Abstract
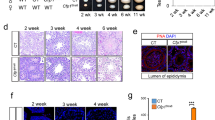
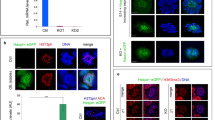
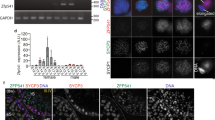
Meiosis is a germ-cell-specific cell division process through which haploid gametes are produced for sexual reproduction1. Before the initiation of meiosis, mouse primordial germ cells undergo a series of epigenetic reprogramming steps2,3, including the global erasure of DNA methylation at the 5-position of cytosine (5mC) in CpG-rich DNA4,5. Although several epigenetic regulators, such as Dnmt3l and the histone methyltransferases G9a and Prdm9, have been reported to be crucial for meiosis6, little is known about how the expression of meiotic genes is regulated and how their expression contributes to normal meiosis. Using a loss-of-function approach in mice, here we show that the 5mC-specific dioxygenase Tet1 has an important role in regulating meiosis in mouse oocytes. Tet1 deficiency significantly reduces female germ-cell numbers and fertility. Univalent chromosomes and unresolved DNA double-strand breaks are also observed in Tet1-deficient oocytes. Tet1 deficiency does not greatly affect the genome-wide demethylation that takes place in primordial germ cells, but leads to defective DNA demethylation and decreased expression of a subset of meiotic genes. Our study thus establishes a function for Tet1 in meiosis and meiotic gene activation in female germ cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Handel, M. A. & Schimenti, J. C. Genetics of mammalian meiosis: regulation, dynamics and impact on fertility. Nature Rev. Genet. 11, 124–136 (2010)
Hayashi, K. & Surani, M. A. Resetting the epigenome beyond pluripotency in the germline. Cell Stem Cell 4, 493–498 (2009)
Sasaki, H. & Matsui, Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nature Rev. Genet. 9, 129–140 (2008)
Hajkova, P. et al. Epigenetic reprogramming in mouse primordial germ cells. Mech. Dev. 117, 15–23 (2002)
Seki, Y. et al. Extensive and orderly reprogramming of genome-wide chromatin modifications associated with specification and early development of germ cells in mice. Dev. Biol. 278, 440–458 (2005)
Kota, S. K. & Feil, R. Epigenetic transitions in germ cell development and meiosis. Dev. Cell 19, 675–686 (2010)
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011)
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010)
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011)
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009)
Dawlaty, M. M. et al. Tet1 is dispensable for maintaining pluripotency and its loss is compatible with embryonic and postnatal development. Cell Stem Cell 9, 166–175 (2011)
Roeder, G. S. & Bailis, J. M. The pachytene checkpoint. Trends Genet. 16, 395–403 (2000)
Edelmann, W. et al. Meiotic pachytene arrest in MLH1-deficient mice. Cell 85, 1125–1134 (1996)
Takada, Y. et al. HP1γ links histone methylation marks to meiotic synapsis in mice. Development 138, 4207–4217 (2011)
Tachibana, M., Nozaki, M., Takeda, N. & Shinkai, Y. Functional dynamics of H3K9 methylation during meiotic prophase progression. EMBO J. 26, 3346–3359 (2007)
Williams, K. et al. TET1 and hydroxymethylcytosine in transcription and DNA methylation fidelity. Nature 473, 343–348 (2011)
Wu, H. et al. Dual functions of Tet1 in transcriptional regulation in mouse embryonic stem cells. Nature 473, 389–393 (2011)
Ramskold, D. et al. Full-length mRNA-Seq from single-cell levels of RNA and individual circulating tumor cells. Nature Biotechnol. 30, 777–782 (2012)
de Vries, F. A. T. et al. Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev. 19, 1376–1389 (2005)
Hayashi, K., Yoshida, K. & Matsui, Y. A histone H3 methyltransferase controls epigenetic events required for meiotic prophase. Nature 438, 374–378 (2005)
Soper, S. F. et al. Mouse maelstrom, a component of nuage, is essential for spermatogenesis and transposon repression in meiosis. Dev. Cell 15, 285–297 (2008)
Wang, H. & Höög, C. Structural damage to meiotic chromosomes impairs DNA recombination and checkpoint control in mammalian oocytes. J. Cell Biol. 173, 485–495 (2006)
Adey, A. & Shendure, J. Ultra-low-input, tagmentation-based whole-genome bisulfite sequencing. Genome Res. 22, 1139–1143 (2012)
Popp, C. et al. Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463, 1101–1105 (2010)
Maatouk, D. M. et al. DNA methylation is a primary mechanism for silencing postmigratory primordial germ cell genes in both germ cell and somatic cell lineages. Development 133, 3411–3418 (2006)
Dahl, J. A. & Collas, P. A rapid micro chromatin immunoprecipitation assay (microChIP). Nature Protocols 3, 1032–1045 (2008)
Markoulaki, S., Meissner, A. & Jaenisch, R. Somatic cell nuclear transfer and derivation of embryonic stem cells in the mouse. Methods 45, 101–114 (2008)
Soper, S. F. C. et al. Mouse maelstrom, a component of nuage, is essential for spermatogenesis and transposon repression in meiosis. Dev. Cell 15, 285–297 (2008)
Kouznetsova, A., Novak, I., Jessberger, R. & Hoog, C. SYCP2 and SYCP3 are required for cohesin core integrity at diplotene but not for centromere cohesion at the first meiotic division. J. Cell Sci. 118, 2271–2278 (2005)
Diep, D. et al. Library-free methylation sequencing with bisulfite padlock probes. Nature Methods 9, 270–272 (2012)
Acknowledgements
We thank J. Wang and W. Jiang for FACS sorting, A. Adey and J. Shendure for sharing reagents and protocols for Tn5mC-seq, and H.-L. Fung for Illumina sequencing. This work was partially supported by U01DK089565 (National Institutes of Health) (to Y.Z.), R01GM097253 and CIRM BRB3-05083 (to K.Z.). Y.Z. is an Investigator of the Howard Hughes Medical Institute. D.D. is a CIRM-UCSD pre-doctoral fellow.
Author information
Authors and Affiliations
Contributions
Y.Z. conceived the project; S.Y., K.H. and Y.Z. designed the experiments; S.Y., K.H., R.L., L.S. and A.I. performed the experiments; S.Y., K.H., R.L., D.D., K.Z. and Y.Z. analysed and interpreted the data; S.Y., K.H., R.L., K.Z. and Y.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-20 and Supplementary Tables 1, 2, 4 and 7 (see separate files for Supplementary Tables 3, 5 and 6). (PDF 13096 kb)
Supplementary Table 3
This file contains a list of differentially expressed genes (DEG) based on RNAseq. (XLSX 115 kb)
Supplementary Table 5
This table contains a List of differentially methylated regions (DMR) in female Tet1Gt/Gt PGC. (XLSX 296 kb)
Supplementary Table 6
This table contains a List of differentially expressed genes (DEG) associated with DMR. (XLSX 46 kb)
Rights and permissions
About this article
Cite this article
Yamaguchi, S., Hong, K., Liu, R. et al. Tet1 controls meiosis by regulating meiotic gene expression. Nature 492, 443–447 (2012). https://doi.org/10.1038/nature11709
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11709
This article is cited by
-
Dynamics of DNA hydroxymethylation and methylation during mouse embryonic and germline development
Nature Genetics (2023)
-
PRC1-mediated epigenetic programming is required to generate the ovarian reserve
Nature Communications (2022)
-
Genome-Wide Differential DNA Methylomes Provide Insights into the Infertility of Triploid Oysters
Marine Biotechnology (2022)
-
Reconstitution of the oocyte transcriptional network with transcription factors
Nature (2021)
-
Roles of Tet2 in meiosis, fertility and reproductive aging
Protein & Cell (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.