. Development 124, 693–702 (1997)
Rana, N. A. & Haltiwanger, R. S. Fringe benefits: functional and structural impacts of O-glycosylation on the extracellular domain of Notch receptors. Curr. Opin. Struct. Biol. 21, 583–589 (2011)
Schwientek, T. et al. Functional conservation of subfamilies of putative UDP-N-acetylgalactosamine:polypeptide N-acetylgalactosaminyltransferases in Drosophila, Caenorhabditis elegans, and mammals. One subfamily composed of l(2)35Aa is essential in Drosophila . J. Biol. Chem. 277, 22623–22638 (2002)
Brou, C. et al. A novel proteolytic cleavage involved in Notch signaling: the role of the disintegrin-metalloprotease TACE. Mol. Cell 5, 207–216 (2000)
Gram Schjoldager, K. T. et al. A systematic study of site-specific GalNAc-type O-glycosylation modulating proprotein convertase processing. J. Biol. Chem. 286, 40122–40132 (2011)
Kato, Y. The multiple roles of Notch signaling during left-right patterning. Cell. Mol. Life Sci. 68, 2555–2567 (2011)
Lopes, S. S. et al. Notch signalling regulates left-right asymmetry through ciliary length control. Development 137, 3625–3632 (2010)
Krebs, L. T. et al. Notch signaling regulates left–right asymmetry determination by inducing Nodal expression. Genes Dev. 17, 1207–1212 (2003)
Pennekamp, P. et al. The ion channel polycystin-2 is required for left-right axis determination in mice. Curr. Biol. 12, 938–943 (2002)
Duldulao, N. A., Lee, S. & Sun, Z. Cilia localization is essential for in vivo functions of the Joubert syndrome protein Arl13b/Scorpion. Development 136, 4033–4042 (2009)
Raya, A. et al. Notch activity acts as a sensor for extracellular calcium during vertebrate left–right determination. Nature 427, 121–128 (2004)
Stubbs, J. L., Oishi, I., Izpisua Belmonte, J. C. & Kintner, C. The forkhead protein Foxj1 specifies node-like cilia in Xenopus and zebrafish embryos. Nature Genet. 40, 1454–1460 (2008)
Bisgrove, B. W., Makova, S., Yost, H. J. & Brueckner, M. RFX2 is essential in the ciliated organ of asymmetry and an RFX2 transgene identifies a population of ciliated cells sufficient for fluid flow. Dev. Biol. 363, 166–178 (2012)
Chung, M. I. et al. RFX2 is broadly required for ciliogenesis during vertebrate development. Dev. Biol. 363, 155–165 (2012)
Shinohara, K. et al. Two rotating cilia in the node cavity are sufficient to break left-right symmetry in the mouse embryo. Nature Commun. 3, 622 (2012)
Kamura, K. et al. Pkd1l1 complexes with Pkd2 on motile cilia and functions to establish the left-right axis. Development 138, 1121–1129 (2011)
Vick, P. et al. Flow on the right side of the gastrocoel roof plate is dispensable for symmetry breakage in the frog Xenopus laevis . Dev. Biol. 331, 281–291 (2009)
del Viso, F. & Khokha, M. Generating diploid embryos from Xenopus tropicalis . Methods Mol. Biol. 917, 33–41 (2012)
Nieuwkoop, P. D. & Faber, J. Normal Table of Xenopus laevis (Daudin): a Systematical and Chronological Survey of the Development from the Fertilized Egg Till the End of Metamorphosis (Garland, 1994)
Khokha, M. K. et al. Techniques and probes for the study of Xenopus tropicalis development. Dev. Dyn. 225, 499–510 (2002)
Vonica, A. & Brivanlou, A. H. The left–right axis is regulated by the interplay of Coco, Xnr1 and derriere in Xenopus embryos. Dev. Biol. 303, 281–294 (2007)
Caspary, T., Larkins, C. E. & Anderson, K. V. The graded response to Sonic Hedgehog depends on cilia architecture. Dev. Cell 12, 767–778 (2007)
Piperno, G. & Fulller, M. T. Monoclonal antibodies specific for an acetylated form of alpha-tubulin recognize the antigen in cilia and flagella from a variety of organisms. J. Cell Biol. 101, 2085–2094 (1985)
Pedersen, J. W. et al. Lectin domains of polypeptide GalNAc transferases exhibit glycopeptide binding specificity. J. Biol. Chem. 286, 32684–32696 (2011)
Walentek, P., Beyer, T., Thumberger, T., Schweickert, A. & Blum, M. ATP4a is required for Wnt-dependent Foxj1 expression and leftward flow in Xenopus left-right development. Cell Rep. 1, 516–527 (2012)
Steentoft, C. et al. Mining the O-glycoproteome using zinc-finger nuclease-glycoengineered SimpleCell lines. Nature Methods 8, 977–982 (2011)
Steentoft, C. et al. Precision mapping of the human O-GalNAc glycoproteome through SimpleCell technology. EMBO J. 32, 1478–1488 (2013)
Acknowledgements
We thank the patients and their families, who are the inspiration for this study. We thank S. Kubek and M. Slocum for animal husbandry. We thank U. Mandel, M. Vester-Christensen, T. D. Madsen and S. B. Levery for help with generation of polyclonal antibodies, recombinant GALNT enzymes, in vitro glycosylation experiments and mass spectrometry. Thanks to the Center for Cellular and Molecular Imaging at Yale for assistance with SEM/TEM (C. Rahner and M. Graham) and confocal imaging. Thanks to C. Kintner and Z. Sun for reagents. Thanks to R. Harland, K. Liem, R. Lifton, J. McGrath, A. Horwich and Z. Sun for critical reading of the manuscript. M.T.B. was supported by grant number TL 1 RR024137 from the National Center for Research Resources/National Institutes of Health (NIH) and NIH Roadmap for Medical Research. S.Y. was supported by NIH training grant 5T32HD00709436. This work was supported by NIH R01HL093280 to M.B. and NIH DE018825, DE018824 to M.K.K., and the Danish National Research Foundation (DNRF107) to H.C.
Author information
Authors and Affiliations
Contributions
M.T.B., S.Y., S.M., M.B. and M.K.K. conceived and designed the embryological experiments in Xenopus and mouse. M.T.B., S.Y., S.M. and M.K.K. performed the Xenopus experiments and M.B. performed mouse experiments. N.B.P., C.K.G. and H.C. performed and evaluated the glycosylation and ADAM studies. All authors wrote and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Galnt11 is enriched in the crown cells at the mouse LRO.
Mouse nodes (E8.0) labelled with anti-Arl13b in red for cilia, Galnt11 in green and nuclei (Hoechst) in blue. a, A maximal Z-projection showing the entire LRO (node), viewed ventrally with posterior to the top and left side to the right. b, Higher magnification of the lateral aspect of the node. c, Orthologue view (y–z axis) of the corresponding z-stack demonstrating Galnt11 expression bordering the cilia in the node pit. d, A maximal Z-projection showing the entire crown and pit of the node of another E8.0 embryo, viewed ventrally with posterior to the top and left to the right. e, Higher magnification of the lateral aspect of the node.
Extended Data Figure 2 galnt11 knockdown does not affect epidermal cilia structure.
a–d, Wild-type (a, c) or galnt11 morphant (b, d) multiciliated epidermal cells were imaged using scanning electron microscopy (a, b) or transmission electron microscopy (c, d). No consistent abnormalities in axonemal structure were noted in galnt11 morphants.
Extended Data Figure 3 Galnt11 and Notch affect left–right development similarly.
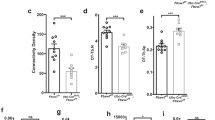
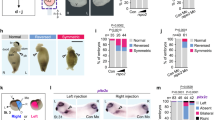
Histograms indicating the percentage of embryos with abnormal coco, pitx2c and cardiac looping with gain or loss of function of galnt11 or notch1. *P < 0.05, **P < 0.01, ***P < 0.001. Graphs depict means. Additional details are in Methods. coco expression: uninjected control (UC) n = 21, galnt11 MO n = 24, notch1 MO n = 30, GALNT11 RNA n = 23, nicd RNA n = 22. pitx2c expression: UC n = 45, galnt11 MO n = 54, notch1 MO n = 50, GALNT11 RNA n = 34, nicd RNA n = 30. Heart looping UC n = 276, galnt11 MO n = 68, notch1 MO n = 75, GALNT11 RNA n = 91, nicd RNA n = 50.
Extended Data Figure 4 Glycosylation of peptides in the NOTCH1 extracellular domain.
Schematic of the extracellular domain of human NOTCH1. The 36 epidermal-like growth factor (EGF) repeats are numbered to the left with their six conserved cysteine residues marked in brown. Underlined are peptides (designated numbers at right side) analysed as substrates by in vitro glycosylation with recombinant GALNTs. Yellow squares indicate the position of GalNAc residues incorporated as evaluated by electrospray ionization-linear ion trap-Fourier transform mass spectrometry (ESI-LIT-FT-MS). Far right column lists GALNT isoforms tested that incorporated GalNAc residues into the peptides. Cases in which sites of GalNAc incorporation were determined are marked with an asterisk. Scissor symbol indicates the S2 cleavage site. TM, transmembrane domain as predicted by TMHMM (http://www.cbs.dtu.dk/services/TMHMM/). Simplified consensus sequence motifs for O-fucosylation (red triangle), O-glucosylation (blue circle) and O-GlcNAc (blue square) are shown at the top. The complete consensus motifs are: O-glucosylation, C1XSX(A/P)C2; O-fucosylation, C2X3(A/G/S)(S/T)C3; O-GlcNAc, C5XX(G/H)(Y/F/L)(S/T)GX2–5C6, with underlining indicating the O-glycan attachment site.
Extended Data Figure 5 Cleavage controls and human GALNT11 catalytic site.
a, Typically GalNAc glycosylation inhibits cleavage of peptides such as this example from TNR16. b, ADAM17 is necessary for cleavage of the NOTCH1 juxtamembrane peptide. 2-h incubation of the naked peptide and the glycopeptides without ADAM17 does not give rise to the cleavage products seen in Fig. 2. c, Mutation of the catalytic site DSH to DSA abrogates the activity of GALNT11 to alter left–right patterning. Histograms indicate the number of embryos with abnormal pitx2c expression or cardiac looping. ***P < 0.001. Graphs depict means. pitx2c expression: UC n = 45, GALNT11 wild type n = 34, GALNT11(H247A) n = 40. Heart looping: UC n = 243, GALNT11 wild type n = 91, GALNT11(H247A) n = 133. Additional details are in Methods. d, Expression of the mutant GALNT11(H247A) in transfected IMCD3 cells has similar expression levels and subcellular localization as wild-type GALNT11 on the basis of immunofluorescence studies. Red, Galnt11; blue, Hoechst.
Extended Data Figure 6 Imaging motile and immotile cilia in the LRO and effects on total cilia number.
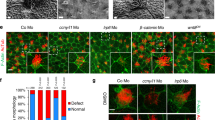
a, A maximal stack projection through a high-speed fluorescent acquisition of a wild-type LRO explant expressing arl13b-mCherry (cyan) reveals distinct populations of immotile and motile cilia (elapsed time = 1 s). Cyan box highlights a single motile cilium and magenta box highlights a single immotile cilium. b, c, Spatial mapping of cilia in the same wild-type LRO for immotile cilia (magenta) (b) and motile cilia (cyan) (c). d, Two-colour merge of immotile cilia (magenta) and motile cilia (cyan) spatial maps. e, Total area of the LRO (magenta). AU, arbitrary units. f, Centre of the LRO. g, Combining e and f identifies centre and periphery. h, The periphery (light magenta) and centre outlines (light cyan) are merged back onto the original wild-type LRO acquisition data and the numbers of motile (cyan) and immotile (magenta) cilia within each defined area are scored. i, Graph representing the mean cilia number per LRO in vehicle, nicd overexpressants, notch1 morphants and galnt11 morphants. Although modest changes in total cilia numbers were observed in experimental samples, they were not statistically significant. Data were quantified from the same data set presented in Fig. 3g. j, Total numbers of cilia scored on right or left of the wild-type LRO represented as percentage of total population. No significant change in total cilia number is present across left–right axis. k, Motile cilia scored on right or left of the wild-type LRO represented as percentage of total motile cilia population. NS, P ≥ 0.05.
Extended Data Figure 7 Early markers of the LRO and motile cilia.
a–h, In situ hybridization for foxj1 (a–d) and rfx2 (e–h) at the Xenopus LRO in uninjected control (a, e), nicd overexpressants (b, f), GALNT11 overexpressants (c, g) and GALNT11(H247A) overexpressants (d, h). GALNT11 or nicd reduce the expression of foxj1 and rfx2, which are upstream transcription factors for ciliary motility proteins. Pink stain indicates co-injection with lacZ to trace the RNA. Overexpression of GALNT11(H247A) does not affect either foxj1 or rfx2. All GRPs are viewed ventrally with anterior to the top (right side of embryo to left of figure). i, Schematic of LRO border markers at stage 16 and stage 20. j–u, Expression of peripheral LRO markers in galnt11 morphants and uninjected controls. coco expression (j, k, p, q) is either asymmetric in j, p or symmetric in k, q for stage 16 embryos. xnr1 (l, r) and gdf3 (m, s) expression is symmetric at stage 16. coco expression in stage 20 embryos is asymmetric (n, t) or symmetric (o, u) but the breadth of expression is not dramatically different. All GRPs (wild type, j–o; galnt11 morphants, p–u) are viewed ventrally with anterior to the top (right side of embryo to left of figure).
Extended Data Figure 8 Galnt11 modifies Notch to alter cilia type and left–right patterning.
a, Model describing the relation between motile cilia and non-motile (sensory) cilia and the relation between Galnt11 glycosylation and Notch signalling. Galnt11 activates Notch signalling, increasing immotile cilia while suppressing motile cilia, probably via foxj1/rfx2. b, In wild-type embryos, motile and immotile cilia numbers and spatial distribution are balanced by Galnt11/Notch signalling. c, Activation of Galnt11/Notch decreases motile cilia, resulting in phenotypes similar to primary ciliary dyskinesia—symmetric or reversed laterality. d, Reduction of Galnt11/Notch signalling increases motile cilia, resulting in loss of laterality, a phenotype similar to loss of ciliary sensation in Pkd2 mutants.
Supplementary information
Wildtype Xenopus embryos display a balanced population of immotile and motile cilia types in the LRO
Live high-speed laser confocal acquisition of wildtype embryos co-injected with arl13b-mCherry RNA (cilia; cyan) and water (vehicle). Yellow trace outlines the GRP/LRO. A: anterior, P: posterior, L: left, R: right. (MOV 10384 kb)
nicd overexpression decreases the ratio of motile to immotile cilia in the LRO
Live high-speed laser confocal acquisition of wildtype embryos co-injected with arl13b-mCherry RNA (cilia; cyan) and nicd RNA. Yellow trace outlines the GRP/LRO. A: anterior, P: posterior, L: left, R: right. (MOV 10640 kb)
notch1 knockdown increases the ratio of motile to immotile cilia in the LRO
Live high-speed laser confocal acquisition of wildtype embryos co-injected with arl13b-mCherry RNA (cilia; cyan) and notch1 MO. Yellow trace outlines the GRP/LRO. A: anterior, P: posterior, L: left, R: right. (MOV 10486 kb)
galnt11 knockdown increases the ratio of motile to immotile cilia in the LRO
Live high-speed laser confocal acquisition of wildtype embryos co-injected with arl13b-mCherry RNA (cilia; cyan) and galnt11 MO. Yellow trace outlines the GRP/LRO. A: anterior, P: posterior, L: left, R: right. (MOV 10470 kb)
Rights and permissions
About this article
Cite this article
Boskovski, M., Yuan, S., Pedersen, N. et al. The heterotaxy gene GALNT11 glycosylates Notch to orchestrate cilia type and laterality. Nature 504, 456–459 (2013). https://doi.org/10.1038/nature12723
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12723
This article is cited by
-
Identification of global inhibitors of cellular glycosylation
Nature Communications (2023)
-
A change of heart: new roles for cilia in cardiac development and disease
Nature Reviews Cardiology (2022)
-
The second DDOST-CDG patient with lactose intolerance, developmental delay, and situs inversus totalis
Journal of Human Genetics (2022)
-
Disruption of GMNC-MCIDAS multiciliogenesis program is critical in choroid plexus carcinoma development
Cell Death & Differentiation (2022)
-
Cystic kidney diseases associated with mutations in phosphomannomutase 2 promotor: a large spectrum of phenotypes
Pediatric Nephrology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.