Abstract
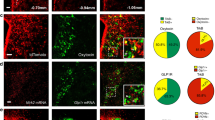
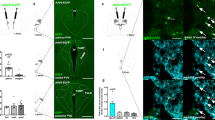
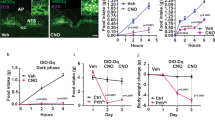
Appetite suppression occurs after a meal and in conditions when it is unfavourable to eat, such as during illness or exposure to toxins. A brain region proposed to play a role in appetite suppression is the parabrachial nucleus1,2,3, a heterogeneous population of neurons surrounding the superior cerebellar peduncle in the brainstem. The parabrachial nucleus is thought to mediate the suppression of appetite induced by the anorectic hormones amylin and cholecystokinin2, as well as by lithium chloride and lipopolysaccharide, compounds that mimic the effects of toxic foods and bacterial infections, respectively4,5,6. Hyperactivity of the parabrachial nucleus is also thought to cause starvation after ablation of orexigenic agouti-related peptide neurons in adult mice1,7. However, the identities of neurons in the parabrachial nucleus that regulate feeding are unknown, as are the functionally relevant downstream projections. Here we identify calcitonin gene-related peptide-expressing neurons in the outer external lateral subdivision of the parabrachial nucleus that project to the laterocapsular division of the central nucleus of the amygdala as forming a functionally important circuit for suppressing appetite. Using genetically encoded anatomical, optogenetic8 and pharmacogenetic9 tools, we demonstrate that activation of these neurons projecting to the central nucleus of the amygdala suppresses appetite. In contrast, inhibition of these neurons increases food intake in circumstances when mice do not normally eat and prevents starvation in adult mice whose agouti-related peptide neurons are ablated. Taken together, our data demonstrate that this neural circuit from the parabrachial nucleus to the central nucleus of the amygdala mediates appetite suppression in conditions when it is unfavourable to eat. This neural circuit may provide targets for therapeutic intervention to overcome or promote appetite.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wu, Q., Clark, M. S. & Palmiter, R. D. Deciphering a neuronal circuit that mediates appetite. Nature 483, 594–597 (2012)
Becskei, C., Grabler, V., Edwards, G. L., Riediger, T. & Lutz, T. A. Lesion of the lateral parabrachial nucleus attenuates the anorectic effect of peripheral amylin and CCK. Brain Res. 1162, 76–84 (2007)
DiPatrizio, N. V. & Simansky, K. J. Activating parabrachial cannabinoid CB1 receptors selectively stimulates feeding of palatable foods in rats. J. Neurosci. 28, 9702–9709 (2008)
Yamamoto, T. et al. C-fos expression in the rat brain after intraperitoneal injection of lithium chloride. Neuroreport 3, 1049–1052 (1992)
Elmquist, J. K., Scammell, T. E., Jacobson, C. D. & Saper, C. B. Distribution of Fos-like immunoreactivity in the rat brain following intravenous lipopolysaccharide administration. J. Comp. Neurol. 371, 85–103 (1996)
Paues, J., Mackerlova, L. & Blomqvist, A. Expression of melanocortin-4 receptor by rat parabrachial neurons responsive to immune and aversive stimuli. Neuroscience 141, 287–297 (2006)
Wu, Q., Boyle, M. P. & Palmiter, R. D. Loss of GABAergic signaling by AgRP neurons to the parabrachial nucleus leads to starvation. Cell 137, 1225–1234 (2009)
Yizhar, O., Fenno, L. E., Davidson, T. J., Mogri, M. & Deisseroth, K. Optogenetics in neural systems. Neuron 71, 9–34 (2011)
Armbruster, B. N., Li, X., Pausch, M. H., Herlitze, S. & Roth, B. L. Evolving the lock to fit the key to create a family of G protein-coupled receptors potently activated by an inert ligand. Proc. Natl Acad. Sci. USA 104, 5163–5168 (2007)
Rosen, A. M., Victor, J. D. & Di Lorenzo, P. M. Temporal coding of taste in the parabrachial nucleus of the pons of the rat. J. Neurophysiol. 105, 1889–1896 (2011)
Tokita, K. & Boughter, J. D., Jr Sweet-bitter and umami-bitter taste interactions in single parabrachial neurons in C57BL/6J mice. J. Neurophysiol. 108, 2179–2190 (2012)
Geerling, J. C. & Loewy, A. D. Sodium deprivation and salt intake activate separate neuronal subpopulations in the nucleus of the solitary tract and the parabrachial complex. J. Comp. Neurol. 504, 379–403 (2007)
Geerling, J. C. et al. FoxP2 expression defines dorsolateral pontine neurons activated by sodium deprivation. Brain Res. 1375, 19–27 (2011)
Chamberlin, N. L. & Saper, C. B. Topographic organization of respiratory responses to glutamate microstimulation of the parabrachial nucleus in the rat. J. Neurosci. 14, 6500–6510 (1994)
Hermanson, O. & Blomqvist, A. Subnuclear localization of FOS-like immunoreactivity in the rat parabrachial nucleus after nociceptive stimulation. J. Comp. Neurol. 368, 45–56 (1996)
Richard, S., Engblom, D., Paues, J., Mackerlova, L. & Blomqvist, A. Activation of the parabrachio-amygdaloid pathway by immune challenge or spinal nociceptive input: a quantitative study in the rat using Fos immunohistochemistry and retrograde tract tracing. J. Comp. Neurol. 481, 210–219 (2005)
Nakamura, K. & Morrison, S. F. A thermosensory pathway that controls body temperature. Nature Neurosci. 11, 62–71 (2008)
Nakamura, K. & Morrison, S. F. A thermosensory pathway mediating heat-defense responses. Proc. Natl Acad. Sci. USA 107, 8848–8853 (2010)
Luquet, S., Perez, F. A., Hnasko, T. S. & Palmiter, R. D. NPY/AgRP neurons are essential for feeding in adult mice but can be ablated in neonates. Science 310, 683–685 (2005)
Ng, L. et al. An anatomic gene expression atlas of the adult mouse brain. Nature Neurosci. 12, 356–362 (2009)
Jacobs, J. W. et al. Calcitonin messenger RNA encodes multiple polypeptides in a single precursor. Science 213, 457–459 (1981)
Paues, J., Engblom, D., Mackerlova, L., Ericsson-Dahlstrand, A. & Blomqvist, A. Feeding-related immune responsive brain stem neurons: association with CGRP. Neuroreport 12, 2399–2403 (2001)
Krashes, M. J. et al. Rapid, reversible activation of AgRP neurons drives feeding behavior in mice. J. Clin. Invest. 121, 1424–1428 (2011)
D’Hanis, W., Linke, R. & Yilmazer-Hanke, D. M. Topography of thalamic and parabrachial calcitonin gene-related peptide (CGRP) immunoreactive neurons projecting to subnuclei of the amygdala and extended amygdala. J. Comp. Neurol. 505, 268–291 (2007)
Schwaber, J. S., Sternini, C., Brecha, N. C., Rogers, W. T. & Card, J. P. Neurons containing calcitonin gene-related peptide in the parabrachial nucleus project to the central nucleus of the amygdala. J. Comp. Neurol. 270, 416–426 (1988)
Elmquist, J. K., Coppari, R., Balthasar, N., Ichinose, M. & Lowell, B. B. Identifying hypothalamic pathways controlling food intake, body weight, and glucose homeostasis. J. Comp. Neurol. 493, 63–71 (2005)
Atasoy, D., Betley, J. N., Su, H. H. & Sternson, S. M. Deconstruction of a neural circuit for hunger. Nature 488, 172–177 (2012)
Soudais, C., Laplace-Builhe, C., Kissa, K. & Kremer, E. J. Preferential transduction of neurons by canine adenovirus vectors and their efficient retrograde transport in vivo. FASEB J. 15, 2283–2285 (2001)
Gao, Q. & Horvath, T. L. Neurobiology of feeding and energy expenditure. Annu. Rev. Neurosci. 30, 367–398 (2007)
Neugebauer, V., Li, W., Bird, G. C. & Han, J. S. The amygdala and persistent pain. Neuroscientist 10, 221–234 (2004)
Kremer, E. J., Boutin, S., Chillon, M. & Danos, O. Canine adenovirus vectors: an alternative for adenovirus-mediated gene transfer. J. Virol. 74, 505–512 (2000)
Paxinos, G. & Franklin, K. B. J. The Mouse Brain in Stereotaxic Coordinates 4th edn (Elsevier, 2013)
Aravanis, A. et al. An optical neural interface: in vivo control of rodent motor cortex with integrated fiberoptic and optogenetic technology. J. Neural Eng. 4, S143–S156 (2007)
Lenth, R. V. Some practical guidelines for effective sample size determination. Am. Stat. 55, 187–193 (2001)
Acknowledgements
We thank B. Roth for hM3Dq–mCherry and hM4Di–mCherry constructs, and K. Deisseroth for mCherry and ChR2–mCherry constructs. E. Allen, J. Resnick, M. Soleiman and S. Padilla assisted with histology, E. Allen and A. Rainwater assisted with animal husbandry, and J. Shulkin provided suggestions and advice. We thank members of the Palmiter and Zweifel laboratories for feedback on the manuscript. M.E.C. is financed by a fellowship from the Hilda and Preston Davis Foundation. L.S.Z. is financed by a grant from the National Institutes of Health (R01MH094536). R.D.P is supported in part by grants from the National Institutes of Health (R01DA024908) and the Klarman Family Foundation.
Author information
Authors and Affiliations
Contributions
M.E.C. and R.D.P. conceived and designed the study. M.E.C. performed and analysed histological and behavioural experiments, M.E.S. performed electrophysiology experiments and R.D.P. generated CalcaCre knock-in mice. L.S.Z. and R.D.P. provided equipment, reagents and expertise. M.E.C. wrote the manuscript in collaboration with the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-17 and Supplementary Statistical Analyses. (PDF 8023 kb)
Stimulation of PBelo CGRP neurons with ChR2 during consumption of palatable food
This video is representative of data from a single trial in Figure 2b. Left (near) lickometer port contains palatable liquid diet; right (far) lickometer port contains water. Video filmed 5 minutes after start of the active period. (MOV 3696 kb)
Rights and permissions
About this article
Cite this article
Carter, M., Soden, M., Zweifel, L. et al. Genetic identification of a neural circuit that suppresses appetite. Nature 503, 111–114 (2013). https://doi.org/10.1038/nature12596
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12596
This article is cited by
-
Parabrachial tachykinin1-expressing neurons involved in state-dependent breathing control
Nature Communications (2023)
-
Electroacupuncture improves swallowing function in a post-stroke dysphagia mouse model by activating the motor cortex inputs to the nucleus tractus solitarii through the parabrachial nuclei
Nature Communications (2023)
-
Advanced neurobiological tools to interrogate metabolism
Nature Reviews Endocrinology (2023)
-
Intestinal gluconeogenesis: metabolic benefits make sense in the light of evolution
Nature Reviews Gastroenterology & Hepatology (2023)
-
Novel Insights into the Physiology of Nutrient Sensing and Gut-Brain Communication in Surgical and Experimental Obesity Therapy
Obesity Surgery (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.