Abstract
Influenza A virus-specific B lymphocytes and the antibodies they produce protect against infection1. However, the outcome of interactions between an influenza haemagglutinin-specific B cell via its receptor (BCR) and virus is unclear. Through somatic cell nuclear transfer we generated mice that harbour B cells with a BCR specific for the haemagglutinin of influenza A/WSN/33 virus (FluBI mice). Their B cells secrete an immunoglobulin gamma 2b that neutralizes infectious virus. Whereas B cells from FluBI and control mice bind equivalent amounts of virus through interaction of haemagglutinin with surface-disposed sialic acids, the A/WSN/33 virus infects only the haemagglutinin-specific B cells. Mere binding of virus is not sufficient for infection of B cells: this requires interactions of the BCR with haemagglutinin, causing both disruption of antibody secretion and FluBI B-cell death within 18 h. In mice infected with A/WSN/33, lung-resident FluBI B cells are infected by the virus, thus delaying the onset of protective antibody release into the lungs, whereas FluBI cells in the draining lymph node are not infected and proliferate. We propose that influenza targets and kills influenza-specific B cells in the lung, thus allowing the virus to gain purchase before the initiation of an effective adaptive response.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Valkenburg, S. A. et al. Immunity to seasonal and pandemic influenza A viruses. Microbes Infect. 13, 489–501 (2011)
Onodera, T. et al. Memory B cells in the lung participate in protective humoral immune responses to pulmonary influenza virus reinfection. Proc. Natl Acad. Sci. USA 109, 2485–2490 (2012)
Manicassamy, B. et al. Analysis of in vivo dynamics of influenza virus infection in mice using a GFP reporter virus. Proc. Natl Acad. Sci. USA 107, 11531–11536 (2010)
Popp, M. W., Karssemeijer, R. A. & Ploegh, H. L. Chemoenzymatic site-specific labeling of influenza glycoproteins as a tool to observe virus budding in real time. PLoS Pathog. 8, e1002604 (2012)
Popp, M. W. & Ploegh, H. L. Making and breaking peptide bonds: protein engineering using sortase. Angew. Chem. Int. Edn Engl. 50, 5024–5032 (2011)
Antos, J. M., Miller, G. M., Grotenbreg, G. M. & Ploegh, H. L. Lipid modification of proteins through sortase-catalyzed transpeptidation. J. Am. Chem. Soc. 130, 16338–16343 (2008)
Dougan, S. K. et al. IgG1+ ovalbumin-specific B-cell transnuclear mice show class switch recombination in rare allelically included B cells. Proc. Natl Acad. Sci. USA 109, 13739–13744 (2012)
Kirak, O. et al. Transnuclear mice with predefined T cell receptor specificities against Toxoplasma gondii obtained via SCNT. Science 328, 243–248 (2010)
Dougan, S. K. et al. Transnuclear TRP1-specific CD8 T cells with high or low affinity TCRs show equivalent anti-tumor activity. Cancer Immunol. Res. 1, 99–111 (2013)
Wiley, D. C. & Skehel, J. J. The structure and function of the hemagglutinin membrane glycoprotein of influenza virus. Annu. Rev. Biochem. 56, 365–394 (1987)
Stray, S. J., Cummings, R. D. & Air, G. M. Influenza virus infection of desialylated cells. Glycobiology 10, 649–658 (2000)
Thompson, C. I., Barclay, W. S., Zambon, M. C. & Pickles, R. J. Infection of human airway epithelium by human and avian strains of influenza a virus. J. Virol. 80, 8060–8068 (2006)
Pedroso de Lima, M. C. et al. Target cell membrane sialic acid modulates both binding and fusion activity of influenza virus. Biochim. Biophys. Acta 1236, 323–330 (1995)
Huang, R. T., Lichtenberg, B. & Rick, O. Involvement of annexin V in the entry of influenza viruses and role of phospholipids in infection. FEBS Lett. 392, 59–62 (1996)
Chu, V. C. & Whittaker, G. R. Influenza virus entry and infection require host cell N-linked glycoprotein. Proc. Natl Acad. Sci. USA 101, 18153–18158 (2004)
Londrigan, S. L. et al. N-linked glycosylation facilitates sialic acid-independent attachment and entry of influenza A viruses into cells expressing DC-SIGN or L-SIGN. J. Virol. 85, 2990–3000 (2011)
Eierhoff, T., Hrincius, E. R., Rescher, U., Ludwig, S. & Ehrhardt, C. The epidermal growth factor receptor (EGFR) promotes uptake of influenza A viruses (IAV) into host cells. PLoS Pathog. 6, e1001099 (2010)
Harwood, N. E. & Batista, F. D. Early events in B cell activation. Annu. Rev. Immunol. 28, 185–210 (2010)
Witte, M. D. et al. Preparation of unnatural N-to-N and C-to-C protein fusions. Proc. Natl Acad. Sci. USA 109, 11993–11998 (2012)
Moltedo, B. et al. Cutting edge: stealth influenza virus replication precedes the initiation of adaptive immunity. J. Immunol. 183, 3569–3573 (2009)
Moltedo, B., Li, W., Yount, J. S. & Moran, T. M. Unique type I interferon responses determine the functional fate of migratory lung dendritic cells during influenza virus infection. PLoS Pathog. 7, e1002345 (2011)
Joo, H. M., He, Y. & Sangster, M. Y. Broad dispersion and lung localization of virus-specific memory B cells induced by influenza pneumonia. Proc. Natl Acad. Sci. USA 105, 3485–3490 (2008)
Jones, P. D. & Ada, G. L. Influenza-specific antibody-secreting cells and B cell memory in the murine lung after immunization with wild-type, cold-adapted variant and inactivated influenza viruses. Vaccine 5, 244–248 (1987)
Popp, M. W., Antos, J. M. & Ploegh, H. L. Site-specific protein labeling via sortase-mediated transpeptidation. Curr. Protoc. Protein Sci. 15, Unit 15.13. (2009)
Boes, M. et al. T-cell engagement of dendritic cells rapidly rearranges MHC class II transport. Nature 418, 983–988 (2002)
Morgan, D. J., McLain, L. & Dimmock, N. J. Protection of three strains of mice against lethal influenza in vivo by defective interfering virus. Virus Res. 29, 179–193 (1993)
Kirak, O. et al. Transnuclear mice with pre-defined T cell receptor specificities against Toxoplasma gondii obtained via SCNT. J. Vis. Exp. 43, e2168 (2010)
Sehrawat, S. et al. CD8+ T cells from mice transnuclear for a TCR that recognizes a single H-2Kb-restricted MHV68 epitope derived from gB-ORF8 help control infection. Cell Rep. 1, 461–471 (2012)
Kishigami, S. et al. Production of cloned mice by somatic cell nuclear transfer. Nature Protocols 1, 125–138 (2006)
Simons, K., Helenius, A., Leonard, K., Sarvas, M. & Gething, M. J. Formation of protein micelles from amphiphilic membrane proteins. Proc. Natl Acad. Sci. USA 75, 5306–5310 (1978)
Acknowledgements
S.K.D. and C.G. were funded by the Cancer Research Institute. J.J.C. was funded by the Human Frontiers Science Program. F.W.A., R.J. and H.L.P. are funded by grants from the National Institutes of Health. F.W.A. is a Howard Hughes Medical Institute investigator. S.K.D. and H.L.P. are funded by the American Association for Cancer Research-Pancreatic Cancer Action Network. We are grateful to P. Wisniewski for cell sorting, to J. Jackson for mouse husbandry, to G. Bell for statistical analysis, to N. Watson for electron microscopy and to M. Witte for sortase nucleophiles.
Author information
Authors and Affiliations
Contributions
S.K.D. and J.A. contributed equally. Somatic cell nuclear transfer was performed by S.K.D.; S.K.D., J.A., R.A.K., M.W.P., M.B. and A.F.A. performed experiments. A.M.A. generated hybridomas. J.R.I. and J.J.C. generated flu-specific VHHs. C.G. sequenced the BCR loci. F.W.A. and R.J. provided advice and reagents. S.K.D., J.A. and H.L.P. designed experiments, analysed data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Flu micelles stain HA-specific B cells.
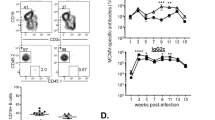
a, Schematic for preparation of glycoprotein micelles from HA–SRTAlexa 647 virus. b, Immunoprecipitation of HA–Alexa 647 with anti- Alexa 647 monoclonal antibody. Triton X100-disrupted virions were incubated with 400 μg anti-Alexa 647 overnight and HA–Alexa 647 was then recovered using protein G-Sepharose. Bound proteins were eluted with 0.1 M glycine pH 2.8. W, wash; E, elution. c, Typhoon image of the fractions obtained from a linear sucrose gradient after 20 h centrifugation (107,900g). d, Fraction 8 from the sucrose gradient was concentrated and sucrose-depleted by centrifugation over a 30 kDa filter (Amicon UltraCel). The preparation was stained with phosphotungstate and examined by transmission electron microscopy (×150,000 magnification). e, Splenocytes from mice infected with A/WSN/33 or control mice were stained with anti-CD19 and HA–Alexa 647 micelles and analysed by cytofluorometry. Plots are representative of 6 mice per group.
Extended Data Figure 2 FluBI antibody is of the IgG2b subclass.
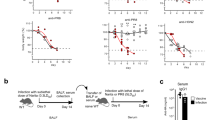
ELISA plates were coated with A/WSN/33-infected MDCK cell lysate and exposed to 1:100 diluted serum from a single C57BL/6 (wt), FluBI, FluBI;Rag2−/−, or wild-type mouse infected with A/WSN/33. Plates were washed and probed with isotype-specific secondary antibodies. Uninfected wild-type mice have flu-reactive antibodies of the IgM subclass. Flu-specific IgE was not detected in any sample. Error bars are s.d. of samples analysed in triplicate.
Extended Data Figure 3 Sequence of the VDJ and VJ segments of the FluBI antibody.
Genomic DNA was prepared from tails of FluBI mice. The heavy and light chain rearrangements were first identified by amplifying and sequencing of the segments with degenerate primers: for heavy chain: forward 5′-ARGCCTGGGRCTTCAGTGAAG-3′ and reverse 5′-AGGCTCTGAGATCCCTAGACAG-3′; for light chain: forward 5′-GGCTGCAGSTTCAGTGGCAGTGGRTCWGGRAC-3′ and reverse 5′-ATGCGACGTCAACTGATAATGAGCCCTCTCC-3′. Then the full sequences of the rearranged heavy and light chain segments were obtained using specific primers: forward 5′-TTACTGAGCACACAGGACCTC-3′ and reverse 5′-AGGCTCTGAGATCCCTAGACAG-3′; for light chain: forward 5′-CAGCCCATATTCTCCCATGT-3′ and reverse 5′-ATGCGACGTCAACTGATAATGAGCCCTCTCC-3′. Amplified products were agarose gel-purified and sequenced. Sequences were aligned to the NCBI mouse V, D and J genes using IgBlast. Sequences were deposited in GenBank (accession numbers KF419287 and KF419288).
Extended Data Figure 4 FluBI mice lack B-1a B cells, but show near-normal development of follicular B cells.
Cells were isolated from spleen, lymph node (LN, pooled mesenteric and cervical), peritoneal cavity and bone marrow of FluBI, FluBI Rag2−/− or C57BL/6 mice. Erythrocytes were lysed and cells were stained with the indicated antibodies and 7-AAD viability dye. LN plots were gated on total live cells. All other populations were gated on CD19+ live cells. Numbers indicated the percentage of cells in the indicated gates. B-1a B cells (CD5+) are absent and B-1b B cells (CD5−CD11b+) are reduced in the peritoneal cavity of FluBI and FluBI Rag2−/− mice. Plots are representative of 5 mice per group.
Extended Data Figure 5 FluBI B cells are infected by A/WSN/33.
CD40-activated OBI or FluBI B cells were incubated with A/WSN/33 virus at an MOI of 1.0 for 30 min on ice. Cells were then washed and incubated at 37 °C in RPMI (0.2% BSA). At 2 h.p.i., cells were fixed, permeabilized and stained with anti-IgG and TAMRA-conjugated anti-NP (VHH54, derived from alpaca; see Extended Data Fig. 9). a, Cells were visualized by confocal microscopy. b, Cells from a were scored as VHH54-positive or -negative. Error bars represent s.d. of positive cells counted per field (3 fields counted; ∼200 total cells were counted per group).
Extended Data Figure 6 Antibody secreted by FluBI B cells does not cross-react with other strains of influenza virus.
ELISA plates were coated with A/WSN/33 (H1N1), A/Udorn/307/1972 (H3N2) or A/Puerto Rico/8/1934 (H1N1) overnight at 4°. Plates were then washed, blocked with 10% fetal bovine serum and exposed to FluBI hybridoma supernatant or WSN-infected serum at the indicated dilutions. Bound antibody was detected using horseradish peroxidase-coupled anti-IgG2b secondary reagent.
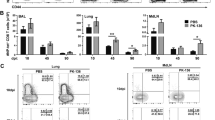
Extended Data Figure 7 FluBI B cells are not infected with A/Puerto Rico/8/1934 virus in vivo.
C57BL/6 mice were administered 5 × 106 MHCII–GFP+ FluBI B cells 2 h before intranasal infection with 2 × 105 p.f.u. per mouse of either A/WSN/33 (WSN) or A/Puerto Rico/8/1934 (PR8). Mice were euthanized 3 days post-infection, and lung resident cells were stained with anti-CD19 and TAMRA-conjugated VHH68 (anti-HA) or TAMRA-conjugated VHH52/54 (anti-NP). a, Representative plots gated on CD19+ cells. b, Quantification of flu-antigen positive cells as shown in a. n = 3. Error bars are s.d. p = 0.06 using two-sided t-test.
Extended Data Figure 8 Proliferating FluBI cells in the mediastinal lymph node are plasmablasts.
a, Mediastinal lymph node cells from day 6 post live infection mice described in Fig. 4 were analysed by confocal microscopy. GFP+ cells displayed a morphology consistent with plasmablasts. b, MSLN cells from day 6 post live infection mice described in Figure 4 were analysed by cytofluorometry. Proliferating (violet low) cells were B220low and CD138+.
Extended Data Figure 9 Alpaca-derived VHHs recognize HA and NP from A/WSN/33.
a, An alpaca was immunized with ethanol-fixed influenza virus. Phage display libraries were constructed from selectively amplified VHH-specific complementary DNA using peripheral blood lymphocytes as starting material, and panned twice against sortase labelled influenza HA–SRTbiotin virus bound to streptavidin coupled beads. VHH sequences obtained from specific binders were expressed with a sortase recognition motif to allow direct conjugation of biotin or fluorophores. b, VHH54 and VHH68 conjugated directly to agarose beads were used to precipitate lysates of A/WSN/33 infected, [35S]cysteine/methionine-labelled MDCK cells.
Extended Data Figure 10 Flu-specific VHHs can stain infected FluBI B cells.
B cells from OBI or FluBI mice were cultured for 24 h in RPMI containing anti-CD40 (1 μg ml−1) before exposure to A/WSN/33. OBI B cells, FluBI B cells and MDCK cells were incubated with A/WSN/33 at an MOI of 1.0 for 30 min on ice, washed once with PBS, and transferred to 37 °C in RPMI (0.2% BSA). At 5 h post infection, cells were washed, permeabilized, fixed and stained using TAMRA-conjugated flu-specific VHHs (1 μg in 50 μl). Infected MDCK cells were analysed in parallel as a positive control. Cells were analysed by cytofluorometry using a BD Fortessa.
Rights and permissions
About this article
Cite this article
Dougan, S., Ashour, J., Karssemeijer, R. et al. Antigen-specific B-cell receptor sensitizes B cells to infection by influenza virus. Nature 503, 406–409 (2013). https://doi.org/10.1038/nature12637
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12637
This article is cited by
-
Elucidating the characteristics of Mx1 and resistance to influenza A virus subtype H1N1 in the newly developed KWM/Hym mice
Laboratory Animal Research (2022)
-
Influenza A virus-induced apoptosis and virus propagation
Apoptosis (2020)
-
Nanobody-based sandwich reporter system for living cell sensing influenza A virus infection
Scientific Reports (2019)
-
Tumor-educated B cells selectively promote breast cancer lymph node metastasis by HSPA4-targeting IgG
Nature Medicine (2019)
-
Viral subversion of B cell responses within secondary lymphoid organs
Nature Reviews Immunology (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.