Abstract
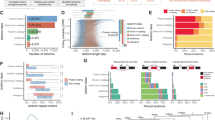
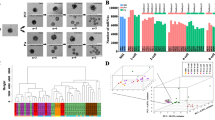
Mammalian pre-implantation development is a complex process involving dramatic changes in the transcriptional architecture1,2,3,4. We report here a comprehensive analysis of transcriptome dynamics from oocyte to morula in both human and mouse embryos, using single-cell RNA sequencing. Based on single-nucleotide variants in human blastomere messenger RNAs and paternal-specific single-nucleotide polymorphisms, we identify novel stage-specific monoallelic expression patterns for a significant portion of polymorphic gene transcripts (25 to 53%). By weighted gene co-expression network analysis5,6, we find that each developmental stage can be delineated concisely by a small number of functional modules of co-expressed genes. This result indicates a sequential order of transcriptional changes in pathways of cell cycle, gene regulation, translation and metabolism, acting in a step-wise fashion from cleavage to morula. Cross-species comparisons with mouse pre-implantation embryos reveal that the majority of human stage-specific modules (7 out of 9) are notably preserved, but developmental specificity and timing differ between human and mouse. Furthermore, we identify conserved key members (or hub genes) of the human and mouse networks. These genes represent novel candidates that are likely to be key in driving mammalian pre-implantation development. Together, the results provide a valuable resource to dissect gene regulatory mechanisms underlying progressive development of early mammalian embryos.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
29 August 2013
Ref. 26 and the associated sentence in the text were corrected; two reference citations in the online Methods were corrected.
References
Niakan, K. K., Han, J., Pedersen, R. A., Simon, C. & Pera, R. A. Human pre-implantation embryo development. Development 139, 829–841 (2012)
Hamatani, T., Carter, M. G., Sharov, A. A. & Ko, M. S. Dynamics of global gene expression changes during mouse preimplantation development. Dev. Cell 6, 117–131 (2004)
Wang, Q. T. et al. A genome-wide study of gene activity reveals developmental signaling pathways in the preimplantation mouse embryo. Dev. Cell 6, 133–144 (2004)
Zeng, F., Baldwin, D. A. & Schultz, R. M. Transcript profiling during preimplantation mouse development. Dev. Biol. 272, 483–496 (2004)
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9, 559 (2008)
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol.. http://dx.doi.org/10.2202/1544-6115.1128 (2005)
Walser, C. B. & Lipshitz, H. D. Transcript clearance during the maternal-to-zygotic transition. Curr. Opin. Genet. Dev. 21, 431–443 (2011)
Schier, A. F. The maternal-zygotic transition: death and birth of RNAs. Science 316, 406–407 (2007)
Schultz, R. M. The molecular foundations of the maternal to zygotic transition in the preimplantation embryo. Hum. Reprod. Update 8, 323–331 (2002)
Vassena, R. et al. Waves of early transcriptional activation and pluripotency program initiation during human preimplantation development. Development 138, 3699–3709 (2011)
Xie, D. et al. Rewirable gene regulatory networks in the preimplantation embryonic development of three mammalian species. Genome Res. 20, 804–815 (2010)
Zhang, P. et al. Transcriptome profiling of human pre-implantation development. PLoS ONE 4, e7844 (2009)
Dobson, A. T. et al. The unique transcriptome through day 3 of human preimplantation development. Hum. Mol. Genet. 13, 1461–1470 (2004)
Tang, F. et al. RNA-Seq analysis to capture the transcriptome landscape of a single cell. Nature Protocols 5, 516–535 (2010)
Islam, S. et al. Characterization of the single-cell transcriptional landscape by highly multiplex RNA-seq. Genome Res. 21, 1160–1167 (2011)
Ramskold, D. et al. Full-length mRNA-seq from single-cell levels of RNA and individual circulating tumor cells. Nature Biotech. 30, 777–782 (2012)
Hashimshony, T., Wagner, F., Sher, N. & Yanai, I. CEL-Seq: single-cell RNA-seq by multiplexed linear amplification. Cell Rep. 2, 666–673 (2012)
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010)
Li, Y. et al. Resequencing of 200 human exomes identifies an excess of low-frequency non-synonymous coding variants. Nature Genet. 42, 969–972 (2010)
Gabriel, S. B. et al. The structure of haplotype blocks in the human genome. Science 296, 2225–2229 (2002)
Altshuler, D. M. et al. Integrating common and rare genetic variation in diverse human populations. Nature 467, 52–58 (2010)
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nature Protocols 4, 1073–1081 (2009)
Langfelder, P., Luo, R., Oldham, M. C. & Horvath, S. Is my network module preserved and reproducible? PLOS Comput. Biol. 7, e1001057 (2011)
Tang, F. et al. Deterministic and stochastic allele specific gene expression in single mouse blastomeres. PLoS ONE 6, e21208 (2011)
Hu, J. et al. Novel importin-α family member Kpna7 is required for normal fertility and fecundity in the mouse. J. Biol. Chem. 285, 33113–33122 (2010)
Dannenberg, J. H. et al. mSin3A corepressor regulates diverse transcriptional networks governing normal and neoplastic growth and survival. Genes Dev. 19, 1581–1595 (2005)
Miller, J. A., Horvath, S. & Geschwind, D. H. Divergence of human and mouse brain transcriptome highlights Alzheimer disease pathways. Proc. Natl Acad. Sci. USA 107, 12698–12703 (2010)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature Methods 5, 621–628 (2008)
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010)
Yip, A. M. & Horvath, S. Gene network interconnectedness and the generalized topological overlap measure. BMC Bioinformatics 8, 22 (2007)
Langfelder, P., Zhang, B. & Horvath, S. Defining clusters from a hierarchical cluster tree: the Dynamic Tree Cut package for R. Bioinformatics 24, 719–720 (2008)
Eppig, J. T., Blake, J. A., Bult, C. J., Kadin, J. A. & Richardson, J. E. The Mouse Genome Database (MGD): comprehensive resource for genetics and genomics of the laboratory mouse. Nucleic Acids Res. 40, D881–D886 (2012)
Horvath, S. & Dong, J. Geometric interpretation of gene coexpression network analysis. PLOS Comput. Biol. 4, e1000117 (2008)
Hu, Z., Mellor, J., Wu, J. & DeLisi, C. VisANT: an online visualization and analysis tool for biological interaction data. BMC Bioinformatics 5, 17 (2004)
Huang, D., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID Bioinformatics Resources. Nature Protoc. 4, 44–57 (2009)
Acknowledgements
We thank many of our colleagues for invaluable discussions and comments on our study, in particular, P. Pajukanta at UCLA for very helpful suggestions on SNP analyses and D. Geschwind for critically reading the manuscript. This work was supported by 973 Grant Programs 2012CB966300, 2011CB966204 and 2011CB965102 from the Ministry of Science and Technology in China; the International Science and Technology Cooperation Program of China (no. 2011DFB30010); and the National Natural Science Foundation of China (81271258).
Author information
Authors and Affiliations
Contributions
Z.X., K.H., J.L. and G.F. designed the study. Z.X., L.C., C.J., Y.F., Z.L., Q.Z., L.C. and Y.E.S. carried out experiments or contributed critical reagents and protocols. K.H., C.C. and S.H. analysed the data and performed statistical analyses. K.H. and G.F. wrote the manuscript in discussion with all the authors. All the authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text, Supplementary Figures 1-9 and Supplementary References. (PDF 884 kb)
Supplementary Table 1
This file contains single-cell RNA-Seq statistics. (XLS 20 kb)
Supplementary Table 2
This file contains inferred Haplotype blocks as described in the methods. (XLS 52 kb)
Supplementary Table 3
This file contains a summary of hub gene data sets. (XLS 56 kb)
Rights and permissions
About this article
Cite this article
Xue, Z., Huang, K., Cai, C. et al. Genetic programs in human and mouse early embryos revealed by single-cell RNA sequencing. Nature 500, 593–597 (2013). https://doi.org/10.1038/nature12364
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12364
This article is cited by
-
Interaction network of human early embryonic transcription factors
EMBO Reports (2024)
-
Self-renewing human naïve pluripotent stem cells dedifferentiate in 3D culture and form blastoids spontaneously
Nature Communications (2024)
-
KLF4 facilitates chromatin accessibility remodeling in porcine early embryos
Science China Life Sciences (2024)
-
Sperm-borne microRNA-34c regulates maternal mRNA degradation and preimplantation embryonic development in mice
Reproductive Biology and Endocrinology (2023)
-
multiWGCNA: an R package for deep mining gene co-expression networks in multi-trait expression data
BMC Bioinformatics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.