Abstract
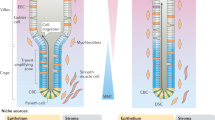
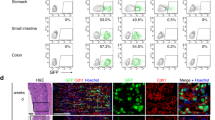
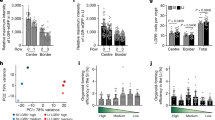
The rapid cell turnover of the intestinal epithelium is achieved from small numbers of stem cells located in the base of glandular crypts. These stem cells have been variously described as rapidly cycling or quiescent. A functional arrangement of stem cells that reconciles both of these behaviours has so far been difficult to obtain. Alternative explanations for quiescent cells have been that they act as a parallel or reserve population that replace rapidly cycling stem cells periodically or after injury; their exact nature remains unknown. Here we show mouse intestinal quiescent cells to be precursors that are committed to mature into differentiated secretory cells of the Paneth and enteroendocrine lineage. However, crucially we find that after intestinal injury they are capable of extensive proliferation and can give rise to clones comprising the main epithelial cell types. Thus, quiescent cells can be recalled to the stem-cell state. These findings establish quiescent cells as an effective clonogenic reserve and provide a motivation for investigating their role in pathologies such as colorectal cancers and intestinal inflammation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007)
Roth, S. et al. Paneth cells in intestinal homeostasis and tissue injury. PLoS ONE 7, e38965 (2012)
Powell, A. E. et al. The Pan-ErbB negative regulator Lrig1 is an intestinal stem cell marker that functions as a tumor suppressor. Cell 149, 146–158 (2012)
Sangiorgi, E. & Capecchi, M. R. Bmi1 is expressed in vivo in intestinal stem cells. Nature Genet. 40, 915–920 (2008)
Montgomery, R. K. et al. Mouse telomerase reverse transcriptase (mTert) expression marks slowly cycling intestinal stem cells. Proc. Natl Acad. Sci. USA 108, 179–184 (2011)
Takeda, N. et al. Interconversion between intestinal stem cell populations in distinct niches. Science 334, 1420–1424 (2011)
Wong, V. W. et al. Lrig1 controls intestinal stem-cell homeostasis by negative regulation of ErbB signalling. Nature Cell Biol. 14, 401–408 (2012)
Lopez-Garcia, C., Klein, A. M., Simons, B. D. & Winton, D. J. Intestinal stem cell replacement follows a pattern of neutral drift. Science 330, 822–825 (2010)
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010)
Li, L. & Clevers, H. Coexistence of quiescent and active adult stem cells in mammals. Science 327, 542–545 (2010)
Tian, H. et al. A reserve stem cell population in small intestine renders Lgr5-positive cells dispensable. Nature 478, 255–259 (2011)
Potten, C. S., Hume, W. J., Reid, P. & Cairns, J. The segregation of DNA in epithelial stem cells. Cell 15, 899–906 (1978)
Potten, C. S., Owen, G. & Booth, D. Intestinal stem cells protect their genome by selective segregation of template DNA strands. J. Cell Sci. 115, 2381–2388 (2002)
Demidov, O. N. et al. Wip1 phosphatase regulates p53-dependent apoptosis of stem cells and tumorigenesis in the mouse intestine. Cell Stem Cell 1, 180–190 (2007)
He, X. C. et al. BMP signaling inhibits intestinal stem cell self-renewal through suppression of Wnt-β-catenin signaling. Nature Genet. 36, 1117–1121 (2004)
He, X. C. et al. PTEN-deficient intestinal stem cells initiate intestinal polyposis. Nature Genet. 39, 189–198 (2007)
Muñoz, J. et al. The Lgr5 intestinal stem cell signature: robust expression of proposed quiescent '+4' cell markers. EMBO J. 31, 3079–3091 (2012)
Ireland, H., Houghton, C., Howard, L. & Winton, D. J. Cellular inheritance of a Cre-activated reporter gene to determine Paneth cell longevity in the murine small intestine. Dev. Dyn. 233, 1332–1336 (2005)
Foudi, A. et al. Analysis of histone 2B-GFP retention reveals slowly cycling hematopoietic stem cells. Nature Biotechnol. 27, 84–90 (2009)
Sato, T. et al. Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature 469, 415–418 (2011)
Cheng, H. & Leblond, C. P. Origin, differentiation and renewal of the four main epithelial cell types in the mouse small intestine. V. Unitarian Theory of the origin of the four epithelial cell types. Am. J. Anat. 141, 537–561 (1974)
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009)
Jullien, N., Sampieri, F., Enjalbert, A. & Herman, J. P. Regulation of Cre recombinase by ligand-induced complementation of inactive fragments. Nucleic Acids Res. 31, e131 (2003)
van Es, J. H. et al. Dll1+ secretory progenitor cells revert to stem cells upon crypt damage. Nature Cell Biol. 14, 1099–1104 (2012)
Lewis, A. et al. Severe polyposis in Apc(1322T) mice is associated with submaximal Wnt signalling and increased expression of the stem cell marker Lgr5. Gut 59, 1680–1686 (2010)
Campbell, S. J. et al. Regulation of the CYP1A1 promoter in transgenic mice: an exquisitely sensitive on-off system for cell specific gene regulation. J. Cell Sci. 109, 2619–2625 (1996)
Vooijs, M., Jonkers, J. & Berns, A. A highly efficient ligand-regulated Cre recombinase mouse line shows that LoxP recombination is position dependent. EMBO Rep. 2, 292–297 (2001)
McCutcheon, S. C. et al. Characterization of a heat resistant β-glucosidase as a new reporter in cells and mice. BMC Biol. 8, 89 (2010)
Gracz, A. D., Ramalingam, S. & Magness, S. T. Sox9 expression marks a subset of CD24-expressing small intestine epithelial stem cells that form organoids in vitro. Am. J. Physiol. Gastrointest. Liver Physiol. 298, G590–G600 (2010)
Potten, C. S. & Loeffler, M. Stem cells: attributes, cycles, spirals, pitfalls and uncertainties. Lessons for and from the crypt. Development 110, 1001–1020 (1990)
Bjerknes, M. & Cheng, H. Methods for the isolation of intact epithelium from the mouse intestine. Anat. Rec. 199, 565–574 (1981)
Irizarry, R. A. et al. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 31, e15 (2003)
Pruitt, K. D., Tatusova, T., Klimke, W. & Maglott, D. R. NCBI Reference Sequences: current status, policy and new initiatives. Nucleic Acids Res. 37, D32–D36 (2009)
Lockstone, H. E. Exon array data analysis using Affymetrix power tools and R statistical software. Brief. Bioinform. 12, 634–644 (2011)
R Development Core Team. R: A Language and Environment for Statistical Computing (2009)
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
Smyth, G. K. in Bioinformatics and Computational Biology Solutions using R and Bioconductor Solutions using R and Bioconductor (eds Carey, V., Gentleman, R., Dudoit, S., Huber, W. & Izzarry, R.) 397–420 (Springer, 2005)
Jensen, K. B. & Watt, F. M. Single-cell expression profiling of human epidermal stem and transit-amplifying cells: Lrig1 is a regulator of stem cell quiescence. Proc. Natl Acad. Sci. USA 103, 11958–11963 (2006)
Acknowledgements
This research was supported by Cancer Research UK (S.J.A.B., H.I.Z., A.M.N., R.R., R.K. and D.J.W.). L.V. was supported by a KWF fellowship. We thank D. Tan for providing the single-cell RNA amplification protocol. We also thank M. de la Roche, R. von Furstenberg and S. Henning for advice with the in vitro culture work. We acknowledge the following core facilities at CRUK/CRI: The Transgenic Laboratory, Biological Resource Unit, Flow Cytometry, Histopathology, Microscopy, Genomics and Bioinformatics and the CRUK Paterson Institute Microarray Facility. We thank R. J. Davies, A. Klein and A. Ibrahim for manuscript discussions.
Author information
Authors and Affiliations
Contributions
S.J.A.B. designed and performed experiments and wrote the paper. H.I.Z. designed and developed the H2B–YFP model and performed experiments. R.R. performed the bioinformatic analysis. A.M.N. and L.V. performed experiments. R.K. designed experiments and performed bioinformatic analysis. D.J.W. designed experiments, developed the diCreAB model and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6. (PDF 2399 kb)
H2B-YFP Paneth cells are non-responsive to βNF induction
Confocal reconstruction video of an isolated crypt immediately following βNF treatment, showing all cells other than Paneth cells express YFP. Yellow=YFP. Red=UEA. (MOV 4370 kb)
Rights and permissions
About this article
Cite this article
Buczacki, S., Zecchini, H., Nicholson, A. et al. Intestinal label-retaining cells are secretory precursors expressing Lgr5. Nature 495, 65–69 (2013). https://doi.org/10.1038/nature11965
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11965
This article is cited by
-
Elevated stress response marks deeply quiescent reserve cells of gastric chief cells
Communications Biology (2023)
-
Advanced Progression for the Heterogeneity and Homeostasis of Intestinal Stem Cells
Stem Cell Reviews and Reports (2023)
-
Stem cell activation during distraction enterogenesis in the murine colon
Pediatric Surgery International (2023)
-
Adult stem cell activity in naked mole rats for long-term tissue maintenance
Nature Communications (2023)
-
Challenges and opportunities targeting mechanisms of epithelial injury and recovery in acute intestinal graft-versus-host disease
Mucosal Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.