Abstract
Mutations in SHANK3 and large duplications of the region spanning SHANK3 both cause a spectrum of neuropsychiatric disorders, indicating that proper SHANK3 dosage is critical for normal brain function. However, SHANK3 overexpression per se has not been established as a cause of human disorders because 22q13 duplications involve several genes. Here we report that Shank3 transgenic mice modelling a human SHANK3 duplication exhibit manic-like behaviour and seizures consistent with synaptic excitatory/inhibitory imbalance. We also identified two patients with hyperkinetic disorders carrying the smallest SHANK3-spanning duplications reported so far. These findings indicate that SHANK3 overexpression causes a hyperkinetic neuropsychiatric disorder. To probe the mechanism underlying the phenotype, we generated a Shank3 in vivo interactome and found that Shank3 directly interacts with the Arp2/3 complex to increase F-actin levels in Shank3 transgenic mice. The mood-stabilizing drug valproate, but not lithium, rescues the manic-like behaviour of Shank3 transgenic mice raising the possibility that this hyperkinetic disorder has a unique pharmacogenetic profile.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Südhof, T. C. Neuroligins and neurexins link synaptic function to cognitive disease. Nature 455, 903–911 (2008)
Zoghbi, H. Y. Postnatal neurodevelopmental disorders: meeting at the synapse? Science 302, 826–830 (2003)
Bourgeron, T. A synaptic trek to autism. Curr. Opin. Neurobiol. 19, 231–234 (2009)
Ting, J. T., Peca, J. & Feng, G. Functional consequences of mutations in postsynaptic scaffolding proteins and relevance to psychiatric disorders. Annu. Rev. Neurosci. 35, 49–71 (2012)
Sheng, M. & Kim, E. The Shank family of scaffold proteins. J. Cell Sci. 113, 1851–1856 (2000)
Sato, D. et al. SHANK1 Deletions in Males with Autism Spectrum Disorder. Am. J. Hum. Genet. 90, 879–887 (2012)
Berkel, S. et al. Mutations in the SHANK2 synaptic scaffolding gene in autism spectrum disorder and mental retardation. Nature Genet. 42, 489–491 (2010)
Grabrucker, A. M., Schmeisser, M. J., Schoen, M. & Boeckers, T. M. Postsynaptic ProSAP/Shank scaffolds in the cross-hair of synaptopathies. Trends Cell Biol. 21, 594–603 (2011)
Durand, C. M. et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nature Genet. 39, 25–27 (2007)
Moessner, R. et al. Contribution of SHANK3 mutations to autism spectrum disorder. Am. J. Hum. Genet. 81, 1289–1297 (2007)
Gauthier, J. et al. Novel de novo SHANK3 mutation in autistic patients. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 150B, 421–424 (2009)
Gauthier, J. et al. De novo mutations in the gene encoding the synaptic scaffolding protein SHANK3 in patients ascertained for schizophrenia. Proc. Natl Acad. Sci. USA 107, 7863–7868 (2010)
Bonaglia, M. C. et al. Disruption of the ProSAP2 gene in a t(12;22)(q24.1;q13.3) is associated with the 22q13.3 deletion syndrome. Am. J. Hum. Genet. 69, 261–268 (2001)
Bonaglia, M. C. et al. Identification of a recurrent breakpoint within the SHANK3 gene in the 22q13.3 deletion syndrome. J. Med. Genet. 43, 822–828 (2006)
Bozdagi, O. et al. Haploinsufficiency of the autism-associated Shank3 gene leads to deficits in synaptic function, social interaction, and social communication. Mol. Autism 1, 15 (2010)
Peça, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011)
Wang, X. et al. Synaptic dysfunction and abnormal behaviors in mice lacking major isoforms of Shank3. Hum. Mol. Genet. 20, 3093–3108 (2011)
Failla, P. et al. Schizophrenia in a patient with subtelomeric duplication of chromosome 22q. Clin. Genet. 71, 599–601 (2007)
Shaltiel, G. et al. Evidence for the involvement of the kainate receptor subunit GluR6 (GRIK2) in mediating behavioral displays related to behavioral symptoms of mania. Mol. Psychiatry 13, 858–872 (2008)
Leibenluft, E. & Rich, B. A. Pediatric bipolar disorder. Annu. Rev. Clin. Psychol. 4, 163–187 (2008)
Martinowich, K., Schloesser, R. J. & Manji, H. K. Bipolar disorder: from genes to behavior pathways. J. Clin. Invest. 119, 726–736 (2009)
Perry, W., Minassian, A., Feifel, D. & Braff, D. L. Sensorimotor gating deficits in bipolar disorder patients with acute psychotic mania. Biol. Psychiatry 50, 418–424 (2001)
Belmaker, R. H. Bipolar disorder. N. Engl. J. Med. 351, 476–486 (2004)
McCormick, D. A. & Contreras, D. On the cellular and network bases of epileptic seizures. Annu. Rev. Physiol. 63, 815–846 (2001)
Sakai, Y. et al. Protein interactome reveals converging molecular pathways among autism disorders. Sci. Transl. Med. 3, 86ra49 (2011)
Collins, M. O. et al. Molecular characterization and comparison of the components and multiprotein complexes in the postsynaptic proteome. J. Neurochem. 97 (Suppl 1). 16–23 (2006)
Bayés, A. et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nature Neurosci. 14, 19–21 (2011)
Campellone, K. G. & Welch, M. D. A nucleator arms race: cellular control of actin assembly. Nature Rev. Mol. Cell Biol. 11, 237–251 (2010)
Proepper, C. et al. Abelson interacting protein 1 (Abi-1) is essential for dendrite morphogenesis and synapse formation. EMBO J. 26, 1397–1409 (2007)
Naisbitt, S. et al. Shank, a novel family of postsynaptic density proteins that binds to the NMDA receptor/PSD-95/GKAP complex and cortactin. Neuron 23, 569–582 (1999)
Sheng, M. & Kim, E. The postsynaptic organization of synapses. Cold Spring Harb. Perspect. Biol. (http://dx.doi.org/10.1101/cshperspect.a005678) (2011)
Giesemann, T. et al. Complex formation between the postsynaptic scaffolding protein gephyrin, profilin, and Mena: a possible link to the microfilament system. J. Neurosci. 23, 8330–8339 (2003)
Neuhoff, H. et al. The actin-binding protein profilin I is localized at synaptic sites in an activity-regulated manner. Eur. J. Neurosci. 21, 15–25 (2005)
Ackermann, M. & Matus, A. Activity-induced targeting of profilin and stabilization of dendritic spine morphology. Nature Neurosci. 6, 1194–1200 (2003)
Jope, R. S. Anti-bipolar therapy: mechanism of action of lithium. Mol. Psychiatry 4, 117–128 (1999)
Rosenberg, G. The mechanisms of action of valproate in neuropsychiatric disorders: can we see the forest for the trees? Cell. Mol. Life Sci. 64, 2090–2103 (2007)
Ramocki, M. B. & Zoghbi, H. Y. Failure of neuronal homeostasis results in common neuropsychiatric phenotypes. Nature 455, 912–918 (2008)
Toro, R. et al. Key role for gene dosage and synaptic homeostasis in autism spectrum disorders. Trends Genet. 26, 363–372 (2010)
Gitlin, M. Treatment-resistant bipolar disorder. Mol. Psychiatry 11, 227–240 (2006)
Dunner, D. L. & Fieve, R. R. Clinical factors in lithium carbonate prophylaxis failure. Arch. Gen. Psychiatry 30, 229–233 (1974)
Gould, T. D. & Manji, H. K. Glycogen synthase kinase-3: a putative molecular target for lithium mimetic drugs. Neuropsychopharmacology 30, 1223–1237 (2005)
Maglóczky, Z. & Freund, T. F. Impaired and repaired inhibitory circuits in the epileptic human hippocampus. Trends Neurosci. 28, 334–340 (2005)
Marín, O. Interneuron dysfunction in psychiatric disorders. Nature Rev. Neurosci. 13, 107–120 (2012)
Benes, F. M. et al. Regulation of the GABA cell phenotype in hippocampus of schizophrenics and bipolars. Proc. Natl Acad. Sci. USA 104, 10164–10169 (2007)
Schloesser, R. J., Martinowich, K. & Manji, H. K. Mood-stabilizing drugs: mechanisms of action. Trends Neurosci. 35, 36–46 (2012)
Warming, S., Costantino, N., Court, D. L., Jenkins, N. A. & Copeland, N. G. Simple and highly efficient BAC recombineering using galK selection. Nucleic Acids Res. 33, e36 (2005)
Choi, J. et al. Regulation of dendritic spine morphogenesis by insulin receptor substrate 53, a downstream effector of Rac1 and Cdc42 small GTPases. J. Neurosci. 25, 869–879 (2005)
Han, K. et al. Regulated RalBP1 binding to RalA and PSD-95 controls AMPA receptor endocytosis and LTD. PLoS Biol. 7, e1000187 (2009)
Chao, H. T. et al. Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 468, 263–269 (2010)
Roberson, E. D. et al. Amyloid-β/Fyn-induced synaptic, network, and cognitive impairments depend on tau levels in multiple mouse models of Alzheimer’s disease. J. Neurosci. 31, 700–711 (2011)
Lu, H., Lim, B. & Poo, M. M. Cocaine exposure in utero alters synaptic plasticity in the medial prefrontal cortex of postnatal rats. J. Neurosci. 29, 12664–12674 (2009)
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003)
Dennis, G. et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 4, P3 (2003)
Huang D. W, Sherman B. T & Lempicki R. A Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nature Protocols 4, 44–57 (2009)
Boone, P. M. et al. Detection of clinically relevant exonic copy-number changes by array CGH. Hum. Mutat. 31, 1326–1342 (2010)
Acknowledgements
We are indebted to the patients and families who participated in this study; to J. W. Belmont and N. Miller for contributing patients to this study; G. Feng for sharing Shank3B mice; G. Schuster for injection of Shank3 BAC; and C. Spencer for behavioural assays training. This project was supported by The Howard Hughes Medical Institute (H.Y.Z.), National Institutes of Health (NIH) ARRA grant (1R01NS070302) (H.Y.Z.), the Baylor Intellectual and Developmental Disabilities Research Center (P30HD024064) confocal, electrophysiology and mouse neurobehavioral cores, and the Cancer Prevention and Research Institute of Texas (CPRIT) RP110784. J.L.H. was supported by an Early Career Award from the Thrasher Research Fund, NIH 2T32NS043124 and the Ting Tsung and Wei Fong Chao Foundation; C.P.S. was supported by the Joan and Stanford Alexander family, the Ting Tsung and Wei Fong Chao Foundation and the Doris Duke Clinical Scientist Development Award.
Author information
Authors and Affiliations
Contributions
K.H., J.L.H., H.L., H.C., J.T., H.-C.L. and H.Y.Z. designed the experiments. K.H., J.L.H., H.L., H.C., J.T., Z.W., S.H. and H.S. performed the research. K.H., J.L.H., C.P.S., H.L., H.C., H.K., J.T., Z.W., S.H., S.W.C., P.Y., A.M.B., A.P., H.-C.L. and H.Y.Z. collected, analysed and interpreted the data. K.H., J.L.H., C.P.S., H.L., H.K., J.T., H.-C.L. and H.Y.Z. wrote and edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Characterization of expression patterns of EGFP–Shank3 and other synaptic proteins in Shank3 transgenic mice.
a, Description of the Shank3 BAC used for transgenic mice generation (image was modified from UCSC genome browser). b, Western blot images show expression of EGFP–Shank3 in the synaptic fraction of Shank3 transgenic brain lysates. EGFP antibody recognized EGFP–Shank3 (∼200 kDa) specifically in synaptic fractions of Shank3 transgenic mice. Asterisk indicates non-specific bands detected by the EGFP antibody. Shank3 antibody detected endogenous Shank3 plus EGFP–Shank3 (arrow) in the transgenic samples. S2, soluble fraction; P2, crude synaptosomal fraction; PSD I, synaptic fraction after one time Triton X-100 washout. c, The brain regional expression pattern of EGFP–Shank3 is similar to that of endogenous Shank3. Cb, cerebellum; Ct, cortex; Hp, hippocampus; St, striatum; Th, thalamus. d, The brain developmental expression pattern of EGFP–Shank3 is similar to that of endogenous Shank3. e, The fold changes of Shank3 proteins in 3-week-old transgenic mice (n = 4) are similar to those of 6-week-old mice (Fig. 1d). f, Female transgenic mice (n = 4) show similar fold changes of Shank3 proteins to male transgenic mice (Fig. 1d). g, h, Expression levels of excitatory (g) and inhibitory (h) synaptic proteins are not significantly altered, except Shank3, in the synaptosomal fraction of 8-week-old transgenic hippocampus and striatum (n = 6). All data are presented as mean ± s.e.m. P values (*P < 0.05) were derived from unpaired, two-tailed Student’s t-test.
Extended Data Figure 2 Behavioural characterization of Shank3 transgenic mice.
a, Transgenic mice showed increased locomotor speed during the 30 min open-field assay. b, Female transgenic mice showed increased home-cage activity. c, Transgenic mice have increased body weight compared to wild-type mice measured at 9-weeks-old (male wild type: 28.1 ± 0.9, male transgenic: 30.6 ± 0.7, female wild type: 20.1 ± 0.3, female transgenic: 21.4 ± 0.4, n = 10–13; *P < 0.05; unpaired two-tailed Student’s t-test) and 15-weeks-old (male wild type: 32.4 ± 1.2, male transgenic: 37.0 ± 1.3, female wild type: 23.6 ± 0.4, female transgenic: 26.6 ± 0.9, n = 10–13; *P < 0.05). d, Increased food intake by transgenic mice. Food intake was measured from single-caged female transgenic or wild-type mice from 6- to 10-weeks-old. At 7-weeks-old (wild type: 3.51 ± 0.05, transgenic: 3.69 ± 0.06, n = 9–11; *P < 0.05; unpaired two-tailed Student’s t-test) and 9-weeks-old (wild type: 3.61 ± 0.07, transgenic: 3.86 ± 0.05, n = 9–11; *P < 0.05), food intake by transgenic mice was significantly higher than wild-type mice. e, f, In the 3-chamber assay, male (e) and female (f) transgenic mice did not show significant preference for novel mice to novel objects. g, Transgenic mice spent significantly less time in close interaction with novel social partners. h, i, Stereotypy time (h) and activity count (i) during the 30 min open-field test were normal in transgenic mice. j, Time spent in grooming during 10 min monitoring was normal in transgenic mice. k, Separation-induced ultrasonic vocalization was measured from wild-type and transgenic mice of postnatal day 6 to 13. Transgenic mice made less calls than wild type at postnatal day 13. l, At postnatal day 13, transgenic mice express more Shank3 proteins than wild-type mice. All data are presented as mean ± s.e.m. *P < 0.05, **P < 0.01, ***P < 0.001. The summary of statistical analyses for behavioural assays is provided in Supplementary Table 1.
Extended Data Figure 3 Behavioural phenotypes of Shank3 transgenic mice were rescued by crossing with Shank3B+/− mice.
a, Schematic diagram shows possible genotypic combinations and their Shank3 expression levels (arrow) from the crossing between Shank3 transgenic mice and Shank3B+/− mice. b, Quantification of the levels of Shank3 proteins in each genotype (n = 4). The fold changes were compared to the wild-type (Shank3B+/+) controls. Crossing of Shank3 transgenic mice with Shank3B+/− mice significantly decreased the levels of α isoform and total Shank3 protein. c, d, Locomotor activities of male (c) and female (d) Shank3 transgenic mice were rescued by crossing with Shank3B+/− mice. e, f, Immobile time in tail-suspension test of male (e) and female (f) Shank3 transgenic mice were rescued by crossing with Shank3B+/− mice. g, Prepulse inhibition (by 78 dB) of female Shank3 transgenic mice was rescued by crossing with Shank3B+/− mice. All data are presented as mean ± s.e.m. *P < 0.05; ***P < 0.001.
Extended Data Figure 4 Basal synaptic transmission and NMDA receptor-dependent synaptic plasticity are normal at the hippocampal Schaffer collateral-CA1 synapses of Shank3 transgenic mice.
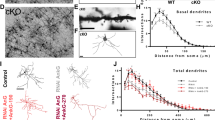
a, Images of cresylviolet staining show normal cytoarchitecture of transgenic brain. b, Normal paired-pulse facilitation ratio (PPR) of evoked EPSCs (eEPSCs) at transgenic Schaffer collateral-CA1 synapses. c, d, Normal synaptic input-output (I-O) relationship at transgenic Schaffer collateral-CA1 synapses. I-O curve (c) and cumulative curve of I-O slope (d) are shown. Red lines in c represent the fitting curves. e, Left, sample traces of action potentials triggered by 200 pA current injection in hippocampal CA1 region of wild-type and transgenic mice. Middle and right, unaltered number of action potentials trigged by the injection of current at different level (middle) and amplitude of the first action potential triggered by 800-ms-long pulse of 200 pA current (right) show normal intrinsic excitability of hippocampal CA1 pyramidal neurons in transgenic mice. f, Amplitude, frequency and decay of mEPSC are not altered in CA1 pyramidal neurons of transgenic mice. g, h, NMDA receptor-dependent long-term potentiation (g) and long-term depression (h) at Schaffer collateral-CA1 pyramidal synapses are normal in transgenic neurons.
Extended Data Figure 5 Generation and characterization of Shank3 in vivo interactome.
a, Isolation of EGFP–Shank3 protein and its interactors from synaptosomal fraction of transgenic mice. b, Venn diagrams show overlaps among the protein lists from Shank3 in vivo immunoprecipitation, Shank3 yeast two-hybrid screening, and either published mouse PSP (postsynaptic proteome) or human PSD. c, Shank3 interactome network. d, Average path length of Shank3 interactome (3.12, red line) is significantly (P < 0.0001) shorter than that of mouse PSP interactome (mean = 3.91, black line), supporting strong connectivity of Shank3 interactome. e, GO and KEGG pathway analysis of Shank3 interactome reveal enrichment of actin cytoskeleton-related function/pathway. The full result of analysis is in Supplementary Table 5. f, Confirmation of the in vivo interactions between Shank3 and actin-related proteins. g, In cultured hippocampal neurons, ARPC2 proteins are co-localized with F-actin (upper panel) and EGFP–Shank3 (lower panel).
Extended Data Figure 6 Shank3 siRNA reversed increased F-actin levels and ARPC2 cluster size in cultured hippocampal pyramidal neurons from Shank3 transgenic mice.
a, Validation of the two siRNAs against Shank3. HEK293T cells were transfected with HA-Shank3 plus control or Shank3 siRNA. EGFP plasmid was co-transfected as an internal control. After 48 h, expression levels of HA–Shank3 were measured by western blot and quantified (n = 3). b, siRNA targeting Shank3 (si-Shank3) reversed the increased F-actin levels of Shank3 transgenic neurons. Image for si-Shank3 #2 is not shown. c, si-Shank3 reversed the increased ARPC2 cluster size of Shank3 transgenic neurons. All data are presented as mean ± s.e.m. from 20–30 neurons per condition. P values (*P < 0.05, **P < 0.01) were derived from one-way ANOVA with post hoc Tukey’s multiple comparison.
Extended Data Figure 7 Normal excitatory and inhibitory synapse numbers of GAD-6-positive inhibitory neurons of Shank3 transgenic mice.
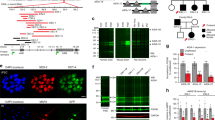
a, Expression of EGFP–Shank3 in GAD-6-positive inhibitory neurons of Shank3 transgenic mice. The yellow circles of dotted line and solid line indicate neuronal cell bodies of a GAD-6-negative excitatory neuron and a GAD-6-positive inhibitory neuron, respectively. The box 1 shows dendritic segment of the excitatory neuron and the box 2 shows that of the inhibitory neuron. b, Quantification of dendritic EGFP–Shank3 intensity in excitatory and inhibitory transgenic neurons. Inhibitory neurons express less EGFP–Shank3 (excitatory neurons: 1.00 ± 0.06, inhibitory neurons: 0.74 ± 0.09, n = 10; *P < 0.05; unpaired two-tailed Student’s t-test). c, Normal excitatory synapse number of GAD-6-positive inhibitory neurons of Shank3 transgenic mice. d, Normal inhibitory synapse number of GAD-6-positive inhibitory neurons of Shank3 transgenic mice. e, Quantification of (c) (wild type: 1.00 ± 0.10, transgenic: 0.99 ± 0.07, n = 18; P > 0.05; unpaired two-tailed Student’s t-test) and (d) (wild type: 1.00 ± 0.12, transgenic: 0.95 ± 0.09, n = 20; P > 0.05; unpaired two-tailed Student’s t-test).
Extended Data Figure 8 Shank3 interactome is connected with gephyrin-interacting actin-related proteins, Mena, profilin1 and profilin2.
Based on a literature search, we selected candidate actin-related proteins (Mena/VASP, profilin1 and profilin2) that directly interact with inhibitory postsynaptic protein gephyrin. To understand potential interactions among these proteins and the Shank3 interactome, we generated interaction network using the information from our yeast two-hybrid screening and IRefIndex PPI database (http://irefindex.org). The interaction network shows that Mena/VASP, profilin1 and profilin2 are connected with Shank3 interactome mainly through actin-related proteins. Note that one of the gephyrin-interacting proteins, profilin2, was also identified by our Shank3 in vivo immunoprecipitation (Supplementary Table 4).
Extended Data Figure 9 Mania-like behaviours of Shank3 transgenic mice are resistant to lithium treatment.
a, Basal and amphetamine-induced locomotor activities of female transgenic mice were not affected by lithium treatment. b–e, Abnormal acoustic startle response (b, d) and PPI (c, e) of male and female transgenic mice were not rescued by lithium treatment. f, g, Immobile time of male (f) and female (g) transgenic mice in tail-suspension test was not rescued by lithium treatment. Wild-type female mice showed decreased immobile time upon lithium treatment, which is an expected response to high dose lithium treatment in wild-type mice. All data are presented as mean ± s.e.m. *P < 0.05; ***P < 0.001.
Extended Data Figure 10 Valproate treatment does not affect synaptic protein and F-actin levels in neurons of Shank3 transgenic mice.
a, One hour after final valproate injection (200 mg per kg), synaptosomal fraction was prepared from brains, and indicated proteins were detected by western blotting. Neither wild-type nor transgenic mice showed significant change in the levels of synaptic proteins by valproate treatment (n = 7). b, Representative confocal images of Shank3 transgenic cultured hippocampal pyramidal neurons (DIV 14) treated with different concentrations of valproate (0, 0.01, 0.1 and 1 mM) for 30 or 60 min. c, Quantification of (b). There was no significant change in F-actin levels by valproate treatment (n = 16 per condition). All data are presented as mean ± s.e.m.
Supplementary information
Supplementary Data
This file contains Supplementary data and Supplementary Tables 4-5. (PDF 406 kb)
Supplementary Data
This file contains Supplementary Tables 1-3. (XLSX 35 kb)
Spontaneous seizure of Shank3 transgenic mice
This video shows spontaneous seizure from an 8-week old female Shank3 transgenic mouse in the home-cage. (MP4 5938 kb)
Rights and permissions
About this article
Cite this article
Han, K., Holder Jr, J., Schaaf, C. et al. SHANK3 overexpression causes manic-like behaviour with unique pharmacogenetic properties. Nature 503, 72–77 (2013). https://doi.org/10.1038/nature12630
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12630
This article is cited by
-
Lithium rescues dendritic abnormalities in Ank3 deficiency models through the synergic effects of GSK3β and cyclic AMP signaling pathways
Neuropsychopharmacology (2023)
-
HSPA12A controls cerebral lactate homeostasis to maintain hippocampal neurogenesis and mood stabilization
Translational Psychiatry (2023)
-
Disrupted extracellular matrix and cell cycle genes in autism-associated Shank3 deficiency are targeted by lithium
Molecular Psychiatry (2023)
-
Mutations affecting the N-terminal domains of SHANK3 point to different pathomechanisms in neurodevelopmental disorders
Scientific Reports (2022)
-
SHANK family on stem cell fate and development
Cell Death & Disease (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.