Abstract
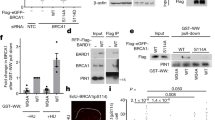
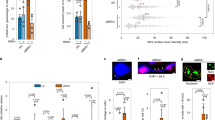
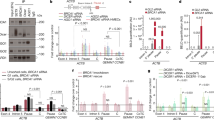
Recent exon-sequencing studies of human tumours have revealed that subunits of BAF (mammalian SWI/SNF) complexes are mutated in more than 20% of all human malignancies1,2, but the mechanisms involved in tumour suppression are unclear. BAF chromatin-remodelling complexes are polymorphic assemblies that use energy provided by ATP hydrolysis to regulate transcription through the control of chromatin structure3 and the placement of Polycomb repressive complex 2 (PRC2) across the genome4,5. Several proteins dedicated to this multisubunit complex, including BRG1 (also known as SMARCA4) and BAF250a (also known as ARID1A), are mutated at frequencies similar to those of recognized tumour suppressors. In particular, the core ATPase BRG1 is mutated in 5–10% of childhood medulloblastomas6,7,8,9 and more than 15% of Burkitt’s lymphomas10,11. Here we show a previously unknown function of BAF complexes in decatenating newly replicated sister chromatids, a requirement for proper chromosome segregation during mitosis. We find that deletion of Brg1 in mouse cells, as well as the expression of BRG1 point mutants identified in human tumours, leads to anaphase bridge formation (in which sister chromatids are linked by catenated strands of DNA) and a G2/M-phase block characteristic of the decatenation checkpoint. Endogenous BAF complexes interact directly with endogenous topoisomerase IIα (TOP2A) through BAF250a and are required for the binding of TOP2A to approximately 12,000 sites across the genome. Our results demonstrate that TOP2A chromatin binding is dependent on the ATPase activity of BRG1, which is compromised in oncogenic BRG1 mutants. These studies indicate that the ability of TOP2A to prevent DNA entanglement at mitosis requires BAF complexes and suggest that this activity contributes to the role of BAF subunits as tumour suppressors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kadoch, C. et al. Proteomic and bioinformatic analysis of mammalian SWI/SNF complexes identifies extensive roles in human malignancy. Nature Genet. http://dx.doi.org/10.1038/ng.2628 (5 May 2013)
You, J. S. & Jones, P. A. Cancer genetics and epigenetics: two sides of the same coin? Cancer Cell 22, 9–20 (2012)
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009)
Ho, L. et al. esBAF facilitates pluripotency by conditioning the genome for LIF/STAT3 signalling and by regulating polycomb function. Nature Cell Biol. 13, 903–913 (2011)
Wilson, B. G. et al. Epigenetic antagonism between polycomb and SWI/SNF complexes during oncogenic transformation. Cancer Cell 18, 316–328 (2010)
Parsons, D. W. et al. The genetic landscape of the childhood cancer medulloblastoma. Science 331, 435–439 (2011)
Pugh, T. J. et al. Medulloblastoma exome sequencing uncovers subtype-specific somatic mutations within a broad landscape of genetic heterogeneity. Nature 488, 106–110 (2012)
Jones, D. T. W. et al. Dissecting the genomic complexity underlying medulloblastoma. Nature 488, 100–105 (2012)
Robinson, G. et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 488, 43–48 (2012)
Love, C. et al. The genetic landscape of mutations in Burkitt lymphoma. Nature Genet. 44, 1321–1325 (2012)
Richter, J. et al. Recurrent mutation of the ID3 gene in Burkitt lymphoma identified by integrated genome, exome and transcriptome sequencing. Nature Genet. 44, 1316–1320 (2012)
Carpenter, A. J. & Porter, A. C. Construction, characterization, and complementation of a conditional-lethal DNA topoisomerase IIα mutant human cell line. Mol. Biol. Cell 15, 5700–5711 (2004)
Lou, Z., Minter-Dykhouse, K. & Chen, J. BRCA1 participates in DNA decatenation. Nature Struct. Mol. Biol. 12, 589–593 (2005)
Dawlaty, M. M. et al. Resolution of sister centromeres requires RanBP2-mediated SUMOylation of topoisomerase IIα. Cell 133, 103–115 (2008)
Ramamoorthy, M. et al. RECQL5 cooperates with Topoisomerase II alpha in DNA decatenation and cell cycle progression. Nucleic Acids Res. 40, 1621–1635 (2012)
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is essential for embryonic stem cell self-renewal and pluripotency. Proc. Natl Acad. Sci. USA 106, 5181–5186 (2009)
Johnson, M., Phua, H. H., Bennett, S. C., Spence, J. M. & Farr, C. J. Studying vertebrate topoisomerase 2 function using a conditional knockdown system in DT40 cells. Nucleic Acids Res. 37, e98 (2009)
Downes, C. S. et al. A topoisomerase II-dependent G2 cycle checkpoint in mammalian cells. Nature 372, 467–470 (1994)
Luo, K., Yuan, J., Chen, J. & Lou, Z. Topoisomerase IIα controls the decatenation checkpoint. Nature Cell Biol. 11, 204–210 (2009)
Sakaguchi, A. & Kikuchi, A. Functional compatibility between isoform α and β of type II DNA topoisomerase. J. Cell Sci. 117, 1047–1054 (2004)
Khavari, P. A., Peterson, C. L., Tamkun, J. W., Mendel, D. B. & Crabtree, G. R. BRG1 contains a conserved domain of the SWI2/SNF2 family necessary for normal mitotic growth and transcription. Nature 366, 170–174 (1993)
Kool, M. et al. Molecular subgroups of medulloblastoma: an international meta-analysis of transcriptome, genetic aberrations, and clinical data of WNT, SHH, Group 3, and Group 4 medulloblastomas. Acta Neuropathol. 123, 473–484 (2012)
Northcott, P. A. et al. Medulloblastomics: the end of the beginning. Nature Rev. Cancer 12, 818–834 (2012)
Ho, L. et al. An embryonic stem cell chromatin remodeling complex, esBAF, is an essential component of the core pluripotency transcriptional network. Proc. Natl Acad. Sci. USA 106, 5187–5191 (2009)
Stros, M., Bacikova, A., Polanska, E., Stokrova, J. & Strauss, F. HMGB1 interacts with human topoisomerase IIα and stimulates its catalytic activity. Nucleic Acids Res. 35, 5001–5013 (2007)
Sano, K., Miyaji-Yamaguchi, M., Tsutsui, K. M. & Tsutsui, K. Topoisomerase IIβ activates a subset of neuronal genes that are repressed in AT-rich genomic environment. PLoS ONE 3, e4103 (2008)
Capranico, G., Jaxel, C., Roberge, M., Kohn, K. W. & Pommier, Y. Nucleosome positioning as a critical determinant for the DNA cleavage sites of mammalian DNA topoisomerase II in reconstituted simian virus 40 chromatin. Nucleic Acids Res. 18, 4553–4559 (1990)
Sperling, A. S., Jeong, K. S., Kitada, T. & Grunstein, M. Topoisomerase II binds nucleosome-free DNA and acts redundantly with topoisomerase I to enhance recruitment of RNA Pol II in budding yeast. Proc. Natl Acad. Sci. USA 108, 12693–12698 (2011)
Bourgo, R. J. et al. SWI/SNF deficiency results in aberrant chromatin organization, mitotic failure, and diminished proliferative capacity. Mol. Biol. Cell 20, 3192–3199 (2009)
Janssen, A., van der Burg, M., Szuhai, K., Kops, G. J. & Medema, R. H. Chromosome segregation errors as a cause of DNA damage and structural chromosome aberrations. Science 333, 1895–1898 (2011)
Bultman, S. J., Gebuhr, T. C. & Magnuson, T. A. Brg1 mutation that uncouples ATPase activity from chromatin remodeling reveals an essential role for SWI/SNF-related complexes in β-globin expression and erythroid development. Genes Dev. 19, 2849–2861 (2005)
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010)
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002)
Meyer, L. R. et al. The UCSC Genome Browser database: extensions and updates 2013. Nucleic Acids Res. 41, D64–D69 (2012)
Acknowledgements
We would like to acknowledge the DNA Sequencing Core facility of the National Heart, Lung, and Blood Institute (NHLBI) for sequencing the ChIP-seq libraries. This work was supported by the NIH (G.R.C.), The American Cancer Society (E.C.D.), The Helen Hay Whitney Foundation (D.C.H.) and the Division of Intramural Research program of NHLBI (K.Z.). G.R.C. is an investigator at the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
E.C.D., D.C.H. and G.R.C. designed the experiments, and E.C.D. and D.C.H. performed the experiments. E.M. performed bioinformatic analysis. A.K., M.K., S.P. and Y-J.C. contributed human tumour samples. K.Z. and K.C. performed sequencing of the TOP2A ChIP. E.C.D., D.C.H. and G.R.C. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-6. (PDF 2922 kb)
Rights and permissions
About this article
Cite this article
Dykhuizen, E., Hargreaves, D., Miller, E. et al. BAF complexes facilitate decatenation of DNA by topoisomerase IIα. Nature 497, 624–627 (2013). https://doi.org/10.1038/nature12146
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12146
This article is cited by
-
Brg1 controls stemness and metastasis of pancreatic cancer through regulating hypoxia pathway
Oncogene (2023)
-
Clinical and translational advances in ovarian cancer therapy
Nature Cancer (2023)
-
Targeting replication stress in cancer therapy
Nature Reviews Drug Discovery (2023)
-
Nuclear architecture and the structural basis of mitotic memory
Chromosome Research (2023)
-
Treating ARID1A mutated cancers by harnessing synthetic lethality and DNA damage response
Journal of Biomedical Science (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.