Abstract
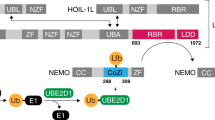
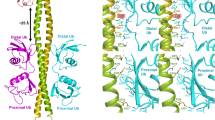
Linear ubiquitin chains are important regulators of cellular signalling pathways that control innate immunity and inflammation through nuclear factor (NF)-κB activation and protection against tumour necrosis factor-α-induced apoptosis1,2,3,4,5. They are synthesized by HOIP, which belongs to the RBR (RING-between-RING) family of E3 ligases and is the catalytic component of LUBAC (linear ubiquitin chain assembly complex), a multisubunit E3 ligase6. RBR family members act as RING/HECT hybrids, employing RING1 to recognize ubiquitin-loaded E2 while a conserved cysteine in RING2 subsequently forms a thioester intermediate with the transferred or ‘donor’ ubiquitin7. Here we report the crystal structure of the catalytic core of HOIP in its apo form and in complex with ubiquitin. The carboxy-terminal portion of HOIP adopts a novel fold that, together with a zinc-finger, forms a ubiquitin-binding platform that orients the acceptor ubiquitin and positions its α-amino group for nucleophilic attack on the E3∼ubiquitin thioester. The C-terminal tail of a second ubiquitin molecule is located in close proximity to the catalytic cysteine, providing a unique snapshot of the ubiquitin transfer complex containing both donor and acceptor ubiquitin. These interactions are required for activation of the NF-κB pathway in vivo, and they explain the determinants of linear ubiquitin chain specificity by LUBAC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Walczak, H., Iwai, K. & Dikic, I. Generation and physiological roles of linear ubiquitin chains. BMC Biol. 10, 23 (2012)
Tokunaga, F. et al. Involvement of linear polyubiquitylation of NEMO in NF-κB activation. Nature Cell Biol. 11, 123–132 (2009)
Ikeda, F. et al. SHARPIN forms a linear ubiquitin ligase complex regulating NF-κB activity and apoptosis. Nature 471, 637–641 (2011)
Tokunaga, F. et al. SHARPIN is a component of the NF-κB-activating linear ubiquitin chain assembly complex. Nature 471, 633–636 (2011)
Gerlach, B. et al. Linear ubiquitination prevents inflammation and regulates immune signalling. Nature 471, 591–596 (2011)
Kirisako, T. et al. A ubiquitin ligase complex assembles linear polyubiquitin chains. EMBO J. 25, 4877–4887 (2006)
Wenzel, D. M., Lissounov, A., Brzovic, P. S. & Klevit, R. E. UBCH7 reactivity profile reveals Parkin and HHARI to be RING/HECT hybrids. Nature 474, 105–108 (2011)
Pickart, C. M. & Fushman, D. Polyubiquitin chains: polymeric protein signals. Curr. Opin. Chem. Biol. 8, 610–616 (2004)
Deshaies, R. J. & Joazeiro, C. A. RING domain E3 ubiquitin ligases. Annu. Rev. Biochem. 78, 399–434 (2009)
Huibregtse, J. M., Scheffner, M., Beaudenon, S. & Howley, P. M. A family of proteins structurally and functionally related to the E6-AP ubiquitin-protein ligase. Proc. Natl Acad. Sci. USA 92, 2563–2567 (1995)
Wang, M. & Pickart, C. M. Different HECT domain ubiquitin ligases employ distinct mechanisms of polyubiquitin chain synthesis. EMBO J. 24, 4324–4333 (2005)
Eisenhaber, B., Chumak, N., Eisenhaber, F. & Hauser, M. T. The ring between ring fingers (RBR) protein family. Genome Biol. 8, 209 (2007)
Wenzel, D. M. & Klevit, R. E. Following Ariadne’s thread: a new perspective on RBR ubiquitin ligases. BMC Biol. 10, 24 (2012)
Stieglitz, B., Morris-Davies, A. C., Koliopoulos, M. G., Christodoulou, E. & Rittinger, K. LUBAC synthesizes linear ubiquitin chains via a thioester intermediate. EMBO Rep. 13, 840–846 (2012)
Smit, J. J. et al. The E3 ligase HOIP specifies linear ubiquitin chain assembly through its RING-IBR-RING domain and the unique LDD extension. EMBO J. 31, 3833–3844 (2012)
Trempe, J. F. et al. Structure of Parkin reveals mechanisms for ubiquitin ligase activation. Science 340, 1451–1455 (2013)
Wauer, T. & Komander, D. Structure of the human Parkin ligase domain in an autoinhibited state. EMBO J. 32, 2099–2112 (2013)
Spratt, D. E. et al. A molecular explanation for the recessive nature of Parkin-linked Parkinson’s disease. Nature Commun. 4, 1983, http://dx.doi.org/10.1038/ncomms2983 (2013)
Riley, B. E. et al. Structure and function of Parkin E3 ubiquitin ligase reveals aspects of RING and HECT ligases. Nature Commun. 4, 1982, http://dx.doi.org/10.1038/ncomms2982 (2013)
Duda, D. M. et al. Structure of HHARI, a RING-IBR-RING ubiquitin ligase: autoinhibition of an Ariadne-family E3 and insights into ligation mechanism. Structure 21, 1030–1041 (2013)
Beasley, S. A., Hristova, V. A. & Shaw, G. S. Structure of the Parkin in-between-ring domain provides insights for E3-ligase dysfunction in autosomal recessive Parkinson’s disease. Proc. Natl Acad. Sci. USA 104, 3095–3100 (2007)
Yunus, A. A. & Lima, C. D. Lysine activation and functional analysis of E2-mediated conjugation in the SUMO pathway. Nature Struct. Mol. Biol. 13, 491–499 (2006)
Wu, P. Y. et al. A conserved catalytic residue in the ubiquitin-conjugating enzyme family. EMBO J. 22, 5241–5250 (2003)
Wickliffe, K. E., Lorenz, S., Wemmer, D. E., Kuriyan, J. & Rape, M. The mechanism of linkage-specific ubiquitin chain elongation by a single-subunit E2. Cell 144, 769–781 (2011)
Eddins, M. J., Carlile, C. M., Gomez, K. M., Pickart, C. M. & Wolberger, C. Mms2-Ubc13 covalently bound to ubiquitin reveals the structural basis of linkage-specific polyubiquitin chain formation. Nature Struct. Mol. Biol. 13, 915–920 (2006)
Petroski, M. D. & Deshaies, R. J. Mechanism of lysine 48-linked ubiquitin-chain synthesis by the cullin-RING ubiquitin-ligase complex SCF-Cdc34. Cell 123, 1107–1120 (2005)
Higuchi, R., Krummel, B. & Saiki, R. K. A general method of in vitro preparation and specific mutagenesis of DNA fragments: study of protein and DNA interactions. Nucleic Acids Res. 16, 7351–7367 (1988)
Raasi, S. & Pickart, C. M. Ubiquitin chain synthesis. Methods Mol. Biol. 301, 47–55 (2005)
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Brown, P. H. & Schuck, P. Macromolecular size-and-shape distributions by sedimentation velocity analytical ultracentrifugation. Biophys. J. 90, 4651–4661 (2006)
Plechanovova, A., Jaffray, E. G., Tatham, M. H., Naismith, J. H. & Hay, R. T. Structure of a RING E3 ligase and ubiquitin-loaded E2 primed for catalysis. Nature 489, 115–120 (2012)
Dou, H., Buetow, L., Sibbet, G. J., Cameron, K. & Huang, D. T. BIRC7-E2 ubiquitin conjugate structure reveals the mechanism of ubiquitin transfer by a RING dimer. Nature Struct. Mol. Biol. 19, 876–883 (2012)
Pruneda, J. N. et al. Structure of an E3:E2∼Ub complex reveals an allosteric mechanism shared among RING/U-box ligases. Mol. Cell 47, 933–942 (2012)
Acknowledgements
We thank S. Strekopytov for inductively coupled plasma mass spectrometry experiments, I. Taylor for help with analytical ultracentrifugation, L. Haire for help with crystallization, S. Martin for circular dichroism spectroscopy, S. Caswell for help with cloning, C. Bayart for technical assistance, P. Walker for data collection, S. Smerdon for discussions and advice, and the Diamond Light Source for synchrotron access. This work was supported by the Medical Research Council (grant U117565398) and the Wellcome Foundation (grant 094112/Z/10/Z) to K.R., and the European Research Council (ERC) under the European Union’s Seventh Framework Programme (FP7/2007-2013) and ERC grant agreement 250241-LineUb to I.D.
Author information
Authors and Affiliations
Contributions
B.S., R.R.R. and M.G.K. purified proteins and performed structural and biochemical analysis. A.C.M.-D. and N.R.B. purified proteins and conducted biochemical analysis. V.S. performed in vivo studies. E.C. produced expression plasmids. S.H. performed MS analysis. I.D. coordinated experimental work and contributed ideas. K.R. contributed to structural analysis and wrote the paper. All authors contributed to data analysis, experimental design and paper writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Comparison between the catalytic activities of the crystallized HOIPCBR-C construct and the HOIPRBR-C construct.
a, Steady-state ubiquitination assays comparing the catalytic activity of the RBR-C construct of HOIP (HOIPRBR-C) with the CBR-C construct (HOIPCBR-C), showing that activity is reduced in HOIPCBR-C with only short ubiquitin chains formed after 1 h. For this reason, all steady-state assays were performed with HOIPRBR-C. To confirm that HOIPCBR-C had retained the ability to specifically synthesize linear chains, a ubiquitination assay was performed with ubiquitin containing an N-terminal His6 tag that was no longer able to produce linear chains (right-hand gel). All gels were stained with Coomassie blue and converted to black and white. b, To ensure that any effects seen in ubiquitination assays were not due to inefficient loading of ubiquitin mutants onto E1 or E2 enzymes, the mutants were tested for their ability to form a thioester quantitatively with E1 and UbcH5A within 5 min. Samples were analysed on SDS gels and loaded in sample buffer in the absence or presence of dithiothreitol to detect the thioester.
Extended Data Figure 2 Topology of HOIPCBR-C, content of the asymmetric unit of the apo crystals, solution behaviour of this fragment and comparison with the ubiquitin bound structure.
a, Topology diagram of HOIPCBR-C maintaining the same colour scheme as in Fig. 1. b, The apo HOIPCBR-C structure contains four molecules of HOIPCBR-C in the asymmetric unit that overlap with root mean squared deviation values of 0.8–1.0 Å. They are related by a local two-fold axis, and two molecules form a disulphide bond (shown in cyan and blue) and coordinate a fifth Zn2+ ion. Boxed: details of disulphide formation and zinc coordination by H887 and H889 of each monomer. Some of the loop regions in the apo structure are disordered. These include: residues 866–868 in two monomers, 880–882 in all monomers, 960–964 in all monomers and 978–983, 973–982, 975–982 and 974–983 in the respective monomers. c, Sedimentation velocity run of the crystallized construct at a sample concentration of 125 μM and derived C(S) distribution (Sapp = 2.38; S20,w = 2.56), indicating that it exists as a monomer in solution. d, Overlap of the apo (blue) and ubiquitin-bound (cyan) structures highlighting the regions that become ordered on complex formation (indicated by a dotted line) with ubiquitin (shown in grey). C885, H887 and H889 are shown as sticks and the Zn2+ ions as spheres.
Extended Data Figure 3 Comparison of CBR domains from different RBR family members.
a, Sequence alignment of different CBR domains. Conserved cysteine and histidine residues in the CBR involved in coordinating zinc 1 and zinc 2 are highlighted in yellow, and residues making hydrophobic contacts with the C-terminal tail of the donor ubiquitin are highlighted in green. The sequence forming ZF1 in HOIP has been removed for clarity and is indicated in cyan. The region around the catalytic C885 and H887, which are both crucial for catalytic activity, is highlighted in red. The glycine preceding C885 is conserved in other RBRs and might be important to allow the ubiquitin-loaded E2 access to the catalytic cysteine. The sequence variation of site 2 is much higher than for site 1, and in particular the sequence between zinc-coordinating residues 2 and 3, which in HOIP accommodates ZF1, varies significantly in length. b, This sequence variation is reflected in the structures of CBRs from HOIP, Parkin (PDB 4K7D)16 and HHARI (PDB 4KC9)20, which overlap well in site 1 and the following first two zinc-coordinating residues of site 2 but subsequently diverge significantly. Nevertheless, the positions of zinc 2 overlap well. The IBR of HOIP (pink; PDB 2CT7)21 is shown for comparison in the overlap. The ZF1 is shown in cyan and the Zn2+ ions are shown as spheres. c, HOIPCBR-C in complex with the donor ubiquitin, overlapped with the CBRs of Parkin and HHARI, showing conserved hydrophobic residues that contact L71 and L73 of ubiquitin in ball-and-stick representation as well as the catalytic cysteine of the ligase. The C-terminal portion of the CBR (site 2) has been omitted for clarity. This overlap shows clearly that the C-terminal tail of the donor ubiquitin could be accommodated in a similar manner in Parkin and HHARI.
Extended Data Figure 4 The HOIPCBR-C–ubiquitin complex in the asymmetric unit and arrangement in the crystal lattice.
a, The asymmetric unit contains one molecule of HOIPCBR-C (blue) and one molecule of ubiquitin (orange). The ubiquitin bound to HOIPCBR-C represents the acceptor ubiquitin; its α-amino group of M1 is located in close proximity to the thioester-forming C885, which is shown in ball-and-stick representation and the Zn2+ ions are shown as grey spheres, including the Zn2+ ion found in the active site. Inset: details of the active site highlighting the proximity of M1 and C885 and H887. The Zn2+ ion found in the active site has been removed for clarity. b, HOIPCBR-C and ubiquitin that constitute the asymmetric unit are shown in blue and orange, respectively. A symmetry-related complex that contributes the donor ubiquitin is shown in yellow (ubiquitin) and light blue (HOIP). All other complexes in the lattice are shown in grey (HOIP) and red (ubiquitin).
Extended Data Figure 5 Active site arrangement including coordination of the fifth Zn2+ ion and ubiquitination assays with wild-type HOIPRBR-C and the H889A mutant.
a, We found residual electron density in the active site of the wild-type HOIPCBR-C–ubiquitin complex (shown in transparent grey), which adopts a tetrahedral coordination and which we interpret as a Zn2+ ion. This Zn2+ is coordinated by the catalytic cysteine, H887, the α-amino group of ubiquitin and an imidazole from the crystallization solution (Imd). The observation that apo and substrate-bound HOIPCBR-C contain a metal ion close to the active site prompted us to investigate a possible role in catalysis. Metal binding was examined by native electrospray mass spectrometry and ICP–MS, which indicated the presence of roughly five Zn2+ ions in the wild-type protein. Mutation of either of the histidines H887A and H889A, which coordinate the additional zinc in the apo structure, decreased the number to four. The structure of the H889A mutant in complex with ubiquitin (in blue and orange, respectively) lacks additional electron density in the active site, while retaining full catalytic activity, indicating that the catalytic step is not metal-dependent. Instead we believe that high reactivity of the active site induces disulphide bond formation and subsequent zinc coordination across the interface during crystallization of apo HOIPCBR-C, whereas in the substrate-bound complex the active site cysteine itself coordinates a metal ion with the help of an imidazole molecule from the crystallization buffer. b, Sigma-A weighted omit map contoured at 1.5σ showing the Zn2+ ion in the active site and its coordination by the α-amino group of ubiquitin M1, C885 and H887 from HOIPCBR-C, and an imidazole molecule from the crystallization buffer. c, Ubiquitination assays comparing the activity of wild-type HOIPRBR-C and the H889A mutant that no longer coordinates a fifth Zn2+ ion but retains full catalytic activity. d, The molecular mass of each construct listed (in daltons) was determined in its native and denatured forms by ESI–MS. The difference in mass of native and denatured proteins was used to calculate the number of Zn2+ ions present (63.4 Da per Zn2+). The calculated mass of the constructs under investigation contains the additional sequence GPG that remains after removal of the glutathione S-transferase tag with PreScission protease. Metal analysis by ICP–MS did not reveal the presence of significant amounts of metal ions apart from Zn2+.
Extended Data Figure 6 Diagram of the HOIPCBR-C–ubiquitin complex interface and complete set of steady-state ubiquitination assays performed with HOIPRBR-C.
a, Diagram of the interface between HOIPCBR-C and donor and acceptor ubiquitin, respectively. The colour scheme used in Fig. 1 has been maintained, with a cut-off of 3.5 Å for polar and 4.0 Å for hydrophobic interactions. The C terminus of the donor ubiquitin is oriented towards the catalytic cysteine through multiple interactions with HOIP. The carbonyl of G76 forms a hydrogen bond with the C885 backbone amide and is 3.5 Å distant from its Sγ. The backbone NH of G76 makes further contact with the loop carrying C885, whereas an extended conformation for the rest of the tail is maintained by interactions with the CBR and β-hairpin. This arrangement is reminiscent of RING ligases that lock the E2∼Ub in a folded-back conformation35,36,37, indicating that this might be a general mechanism to activate a ubiquitin thioester intermediate. b, The interface between HOIP and the acceptor ubiquitin includes residues from the helical base of HOIP that interact with T14, E16, D32 and K33. Mutation of T14 in HOIPRBR-C to alanine has only a modest effect on the steady-state synthesis of linear ubiquitin chains, whereas E16 and D32 show significantly decreased activity; the strongest effect was seen with the K33A mutant. In HOIPRBR-C, mutation of R935 and D936 almost completely abrogates activity, whereas the R1032A mutant has only a minor effect. c, E3-thioester formation and ubiquitin transfer assays. The diagram illustrates the set-up: lanes 1–4, donor Cy5-ubiquitin loading onto E1 and E2, followed by addition of wild-type HOIPRBR-C or mutants to form an E3∼thioester (lanes1 and 2 of each experiment). To monitor ubiquitin transfer, a C-terminally blocked ubiquitin was added (lanes 3 and 4 of each experiment). d, Mutations that disrupt the interaction of the C-terminal tail of the donor ubiquitin with the conserved region of the CBR and the β-hairpin strongly decrease the ability to produce linear ubiquitin chains. In contrast, mutation of E866, which is not conserved in other RBR family members, and its contact R42 has no effect on ubiquitin chain synthesis. Some of the gels included in this figure are also shown in Fig. 3c and are included for comparison.
Extended Data Figure 7 In vitro ubiquitination assays with heterotrimeric LUBAC and in vivo co-immunoprecipitation of HOIP mutants with SHARPIN and HOIL-1L.
a, In vitro ubiquitination assays with heterotrimeric LUBAC, showing that the overall activity of LUBAC is lower than that of isolated HOIPRBR-C, possibly as a result of regulatory roles of the ubiquitin-binding domains that are present in all three LUBAC subunits. Nevertheless, the trends observed with isolated HOIPRBR-C are conserved with LUBAC: those mutations that had only a minor or no effect on chain synthesis (T14A and R42) show the same behaviour, whereas the E16A, K33A, L71A, L73A and R74A mutants show a significant decrease in ubiquitin chain synthesis. b, Co-immuniprecipitation assays show that mutations in HOIP that interfere with the binding of donor or acceptor ubiquitin and interfere with ubiquitin chain synthesis have no effect on complex formation between LUBAC subunits, and hence any effects seen in NF-κB activation and p65 translocation assays are not due to impaired complex formation.
Extended Data Figure 8 pH dependence of ubiquitin transfer with wild-type HOIPRBR-C and the H887A mutant.
Ube1 (0.5 μM) was charged with 1 μM His6-Cy5-ubiquitin (Ub) using 5 mM ATP at pH 7. a, The pre-charged E1-ubiquitin thioester (E1∼Ub) was subsequently mixed with 10 μM UbcH5A and incubated for 5 min under different buffer conditions ranging from pH 7 to pH 11. Complete ubiquitin transthiolation from the E1 onto UbcH5A (E2∼Ub) can be observed at pH 7–9, is impaired at pH 10 and abolished at pH 11. Wild-type HOIPRBR-C (20 μM; E3(wt), top row) or HOIPRBR-C H887A (20 μM; E3(H887A), bottom row) were added to each sample and incubated for a further 5 min. Under these conditions, a thioester intermediate for both wild-type (E3(wt)∼Ub) and mutant HOIPRBR-C (E3(H887A)∼Ub) can be detected at pH 7–9. All samples were finally mixed with 10 μM C-terminal His6-tagged ubiquitin (Ub). HOIPRBR-C wild-type catalyses the formation of di-ubiquitin (Ub2) at pH 7–9 and to some extent at pH 10. In contrast, product formation is absent at pH 7 and pH 8 for HOIPRBR-C H887A, indicating that histidine 887 is required for catalysis under physiological pH conditions. The assay was performed in 5 mM MgCl2, 150 mM NaCl and 200 mM buffer (HEPES pH 7.0, HEPES pH 8.0, CHES pH 9.0, CHES pH 10.0 and CAPS pH 11.0). b, The pre-charged E1-ubiquitin thioester (E1∼Ub) was mixed with 10 μM UbcH5A (E2) and incubated for 5 min in 150 mM sodium acetate buffer ranging from pH 5.2 to pH 6.6 with 0.1 increments. All samples display the same amount of charged UbcH5A (E2∼Ub). Similarly, the addition of 20 μM HOIPRBR-C (E3) shows formation of the thioester charged intermediate (E3∼Ub) to the same extent. Each sample was mixed with 10 μM C-terminal His-tagged diubiquitin (Ub2) as acceptor to allow product formation (Ub3). The discharge of E3-thioester ubiquitin onto the acceptor is clearly impaired at pH values below 5.8. Gels were run under non-reducing conditions.
Extended Data Figure 9 The structural basis of chain linkage specificity.
Surface representation of HOIPCBR-C with the acceptor ubiquitin in orange ribbon representation, indicating the position of all seven lysine residues present in ubiquitin, as well as the N-terminal methionine and the catalytic cysteine, C885, of HOIP. The figure clearly shows that the α-amino group of methionine 1 is closest to the active-site cysteine, explaining the specificity for linear chains.
Rights and permissions
About this article
Cite this article
Stieglitz, B., Rana, R., Koliopoulos, M. et al. Structural basis for ligase-specific conjugation of linear ubiquitin chains by HOIP. Nature 503, 422–426 (2013). https://doi.org/10.1038/nature12638
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12638
This article is cited by
-
Structural snapshots along K48-linked ubiquitin chain formation by the HECT E3 UBR5
Nature Chemical Biology (2024)
-
Myofiber-type-dependent ‘boulder’ or ‘multitudinous pebble’ formations across distinct amylopectinoses
Acta Neuropathologica (2024)
-
Mechanisms underlying linear ubiquitination and implications in tumorigenesis and drug discovery
Cell Communication and Signaling (2023)
-
Genetic deletion and pharmacologic inhibition of E3 ubiquitin ligase HOIP impairs the propagation of myeloid leukemia
Leukemia (2023)
-
The Emerging Roles of E3 Ligases and DUBs in Neurodegenerative Diseases
Molecular Neurobiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.