Abstract
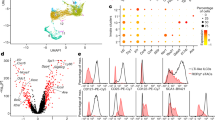
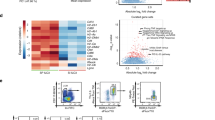
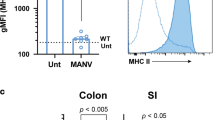
Innate lymphoid cells (ILCs) are a recently characterized family of immune cells that have critical roles in cytokine-mediated regulation of intestinal epithelial cell barrier integrity1,2,3,4,5,6,7,8,9,10. Alterations in ILC responses are associated with multiple chronic human diseases, including inflammatory bowel disease, implicating a role for ILCs in disease pathogenesis3,8,11,12,13. Owing to an inability to target ILCs selectively, experimental studies assessing ILC function have predominantly used mice lacking adaptive immune cells1,2,3,4,5,6,7,8,9,10. However, in lymphocyte-sufficient hosts ILCs are vastly outnumbered by CD4+ T cells, which express similar profiles of effector cytokines. Therefore, the function of ILCs in the presence of adaptive immunity and their potential to influence adaptive immune cell responses remain unknown. To test this, we used genetic or antibody-mediated depletion strategies to target murine ILCs in the presence of an adaptive immune system. We show that loss of retinoic-acid-receptor-related orphan receptor-γt-positive (RORγt+) ILCs was associated with dysregulated adaptive immune cell responses against commensal bacteria and low-grade systemic inflammation. Remarkably, ILC-mediated regulation of adaptive immune cells occurred independently of interleukin (IL)-17A, IL-22 or IL-23. Genome-wide transcriptional profiling and functional analyses revealed that RORγt+ ILCs express major histocompatibility complex class II (MHCII) and can process and present antigen. However, rather than inducing T-cell proliferation, ILCs acted to limit commensal bacteria-specific CD4+ T-cell responses. Consistent with this, selective deletion of MHCII in murine RORγt+ ILCs resulted in dysregulated commensal bacteria-dependent CD4+ T-cell responses that promoted spontaneous intestinal inflammation. These data identify that ILCs maintain intestinal homeostasis through MHCII-dependent interactions with CD4+ T cells that limit pathological adaptive immune cell responses to commensal bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Spits, H. & Cupedo, T. Innate lymphoid cells: emerging insights in development, lineage relationships, and function. Annu. Rev. Immunol. 30, 647–675 (2012)
Buonocore, S. et al. Innate lymphoid cells drive interleukin-23-dependent innate intestinal pathology. Nature 464, 1371–1375 (2010)
Sonnenberg, G. F. et al. Innate lymphoid cells promote anatomical containment of lymphoid-resident commensal bacteria. Science 336, 1321–1325 (2012)
Cella, M. et al. A human natural killer cell subset provides an innate source of IL-22 for mucosal immunity. Nature 457, 722–725 (2009)
Sawa, S. et al. RORγt+ innate lymphoid cells regulate intestinal homeostasis by integrating negative signals from the symbiotic microbiota. Nature Immunol. 12, 320–326 (2011)
Lochner, M. et al. Microbiota-induced tertiary lymphoid tissues aggravate inflammatory disease in the absence of RORγt and LTi cells. J. Exp. Med. 208, 125–134 (2011)
Sonnenberg, G. F., Monticelli, L. A., Elloso, M. M., Fouser, L. A. & Artis, D. CD4(+) lymphoid tissue-inducer cells promote innate immunity in the gut. Immunity 34, 122–134 (2011)
Sonnenberg, G. F. & Artis, D. Innate lymphoid cell interactions with microbiota: implications for intestinal health and disease. Immunity 37, 601–610 (2012)
Spits, H. et al. Innate lymphoid cells—a proposal for uniform nomenclature. Nature Rev. Immunol. 13, 145–149 (2013)
Walker, J. A., Barlow, J. L. & McKenzie, A. N. Innate lymphoid cells—how did we miss them? Nature Rev. Immunol. 13, 75–87 (2013)
Geremia, A. et al. IL-23-responsive innate lymphoid cells are increased in inflammatory bowel disease. J. Exp. Med. 208, 1127–1133 (2011)
Takayama, T. et al. Imbalance of NKp44+NKp46− and NKp44−NKp46+ natural killer cells in the intestinal mucosa of patients with Crohn’s disease. Gastroenterology 139, 882–892 (2010)
Ciccia, F. et al. Interleukin-22 and IL-22-producing NKp44+ NK cells in the subclinical gut inflammation of patients with ankylosing spondylitis. Arthritis Rheum. 64, 1869–1878 (2011)
Sonnenberg, G. F., Fouser, L. A. & Artis, D. Border patrol: regulation of immunity, inflammation and tissue homeostasis at barrier surfaces by IL-22. Nature Immunol. 12, 383–390 (2011)
Eberl, G. & Littman, D. R. Thymic origin of intestinal αβ T cells revealed by fate mapping of RORγt+ cells. Science 305, 248–251 (2004)
Sawa, S. et al. Lineage relationship analysis of RORγt+ innate lymphoid cells. Science 330, 665–669 (2010)
Monticelli, L. A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nature Immunol. 12, 1045–1054 (2011)
Reynders, A. et al. Identity, regulation and in vivo function of gut NKp46+RORγt+ and NKp46+RORγt− lymphoid cells. EMBO J. 30, 2934–2947 (2011)
Yosef, N. et al. Dynamic regulatory network controlling T17 cell differentiation. Nature 496, 461–468 (2013)
Bezman, N. A. et al. Molecular definition of the identity and activation of natural killer cells. Nature Immunol. 13, 1000–1009 (2012)
Klose, C. S. et al. A T-bet gradient controls the fate and function of CCR6-RORγt+ innate lymphoid cells. Nature 494, 261–265 (2013)
Schwartz, R. H. T cell anergy. Annu. Rev. Immunol. 21, 305–334 (2003)
Cong, Y., Feng, T., Fujihashi, K., Schoeb, T. R. & Elson, C. O. A dominant, coordinated T regulatory cell-IgA response to the intestinal microbiota. Proc. Natl Acad. Sci. USA 106, 19256–19261 (2009)
Benoist, C. & Mathis, D. Regulation of major histocompatibility complex class-II genes: X, Y and other letters of the alphabet. Annu. Rev. Immunol. 8, 681–715 (1990)
Mebius, R. E., Rennert, P. & Weissman, I. L. Developing lymph nodes collect CD4+CD3- LTbeta+ cells that can differentiate to APC, NK cells, and follicular cells but not T or B cells. Immunity 7, 493–504 (1997)
Eberl, G. et al. An essential function for the nuclear receptor RORγt in the generation of fetal lymphoid tissue inducer cells. Nature Immunol. 5, 64–73 (2004)
Abt, M. C. et al. Commensal bacteria calibrate the activation threshold of innate antiviral immunity. Immunity 37, 158–170 (2012)
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007)
Huang, D. W. et al. Extracting biological meaning from large gene lists with DAVID. Curr. Protoc. Bioinform. Ch. 13, Unit 13 11. (2009)
Battke, F., Symons, S. & Nieselt, K. Mayday–integrative analytics for expression data. BMC Bioinformatics 11, 121 (2010)
Murphy, D. B. et al. A novel MHC class II epitope expressed in thymic medulla but not cortex. Nature 338, 765–768 (1989)
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nature Methods 7, 335–336 (2010)
Acknowledgements
We thank members of the Sonnenberg and Artis laboratories for discussions and critical reading of the manuscript. We also thank H. L. Ma, L. A. Fouser, S. Olland, R. Zollner, K. Lam and A. Root at Pfizer for critical discussions, valuable advice and the preparation of IL-22 antibodies; M. M. Elloso at Janssen Research and Development for critical discussions, valuable advice and the preparation of IL-17 and IL-23 antibodies; and M.S. Marks for providing the E-alpha protein and Y-Ae antibody. The research is supported by the National Institutes of Health (AI061570, AI087990, AI074878, AI095776, AI102942, AI095466, AI095608 and AI097333 to D.A.; T32-AI055428 to L.A.M.; DK071176 to C.O.E.; and DP5OD012116 to G.F.S.), the Crohn’s and Colitis Foundation of America (to D.A.) and the Burroughs Wellcome Fund Investigator in Pathogenesis of Infectious Disease Award (to D.A.). We also thank the Matthew J. Ryan Veterinary Hospital Pathology Lab, the National Institute of Diabetes and Digestive and Kidney Disease Center for the Molecular Studies in Digestive and Liver Disease Molecular Pathology and Imaging Core (P30DK50306), the Penn Microarray Facility and the Abramson Cancer Center Flow Cytometry and Cell Sorting Resource Laboratory (partially supported by NCI Comprehensive Cancer Center Support Grant (2-P30 CA016520)) for technical advice and support. Human tissue samples were provided by the Cooperative Human Tissue Network, which is funded by the National Cancer Institute.
Author information
Authors and Affiliations
Contributions
M.R.H., L.A.M., T.C.F., D.A. and G.F.S. designed and performed the research. C.G.K.Z. performed analyses of microarray data. A.C., J.M.A. and E.J.W. performed the microarray of naive CD4+ T cells. S.G., R.S. and F.D.B. provided advice, performed and analysed 454 pyrosequencing of intestinal commensal bacteria. A.R.M. assisted with reagents and advice for antigen presentation assays. H.-L.M., P.A.K., C.O.E. and G.E. provided essential mouse strains, valuable advice and technical expertise for these studies. M.R.H., L.A.M., T.C.F., D.A. and G.F.S. analysed the data. M.R.H., D.A. and G.F.S. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-13. (PDF 1812 kb)
Rights and permissions
About this article
Cite this article
Hepworth, M., Monticelli, L., Fung, T. et al. Innate lymphoid cells regulate CD4+ T-cell responses to intestinal commensal bacteria. Nature 498, 113–117 (2013). https://doi.org/10.1038/nature12240
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12240
This article is cited by
-
Increased regulatory activity of intestinal innate lymphoid cells type 3 (ILC3) prevents experimental autoimmune encephalomyelitis severity
Journal of Neuroinflammation (2024)
-
The emerging family of RORγt+ antigen-presenting cells
Nature Reviews Immunology (2024)
-
Disease pathogenesis and barrier functions regulated by group 3 innate lymphoid cells
Seminars in Immunopathology (2024)
-
The crosstalk between enteric nervous system and immune system in intestinal development, homeostasis and diseases
Science China Life Sciences (2024)
-
Virulence factors and mechanisms of paediatric pneumonia caused by Enterococcus faecalis
Gut Pathogens (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.